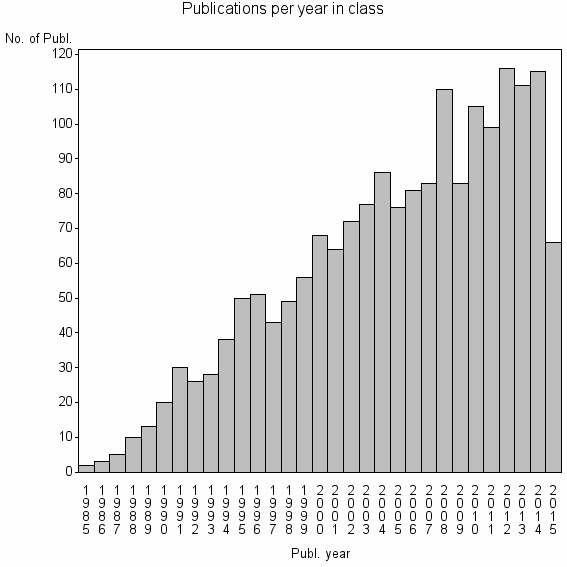
Class information for: |
Basic class information |
| ID | Publications | Average number of references |
Avg. shr. active ref. in WoS |
|---|---|---|---|
| 3769 | 1836 | 51.1 | 93% |
Classes in level above (level 2) |
| ID, lev. above |
Publications | Label for level above |
|---|---|---|
| 195 | 20594 | PRE MRNA SPLICING//RNA LOCALIZATION//SPLICEOSOME |
Terms with highest relevance score |
| Rank | Term | Type of term | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
|---|---|---|---|---|---|---|
| 1 | HUR | Author keyword | 130 | 63% | 7% | 131 |
| 2 | TRISTETRAPROLIN | Author keyword | 94 | 76% | 4% | 65 |
| 3 | AU RICH ELEMENT | Author keyword | 74 | 66% | 4% | 69 |
| 4 | AU RICH ELEMENTS | Author keyword | 40 | 61% | 2% | 43 |
| 5 | ELAV | Author keyword | 33 | 61% | 2% | 35 |
| 6 | MRNA STABILITY | Author keyword | 32 | 19% | 8% | 156 |
| 7 | AUF1 | Author keyword | 27 | 63% | 1% | 27 |
| 8 | PROGRAM BIOMOL | Address | 24 | 82% | 1% | 14 |
| 9 | TIAR | Author keyword | 21 | 71% | 1% | 17 |
| 10 | HU PROTEINS | Author keyword | 14 | 65% | 1% | 13 |
Web of Science journal categories |
Author Key Words |
| Rank | Web of Science journal category | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
LCSH search | Wikipedia search |
|---|---|---|---|---|---|---|---|
| 1 | HUR | 130 | 63% | 7% | 131 | Search HUR | Search HUR |
| 2 | TRISTETRAPROLIN | 94 | 76% | 4% | 65 | Search TRISTETRAPROLIN | Search TRISTETRAPROLIN |
| 3 | AU RICH ELEMENT | 74 | 66% | 4% | 69 | Search AU+RICH+ELEMENT | Search AU+RICH+ELEMENT |
| 4 | AU RICH ELEMENTS | 40 | 61% | 2% | 43 | Search AU+RICH+ELEMENTS | Search AU+RICH+ELEMENTS |
| 5 | ELAV | 33 | 61% | 2% | 35 | Search ELAV | Search ELAV |
| 6 | MRNA STABILITY | 32 | 19% | 8% | 156 | Search MRNA+STABILITY | Search MRNA+STABILITY |
| 7 | AUF1 | 27 | 63% | 1% | 27 | Search AUF1 | Search AUF1 |
| 8 | TIAR | 21 | 71% | 1% | 17 | Search TIAR | Search TIAR |
| 9 | HU PROTEINS | 14 | 65% | 1% | 13 | Search HU+PROTEINS | Search HU+PROTEINS |
| 10 | HNRNP D | 12 | 75% | 0% | 9 | Search HNRNP+D | Search HNRNP+D |
Key Words Plus |
| Rank | Web of Science journal category | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
|---|---|---|---|---|---|
| 1 | AU RICH ELEMENTS | 197 | 44% | 19% | 343 |
| 2 | ELAV LIKE PROTEIN | 154 | 92% | 3% | 60 |
| 3 | AU RICH ELEMENT | 150 | 54% | 10% | 192 |
| 4 | HUR | 97 | 51% | 8% | 138 |
| 5 | TRISTETRAPROLIN | 90 | 51% | 7% | 125 |
| 6 | 3 UNTRANSLATED REGION | 84 | 16% | 26% | 486 |
| 7 | BINDING PROTEIN HUR | 81 | 57% | 5% | 96 |
| 8 | ELAV PROTEIN | 69 | 83% | 2% | 39 |
| 9 | INCREASED CYCLOOXYGENASE 2 EXPRESSION | 68 | 79% | 2% | 44 |
