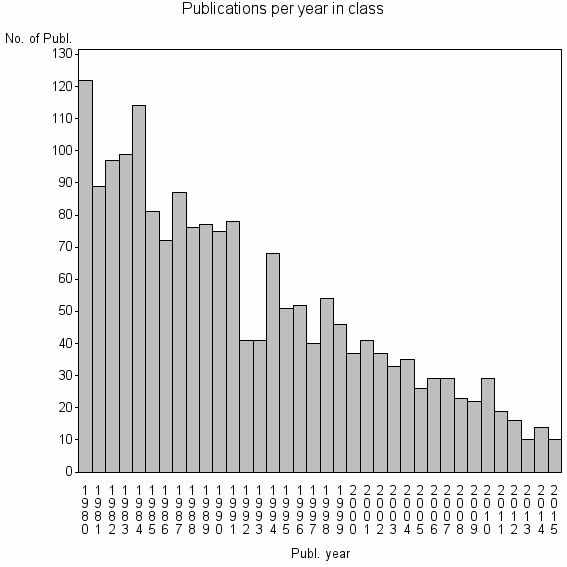
Class information for: |
Basic class information |
| ID | Publications | Average number of references |
Avg. shr. active ref. in WoS |
|---|---|---|---|
| 3612 | 1870 | 36.4 | 59% |
Classes in level above (level 2) |
| ID, lev. above |
Publications | Label for level above |
|---|---|---|
| 501 | 14604 | RNA POLYMERASE//SITE SPECIFIC RECOMBINATION//H NS |
Terms with highest relevance score |
| Rank | Term | Type of term | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
|---|---|---|---|---|---|---|
| 1 | DNA TRANSPOSITION | Author keyword | 65 | 77% | 2% | 44 |
| 2 | PHAGE MU | Author keyword | 30 | 68% | 1% | 27 |
| 3 | TN10 | Author keyword | 21 | 65% | 1% | 20 |
| 4 | MUA TRANSPOSASE | Author keyword | 20 | 100% | 0% | 9 |
| 5 | BACTERIOPHAGE MU | Author keyword | 19 | 64% | 1% | 18 |
| 6 | TRANSPOSASE | Author keyword | 17 | 24% | 3% | 62 |
| 7 | TRANSPOSOSOME | Author keyword | 17 | 100% | 0% | 8 |
| 8 | IS911 | Author keyword | 13 | 80% | 0% | 8 |
| 9 | MU DNA TRANSPOSITION | Author keyword | 11 | 100% | 0% | 6 |
| 10 | TRANSPOSITION | Author keyword | 10 | 12% | 4% | 83 |
Web of Science journal categories |
Author Key Words |
| Rank | Web of Science journal category | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
LCSH search | Wikipedia search |
|---|---|---|---|---|---|---|---|
| 1 | DNA TRANSPOSITION | 65 | 77% | 2% | 44 | Search DNA+TRANSPOSITION | Search DNA+TRANSPOSITION |
| 2 | PHAGE MU | 30 | 68% | 1% | 27 | Search PHAGE+MU | Search PHAGE+MU |
| 3 | TN10 | 21 | 65% | 1% | 20 | Search TN10 | Search TN10 |
| 4 | MUA TRANSPOSASE | 20 | 100% | 0% | 9 | Search MUA+TRANSPOSASE | Search MUA+TRANSPOSASE |
| 5 | BACTERIOPHAGE MU | 19 | 64% | 1% | 18 | Search BACTERIOPHAGE+MU | Search BACTERIOPHAGE+MU |
| 6 | TRANSPOSASE | 17 | 24% | 3% | 62 | Search TRANSPOSASE | Search TRANSPOSASE |
| 7 | TRANSPOSOSOME | 17 | 100% | 0% | 8 | Search TRANSPOSOSOME | Search TRANSPOSOSOME |
| 8 | IS911 | 13 | 80% | 0% | 8 | Search IS911 | Search IS911 |
| 9 | MU DNA TRANSPOSITION | 11 | 100% | 0% | 6 | Search MU+DNA+TRANSPOSITION | Search MU+DNA+TRANSPOSITION |
| 10 | TRANSPOSITION | 10 | 12% | 4% | 83 | Search TRANSPOSITION | Search TRANSPOSITION |
Key Words Plus |
| Rank | Web of Science journal category | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
|---|---|---|---|---|---|
| 1 | PHAGE MU | 132 | 77% | 5% | 90 |
| 2 | BACTERIOPHAGE MU | 109 | 50% | 8% | 156 |
| 3 | INVITRO TRANSPOSITION | 93 | 86% | 3% | 48 |
| 4 | TARGET IMMUNITY | 65 | 90% | 1% | 28 |
| 5 | TN7 ENCODED PROTEINS | 49 | 94% | 1% | 17 |
| 6 | STRAND TRANSFER REACTION | 47 | 81% | 2% | 29 |
