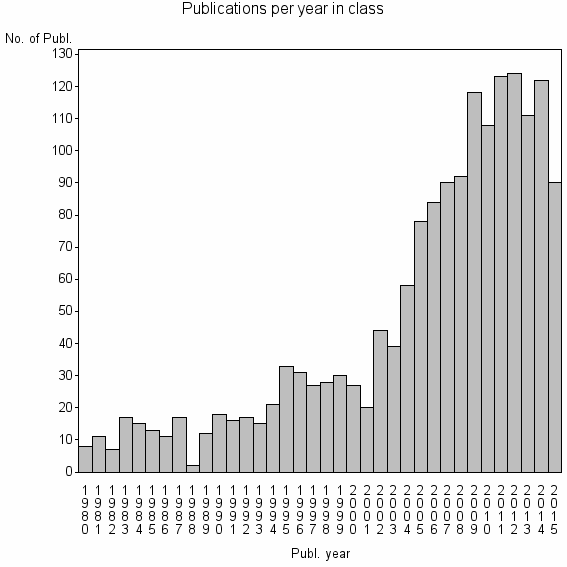
Class information for: |
Basic class information |
| ID | Publications | Average number of references |
Avg. shr. active ref. in WoS |
|---|---|---|---|
| 4652 | 1677 | 47.8 | 81% |
Classes in level above (level 2) |
| ID, lev. above |
Publications | Label for level above |
|---|---|---|
| 42 | 31225 | PROTEINS-STRUCTURE FUNCTION AND BIOINFORMATICS//PROTEIN STRUCTURE PREDICTION//PROTEIN FOLDING |
Terms with highest relevance score |
| Rank | Term | Type of term | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
|---|---|---|---|---|---|---|
| 1 | ELASTIC NETWORK MODEL | Author keyword | 45 | 60% | 3% | 50 |
| 2 | ELASTIC NETWORK MODELS | Author keyword | 44 | 88% | 1% | 21 |
| 3 | NORMAL MODE ANALYSIS | Author keyword | 37 | 30% | 6% | 102 |
| 4 | GAUSSIAN NETWORK MODEL | Author keyword | 20 | 59% | 1% | 23 |
| 5 | LH BAKER BIOINFORMAT BIOL STAT | Address | 12 | 42% | 1% | 23 |
| 6 | ELASTIC NETWORK | Author keyword | 11 | 57% | 1% | 13 |
| 7 | B FACTORS | Author keyword | 9 | 47% | 1% | 15 |
| 8 | MEAN SQUARE FLUCTUATIONS | Author keyword | 9 | 83% | 0% | 5 |
| 9 | NETWORK RIGIDITY | Author keyword | 8 | 75% | 0% | 6 |
| 10 | SACKLER MOL MED | Address | 8 | 15% | 3% | 48 |
Web of Science journal categories |
Author Key Words |
| Rank | Web of Science journal category | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
LCSH search | Wikipedia search |
|---|---|---|---|---|---|---|---|
| 1 | ELASTIC NETWORK MODEL | 45 | 60% | 3% | 50 | Search ELASTIC+NETWORK+MODEL | Search ELASTIC+NETWORK+MODEL |
| 2 | ELASTIC NETWORK MODELS | 44 | 88% | 1% | 21 | Search ELASTIC+NETWORK+MODELS | Search ELASTIC+NETWORK+MODELS |
| 3 | NORMAL MODE ANALYSIS | 37 | 30% | 6% | 102 | Search NORMAL+MODE+ANALYSIS | Search NORMAL+MODE+ANALYSIS |
| 4 | GAUSSIAN NETWORK MODEL | 20 | 59% | 1% | 23 | Search GAUSSIAN+NETWORK+MODEL | Search GAUSSIAN+NETWORK+MODEL |
| 5 | ELASTIC NETWORK | 11 | 57% | 1% | 13 | Search ELASTIC+NETWORK | Search ELASTIC+NETWORK |
| 6 | B FACTORS | 9 | 47% | 1% | 15 | Search B+FACTORS | Search B+FACTORS |
| 7 | MEAN SQUARE FLUCTUATIONS | 9 | 83% | 0% | 5 | Search MEAN+SQUARE+FLUCTUATIONS | Search MEAN+SQUARE+FLUCTUATIONS |
| 8 | NETWORK RIGIDITY | 8 | 75% | 0% | 6 | Search NETWORK+RIGIDITY | Search NETWORK+RIGIDITY |
| 9 | ESSENTIAL DYNAMICS | 7 | 23% | 2% | 28 | Search ESSENTIAL+DYNAMICS | Search ESSENTIAL+DYNAMICS |
| 10 | HINGE BENDING | 7 | 39% | 1% | 14 | Search HINGE+BENDING | Search HINGE+BENDING |
Key Words Plus |
| Rank | Web of Science journal category | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
|---|---|---|---|---|---|
| 1 | SINGLE PARAMETER | 295 | 72% | 14% | 230 |
| 2 | ELASTIC NETWORK MODEL | 144 | 68% | 8% | 126 |
| 3 | FREQUENCY NORMAL MODES | 126 | 67% | 7% | 113 |
| 4 | DOMAIN MOTIONS | 60 | 46% | 6% | 96 |
| 5 | COLLECTIVE MOTIONS | 57 | 49% | 5% | 85 |
| 6 | NORMAL MODE ANALYSIS | 55 | 26% | 11% | 185 |
| 7 | ELASTIC NETWORK MODELS | 52 | 64% | 3% | 51 |
