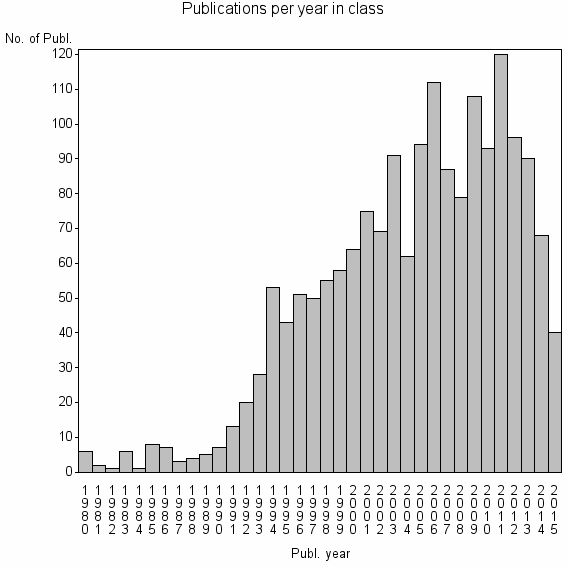
Class information for: |
Basic class information |
| ID | Publications | Average number of references |
Avg. shr. active ref. in WoS |
|---|---|---|---|
| 4108 | 1769 | 27.0 | 80% |
Classes in level above (level 2) |
| ID, lev. above |
Publications | Label for level above |
|---|---|---|
| 1044 | 9634 | ANIMAL GENETICS//GENET CELLULAIRE//ANIM CYTOGENET GENE M PING |
Terms with highest relevance score |
| Rank | Term | Type of term | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
|---|---|---|---|---|---|---|
| 1 | HGEBIET TIERZUCHTUNG BIOTECHNOL | Address | 47 | 88% | 1% | 22 |
| 2 | GENET CELLULAIRE | Address | 41 | 31% | 6% | 111 |
| 3 | ANIMAL GENETICS | Journal | 37 | 11% | 19% | 330 |
| 4 | ANIM BREEDING BIOTECHNOL | Address | 37 | 56% | 2% | 44 |
| 5 | ANIM BIOTECHNOL JIANGXI PROV | Address | 27 | 45% | 3% | 45 |
| 6 | SEZ ALLEVAMENTI ZOOTECN | Address | 18 | 40% | 2% | 35 |
| 7 | INTEGRATED ANIM GENOM | Address | 16 | 29% | 3% | 47 |
| 8 | PROLACTIN RECEPTOR GENE | Author keyword | 13 | 62% | 1% | 13 |
| 9 | KOREAN NATIVE PIG | Author keyword | 12 | 56% | 1% | 15 |
| 10 | MOL BIOL ANIM BREEDING | Address | 12 | 39% | 1% | 24 |
Web of Science journal categories |
Author Key Words |
| Rank | Web of Science journal category | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
LCSH search | Wikipedia search |
|---|---|---|---|---|---|---|---|
| 1 | PROLACTIN RECEPTOR GENE | 13 | 62% | 1% | 13 | Search PROLACTIN+RECEPTOR+GENE | Search PROLACTIN+RECEPTOR+GENE |
| 2 | KOREAN NATIVE PIG | 12 | 56% | 1% | 15 | Search KOREAN+NATIVE+PIG | Search KOREAN+NATIVE+PIG |
| 3 | SSC7 | 9 | 83% | 0% | 5 | Search SSC7 | Search SSC7 |
| 4 | TEAT NUMBER | 9 | 59% | 1% | 10 | Search TEAT+NUMBER | Search TEAT+NUMBER |
| 5 | PRLR | 6 | 41% | 1% | 12 | Search PRLR | Search PRLR |
| 6 | DUROC PIG | 6 | 50% | 0% | 8 | Search DUROC+PIG | Search DUROC+PIG |
| 7 | SSC4 | 6 | 100% | 0% | 4 | Search SSC4 | Search SSC4 |
| 8 | FATNESS | 5 | 13% | 2% | 33 | Search FATNESS | Search FATNESS |
| 9 | COMPARATIVE MAP | 4 | 22% | 1% | 17 | Search COMPARATIVE+MAP | Search COMPARATIVE+MAP |
| 10 | DIVERGENT CROSS | 4 | 75% | 0% | 3 | Search DIVERGENT+CROSS | Search DIVERGENT+CROSS |
Key Words Plus |
| Rank | Web of Science journal category | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
|---|---|---|---|---|---|
| 1 | INFLUENCING ECONOMIC TRAITS | 143 | 86% | 4% | 73 |
| 2 | GENOME SCAN ANALYSIS | 131 | 87% | 4% | 65 |
| 3 | AFFECTING CARCASS COMPOSITION | 89 | 90% | 2% | 38 |
| 4 | ESTROGEN RECEPTOR LOCUS | 82 | 85% | 2% | 44 |
| 5 | PORCINE GENOME | 62 | 65% | 3% | 60 |
| 6 | RESOURCE POPULATION | 57 | 70% | 3% | 47 |
| 7 | BACKFAT THICKNESS | 48 | 49% | 4% | 71 |
| 8 | PIETRAIN RESOURCE POPULATION | 45 | 80% | 2% | 28 |
