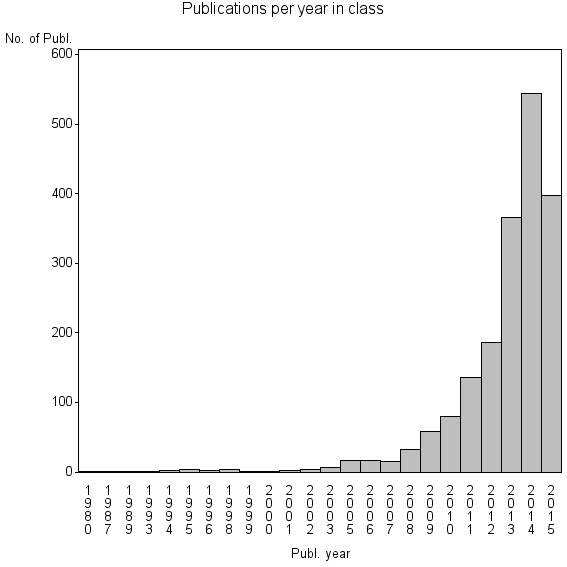
Class information for: |
Basic class information |
| ID | Publications | Average number of references |
Avg. shr. active ref. in WoS |
|---|---|---|---|
| 3570 | 1879 | 47.9 | 93% |
Classes in level above (level 2) |
Terms with highest relevance score |
| Rank | Term | Type of term | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
|---|---|---|---|---|---|---|
| 1 | CAS9 | Author keyword | 190 | 90% | 4% | 82 |
| 2 | GENOME EDITING | Author keyword | 167 | 70% | 7% | 139 |
| 3 | CRISPR | Author keyword | 149 | 63% | 8% | 150 |
| 4 | CRISPR CAS9 | Author keyword | 106 | 76% | 4% | 75 |
| 5 | CRISPR CAS | Author keyword | 96 | 86% | 3% | 49 |
| 6 | TALEN | Author keyword | 96 | 76% | 4% | 68 |
| 7 | CRRNA | Author keyword | 60 | 100% | 1% | 20 |
| 8 | ZINC FINGER NUCLEASE | Author keyword | 58 | 63% | 3% | 58 |
| 9 | TRANSCRIPTION ACTIVATOR LIKE EFFECTOR NUCLEASES | Author keyword | 53 | 95% | 1% | 18 |
| 10 | TALENS | Author keyword | 51 | 79% | 2% | 33 |
Web of Science journal categories |
Author Key Words |
| Rank | Web of Science journal category | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
LCSH search | Wikipedia search |
|---|---|---|---|---|---|---|---|
| 1 | CAS9 | 190 | 90% | 4% | 82 | Search CAS9 | Search CAS9 |
| 2 | GENOME EDITING | 167 | 70% | 7% | 139 | Search GENOME+EDITING | Search GENOME+EDITING |
| 3 | CRISPR | 149 | 63% | 8% | 150 | Search CRISPR | Search CRISPR |
| 4 | CRISPR CAS9 | 106 | 76% | 4% | 75 | Search CRISPR+CAS9 | Search CRISPR+CAS9 |
| 5 | CRISPR CAS | 96 | 86% | 3% | 49 | Search CRISPR+CAS | Search CRISPR+CAS |
| 6 | TALEN | 96 | 76% | 4% | 68 | Search TALEN | Search TALEN |
| 7 | CRRNA | 60 | 100% | 1% | 20 | Search CRRNA | Search CRRNA |
| 8 | ZINC FINGER NUCLEASE | 58 | 63% | 3% | 58 | Search ZINC+FINGER+NUCLEASE | Search ZINC+FINGER+NUCLEASE |
| 9 | TRANSCRIPTION ACTIVATOR LIKE EFFECTOR NUCLEASES | 53 | 95% | 1% | 18 | Search TRANSCRIPTION+ACTIVATOR+LIKE+EFFECTOR+NUCLEASES | Search TRANSCRIPTION+ACTIVATOR+LIKE+EFFECTOR+NUCLEASES |
| 10 | TALENS | 51 | 79% | 2% | 33 | Search TALENS | Search TALENS |
Key Words Plus |
| Rank | Web of Science journal category | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
|---|---|---|---|---|---|
| 1 | CAS SYSTEMS | 275 | 89% | 7% | 126 |
| 2 | ZINC FINGER NUCLEASES | 257 | 39% | 28% | 526 |
| 3 | CAS9 | 253 | 81% | 8% | 154 |
| 4 | SEED SEQUENCE | 230 | 100% | 3% | 57 |
| 5 | ONE STEP GENERATION | 151 | 79% | 5% | 96 |
| 6 | BACTERIAL IMMUNE SYSTEM | 148 | 90% | 3% | 64 |
| 7 | EFFECTOR NUCLEASES | 136 | 81% | 4% | 83 |
| 8 | RNA GUIDED ENDONUCLEASE | 128 | 94% | 2% | 46 |
| 9 | EMBRYO MICROINJECTION | 107 | 82% | 3% | 63 |