| 10 | RICH ELEMENT | 43 | 54% | 3% | 55 |
Journals |
Reviews |
| Title | Publ. year | Cit. | Active references | % act. ref. to same field |
|---|---|---|---|---|
| Tristetraprolin (TTP): Interactions with mRNA and proteins, and current thoughts on mechanisms of action | 2013 | 46 | 197 | 74% |
| AU-rich elements and associated factors: are there unifying principles? | 2005 | 481 | 178 | 78% |
| Diverse molecular functions of Hu proteins | 2008 | 221 | 110 | 66% |
| AU-RICH ELEMENTS - CHARACTERIZATION AND IMPORTANCE IN MESSENGER-RNA DEGRADATION | 1995 | 1334 | 40 | 75% |
| Post-transcriptional control of gene expression by AUF1: Mechanisms, physiological targets, and regulation | 2013 | 22 | 95 | 83% |
| RNA regulons: coordination of post-transcriptional events | 2007 | 506 | 114 | 30% |
| Post-transcriptional control of cytokine production | 2008 | 183 | 100 | 56% |
| Functional Interplay between RNA-Binding Protein HuR and microRNAs | 2012 | 39 | 69 | 62% |
| The involvement of AU-rich element-binding proteins in p38 mitogen-activated protein kinase pathway-mediated mRNA stabilisation | 2004 | 225 | 88 | 81% |
| Post-transcriptional control during chronic inflammation and cancer: a focus on AU-rich elements | 2010 | 68 | 179 | 63% |
Address terms |
| Rank | Address term | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
|---|---|---|---|---|---|
| 1 | PROGRAM BIOMOL | 24 | 82% | 0.8% | 14 |
| 2 | BIOMOL PROGRAM | 9 | 83% | 0.3% | 5 |
| 3 | LCMB | 7 | 34% | 0.9% | 17 |
| 4 | LMBI | 6 | 80% | 0.2% | 4 |
| 5 | GENOM SCREENING | 4 | 56% | 0.3% | 5 |
| 6 | HELMHOLTZ JR GRP POSTTRANSCRIPT CONTROL GENE | 4 | 41% | 0.4% | 7 |
| 7 | BIOL MOL GENE | 3 | 37% | 0.4% | 7 |
| 8 | OFF CLIN | 3 | 11% | 1.3% | 24 |
| 9 | MINERVA CALCIUM BONE METAB | 3 | 23% | 0.5% | 10 |
| 10 | PROGRAM MOL PHARMACOL THER EUT | 2 | 19% | 0.6% | 11 |
Related classes at same level (level 1) |
| Rank | Relatedness score | Related classes |
|---|---|---|
| 1 | 0.0000224606 | MCPIP1//ZC3H12A//MCPIP |
| 2 | 0.0000171110 | DECAPPING//P BODIES//CCR4 NOT |
| 3 | 0.0000129957 | VIGILIN//HNRNP K//HETEROGENEOUS NUCLEAR RIBONUCLEOPROTEIN K |
| 4 | 0.0000101430 | HNRNP B1//HETEROGENEOUS NUCLEAR RIBONUCLEOPROTEINS HNRNPS//RNP CONCENSUS RNA BINDING DOMAIN |
| 5 | 0.0000092578 | CYTOPLASMIC POLYADENYLATION//PUMILIO//CPEB |
| 6 | 0.0000084562 | INTERACTION PROPENSITY//RNA BINDING RESIDUES//COMPUTAT INTELLIGENCE LEARNING DISCOVERY |
| 7 | 0.0000078297 | POLYA BINDING PROTEIN//POLY A BINDING PROTEIN//PABP |
| 8 | 0.0000066560 | MUSASHI1//MUSASHI//MUSASHI 2 |
| 9 | 0.0000059678 | IMP3//IGF2BP3//L523S |
| 10 | 0.0000053098 | UPF1//NMD//NONSENSE MEDIATED MRNA DECAY |