| 9 | LARGE WHITE INTERCROSS | 43 | 79% | 2% | 27 |
| 10 | MEAT QUALITY TRAITS | 39 | 30% | 6% | 108 |
Journals |
| Rank | Web of Science journal category | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
|---|---|---|---|---|---|
| 1 | ANIMAL GENETICS | 37 | 11% | 19% | 330 |
Reviews |
| Title | Publ. year | Cit. | Active references | % act. ref. to same field |
|---|---|---|---|---|
| THE PIGMAP CONSORTIUM LINKAGE MAP OF THE PIG (SUS SCROFA) | 1995 | 204 | 90 | 71% |
| Advances in QTL mapping in pigs | 2007 | 73 | 17 | 65% |
| A comprehensive map of the porcine genome | 1996 | 326 | 88 | 56% |
| Molecular advances in QTL discovery and application in pig breeding | 2013 | 10 | 92 | 54% |
| Porcine genomics delivers new tools and results: This little piggy did more than just go to market | 2004 | 39 | 23 | 96% |
| QTL and candidate genes for fecundity in sows | 2006 | 27 | 47 | 77% |
| A robust linkage map of the porcine autosomes based on gene-associated SNPs | 2009 | 15 | 86 | 69% |
| The Pig Genome Project Has Plenty to Squeal about | 2011 | 7 | 52 | 60% |
| The IGF1R gene: A new marker for reproductive performance traits in sows? | 2011 | 2 | 13 | 77% |
| Genetics of serum and muscle lipids in pigs | 2013 | 3 | 105 | 49% |
Address terms |
| Rank | Address term | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
|---|---|---|---|---|---|
| 1 | HGEBIET TIERZUCHTUNG BIOTECHNOL | 47 | 88% | 1.2% | 22 |
| 2 | GENET CELLULAIRE | 41 | 31% | 6.3% | 111 |
| 3 | ANIM BREEDING BIOTECHNOL | 37 | 56% | 2.5% | 44 |
| 4 | ANIM BIOTECHNOL JIANGXI PROV | 27 | 45% | 2.5% | 45 |
| 5 | SEZ ALLEVAMENTI ZOOTECN | 18 | 40% | 2.0% | 35 |
| 6 | INTEGRATED ANIM GENOM | 16 | 29% | 2.7% | 47 |
| 7 | MOL BIOL ANIM BREEDING | 12 | 39% | 1.4% | 24 |
| 8 | DIPROVAL | 10 | 24% | 2.0% | 35 |
| 9 | SWINE GENET BREEDING | 8 | 22% | 1.9% | 34 |
| 10 | BREEDING BIOL MOL ANIM BREEDING | 8 | 100% | 0.3% | 5 |
Related classes at same level (level 1) |
| Rank | Relatedness score | Related classes |
|---|---|---|
| 1 | 0.0000108879 | SWINE LEUKOCYTE ANTIGEN//MELIM//SWINE LEUKOCYTE ANTIGENS |
| 2 | 0.0000099840 | ANIM CYTOGENET GENE M PING//ANIMAL GENETICS//BIOCHEM GENET CYTOGENET |
| 3 | 0.0000091053 | PRENATAL SURVIVAL//LITTER SIZE//UTERINE CAPACITY |
| 4 | 0.0000090556 | GROWTH HORMONE GENE//GH GENE//CALPASTATIN GENE |
| 5 | 0.0000089557 | ANIMAL BLOOD GROUPS AND BIOCHEMICAL GENETICS//ALPHA PROTEASE INHIBITORS//ALPHA 1 BETA GLYCOPROTEIN |
| 6 | 0.0000083693 | INTERVAL MAPPING//SYSTEMS MAPPING//SELECTIVE GENOTYPING |
| 7 | 0.0000081371 | HALOTHANE GENE//PORK QUALITY//HALOTHANE GENOTYPE |
| 8 | 0.0000076981 | IBERIAN PIG//INTRAMUSCULAR FAT//CHATO MURCIANO |
| 9 | 0.0000070522 | M LONGISSIMUS LUMBORUM//MUSCLE FIBER CHARACTERISTICS//REPROD ANIM ANAT |
| 10 | 0.0000068610 | HN1//CANC 6401 E THOMAS RD//HORMONAL PROFILING |