| 10 | CHIMERIC NUCLEASES | 101 | 84% | 3% | 56 |
Journals |
Reviews |
| Title | Publ. year | Cit. | Active references | % act. ref. to same field |
|---|---|---|---|---|
| Development and Applications of CRISPR-Cas9 for Genome Engineering | 2014 | 224 | 93 | 86% |
| CRIRISPR-Cas systems for editing, regulating and targeting genomes | 2014 | 169 | 91 | 96% |
| ZFN, TALEN, and CRISPR/Cas-based methods for genome engineering | 2013 | 471 | 115 | 78% |
| RNA-guided genetic silencing systems in bacteria and archaea | 2012 | 361 | 78 | 78% |
| A guide to genome engineering with programmable nucleases | 2014 | 86 | 171 | 88% |
| The new frontier of genome engineering with CRISPR-Cas9 | 2014 | 81 | 145 | 82% |
| A Mouse Geneticist's Practical Guide to CRISPR Applications | 2015 | 8 | 66 | 83% |
| CRISPR/Cas, the Immune System of Bacteria and Archaea | 2010 | 400 | 34 | 91% |
| Unravelling the structural and mechanistic basis of CRISPR-Cas systems | 2014 | 47 | 124 | 94% |
| CRISPR-Cas Systems: Prokaryotes Upgrade to Adaptive Immunity | 2014 | 45 | 103 | 94% |
Address terms |
| Rank | Address term | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
|---|---|---|---|---|---|
| 1 | CREAT INITIAT GENOME ENGN | 34 | 93% | 0.7% | 13 |
| 2 | GENOME ENGN | 21 | 34% | 2.7% | 51 |
| 3 | REGULAT INFECT BIOL | 20 | 100% | 0.5% | 9 |
| 4 | INTER PROGRAM BIOINFORMAT COMPUTAT BIO | 15 | 88% | 0.4% | 7 |
| 5 | PROKARYOT SMALL RNA BIOL | 9 | 83% | 0.3% | 5 |
| 6 | MOL PATHOL UNIT | 7 | 13% | 2.6% | 48 |
| 7 | COMPUTAT INTEGRAT BIOL | 6 | 11% | 2.5% | 47 |
| 8 | TSINGHUA FLY | 4 | 67% | 0.2% | 4 |
| 9 | VIROL CBF | 4 | 75% | 0.2% | 3 |
| 10 | OURCE MUTANT MICE | 3 | 35% | 0.4% | 8 |
Related classes at same level (level 1) |
| Rank | Relatedness score | Related classes |
|---|---|---|
| 1 | 0.0000163826 | ARTIFICIAL TRANSCRIPTION FACTOR//ARTIFICIAL TRANSCRIPTION FACTORS//RECOGNITION CODE |
| 2 | 0.0000091589 | SINGLE STRANDED OLIGONUCLEOTIDE//GENE REPAIR//SINGLE STRANDED OLIGONUCLEOTIDES |
| 3 | 0.0000084000 | PROTEIN SPLICING//INTEIN//HOMING ENDONUCLEASE |
| 4 | 0.0000082187 | HAPLOID EMBRYONIC STEM CELLS//CHONGQING PIG IND SCI//INFLAMMATORY LEIOMYOSARCOMA |
| 5 | 0.0000080587 | BLOOMINGTON DROSOPHILA STOCK//ENDS OUT TARGETING//ENDS OUT |
| 6 | 0.0000079663 | MARKER FREE TRANSGENIC PLANTS//MARKER FREE//MANNOSE SELECTION |
| 7 | 0.0000050609 | SYNTHETIC BIOLOGY//ACS SYNTHETIC BIOLOGY//CALIF QUANTITAT BIOL QB3 |
| 8 | 0.0000043054 | TILLING//SURVEYOR NUCLEASE//ECOTILLING |
| 9 | 0.0000041913 | MARINER//SLEEPING BEAUTY//ARNOLD MABEL BECKMAN TRANSPOSON |
| 10 | 0.0000041473 | LENTIVIRAL VECTOR//LENTIVIRAL VECTORS//EXPT HEMATOL |