| 8 | QUANTITATIVE STABILITY FLEXIBILITY RELATIONSHIPS | 35 | 89% | 1% | 16 |
| 9 | MACROMOLECULAR MOTIONS | 29 | 54% | 2% | 38 |
| 10 | FOLDED PROTEINS | 25 | 30% | 4% | 70 |
Journals |
Reviews |
| Title | Publ. year | Cit. | Active references | % act. ref. to same field |
|---|---|---|---|---|
| Comparing the intrinsic dynamics of multiple protein structures using elastic network models | 2015 | 4 | 118 | 83% |
| Global Dynamics of Proteins: Bridging Between Structure and Function | 2010 | 151 | 71 | 85% |
| The ensemble nature of allostery | 2014 | 69 | 91 | 24% |
| Coarse-grained normal mode analysis in structural biology | 2005 | 347 | 63 | 84% |
| Induced fit, conformational selection and independent dynamic segments: an extended view of binding events | 2010 | 238 | 62 | 29% |
| Allostery in Disease and in Drug Discovery | 2013 | 62 | 91 | 24% |
| Comparing proteins by their internal dynamics: Exploring structure-function relationships beyond static structural alignments | 2013 | 21 | 155 | 54% |
| Symmetry, form, and shape: Guiding principles for robustness in macromolecular machines | 2006 | 147 | 102 | 67% |
| Normal Mode Analysis of Biomolecular Structures: Functional Mechanisms of Membrane Proteins | 2010 | 125 | 362 | 31% |
| Usefulness and limitations of normal mode analysis in modeling dynamics of biomolecular complexes | 2005 | 263 | 76 | 59% |
Address terms |
| Rank | Address term | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
|---|---|---|---|---|---|
| 1 | LH BAKER BIOINFORMAT BIOL STAT | 12 | 42% | 1.4% | 23 |
| 2 | SACKLER MOL MED | 8 | 15% | 2.9% | 48 |
| 3 | MOL STRUCT SECT | 8 | 45% | 0.8% | 13 |
| 4 | BAKER BIOINFORMAT BIOL SCI | 6 | 100% | 0.2% | 4 |
| 5 | BIOMED HLTH INFORMAT PROGRAM | 4 | 46% | 0.4% | 6 |
| 6 | ITP SB | 3 | 50% | 0.2% | 4 |
| 7 | COMPUTAT INFORMAT BIOL MED TRAINING PROGRAM | 3 | 60% | 0.2% | 3 |
| 8 | COMPUTAT BIOL UNIT UNI | 2 | 67% | 0.1% | 2 |
| 9 | CANC NANOBIOL PROGRAM | 2 | 10% | 1.2% | 20 |
| 10 | QUANTITAT BIOL MODELING INITIAT | 2 | 50% | 0.2% | 3 |
Related classes at same level (level 1) |
| Rank | Relatedness score | Related classes |
|---|---|---|
| 1 | 0.0000108954 | MARKOV STATE MODELS//TRANSITION PATH SAMPLING//TRANSITION PATH THEORY |
| 2 | 0.0000097068 | BIOPHYS MODELING SIMULAT//ALL ATOM RECONSTRUCTION//BIOL MODELING SIMULAT |
| 3 | 0.0000089811 | CORRELATED MUTATIONS//CONTACT PREDICTION//CONTACT MAP PREDICTION |
| 4 | 0.0000087473 | ENVIRONM IND CHEM//ONE STEP PERTURBATION//SAMPL4 |
| 5 | 0.0000087002 | UNITA RIC ROMA TRE//DYNAMICAL TRANSITION//UFR PHITEM |
| 6 | 0.0000082358 | CROSS CORRELATED RELAXATION//N 15 RELAXATION//NMR SPIN RELAXATION |
| 7 | 0.0000077386 | LOOP PREDICTION//LOOP MODELING//STEPWISE FOLDING |
| 8 | 0.0000076985 | CAPRI//PROTEIN PROTEIN DOCKING//PROTEIN DOCKING |
| 9 | 0.0000074766 | PROTEIN HYDRATION//INTERNAL SOLVENT//SOLVENT MODELING |
| 10 | 0.0000072711 | PROTEIN FOLDING//PHI VALUE//CONTACT ORDER |