| 7 | TN7 TRANSPOSITION | 33 | 58% | 2% | 38 |
| 8 | TN10 TRANSPOSITION | 33 | 56% | 2% | 40 |
| 9 | INSERTION SEQUENCE IS1 | 30 | 100% | 1% | 12 |
| 10 | IS50 | 30 | 84% | 1% | 16 |
Journals |
Reviews |
| Title | Publ. year | Cit. | Active references | % act. ref. to same field |
|---|---|---|---|---|
| Insertion sequences | 1998 | 854 | 337 | 52% |
| Regulation of transposition in bacteria | 2004 | 91 | 89 | 76% |
| Transposon Tn5 | 2008 | 38 | 61 | 92% |
| Tn7: Smarter than we thought | 2001 | 83 | 51 | 78% |
| Comparative architecture of transposase and integrase complexes | 2001 | 112 | 48 | 56% |
| Genetic variation: molecular mechanisms and impact on microbial evolution | 2000 | 131 | 10 | 80% |
| Integrating DNA: Transposases and retroviral integrases | 1999 | 158 | 174 | 51% |
| Structure/function insights into Tn5 transposition | 2004 | 52 | 37 | 76% |
| USES OF TRANSPOSONS WITH EMPHASIS ON TN10 | 1991 | 349 | 53 | 47% |
| TRANSPOSITIONAL RECOMBINATION - MECHANISTIC INSIGHTS FROM STUDIES OF MU AND OTHER ELEMENTS | 1992 | 297 | 127 | 62% |
Address terms |
| Rank | Address term | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
|---|---|---|---|---|---|
| 1 | MICROBIOL GENET MOL | 7 | 12% | 2.7% | 51 |
| 2 | UMR5100 | 4 | 24% | 0.7% | 14 |
| 3 | PROGRAM CELLULAR BIOTECHNOL | 2 | 15% | 0.6% | 12 |
| 4 | UPR 9007 | 2 | 29% | 0.3% | 5 |
| 5 | MICROBIOL ALIMENTAIRE ENVIRONM | 1 | 50% | 0.1% | 2 |
| 6 | MICROORGANISMS GENET ERS | 1 | 100% | 0.1% | 2 |
| 7 | UNIDAD ASOCIADA CIB | 1 | 50% | 0.1% | 2 |
| 8 | UNITE MIXTE RECH 5100 | 1 | 50% | 0.1% | 2 |
| 9 | PROGRAM CELLULAR MOL DEV BIOL | 1 | 20% | 0.3% | 5 |
| 10 | STRUCT BASES GENOME INTEGR GRP | 1 | 40% | 0.1% | 2 |
Related classes at same level (level 1) |
| Rank | Relatedness score | Related classes |
|---|---|---|
| 1 | 0.0000170668 | EPIDEMIOLOGICAL SIGNIFICANCE//SIMILARITY IDENTIFICATION//AIRBORNE ESCHERICHIA COLI |
| 2 | 0.0000130714 | SITE SPECIFIC RECOMBINATION//SERINE RECOMBINASE//PHI C31 INTEGRASE |
| 3 | 0.0000097663 | POLYLINKER//STREPTOMYCIN SENSITIVITY//INSERTIONAL INACTIVATION |
| 4 | 0.0000093751 | TERMINATION OF REPLICATION//REPLICATION TERMINATOR PROTEIN//FORK ARREST |
| 5 | 0.0000084520 | MARINER//SLEEPING BEAUTY//ARNOLD MABEL BECKMAN TRANSPOSON |
| 6 | 0.0000081492 | REPLICATION CONTROL//PLASMID R6K//PLASMID RK2 |
| 7 | 0.0000075604 | REX EXCLUSION//BACTERIOPHAGE LAMBDA LAMBDA//CI REXA REXB OPERON |
| 8 | 0.0000060731 | H NS//HISTONE LIKE PROTEIN//BACTERIAL CHROMATIN |
| 9 | 0.0000060295 | EMANUEL SYNDROME//PARTIAL TRISOMY 11Q//DUPLICATION 11Q |
| 10 | 0.0000055998 | SKIRBALL PROGRAM MOL PATHOGENESIS//BACTERIOPHAGE P2//RETRON |