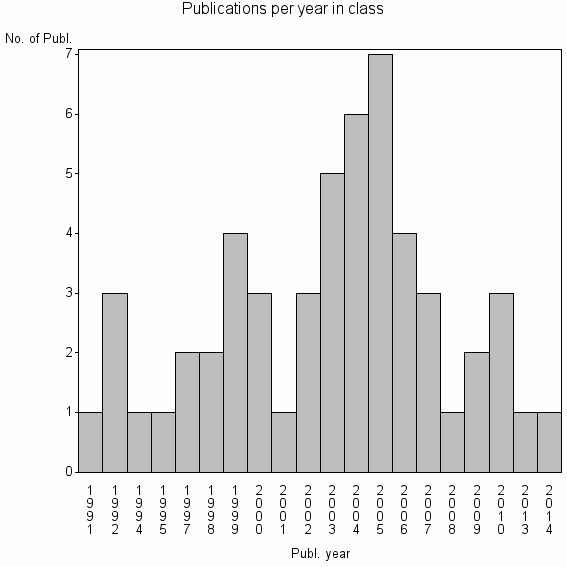
Class information for: |
Basic class information |
| ID | Publications | Average number of references |
Avg. shr. active ref. in WoS |
|---|---|---|---|
| 35063 | 54 | 18.9 | 69% |
Classes in level above (level 2) |
| ID, lev. above |
Publications | Label for level above |
|---|---|---|
| 1339 | 7890 | BIOTECHNIQUES//PCR-METHODS AND APPLICATIONS//DIFFERENTIAL DISPLAY |
Terms with highest relevance score |
| Rank | Term | Type of term | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
|---|---|---|---|---|---|---|
| 1 | TAISHO FUNCT GENOM | Address | 5 | 35% | 20% | 11 |
| 2 | MULTIPHOTON DETECTION | Author keyword | 2 | 50% | 6% | 3 |
| 3 | ADAPTER TAGGED COMPETITIVE PCR | Author keyword | 1 | 100% | 4% | 2 |
| 4 | ADAPTOR TAGGED COMPETITIVE POLYMERASE CHAIN REACTION | Author keyword | 1 | 100% | 4% | 2 |
| 5 | CARDINALITY OF ROW SPACE | Author keyword | 1 | 100% | 4% | 2 |
| 6 | ATAC PCR | Author keyword | 1 | 33% | 6% | 3 |
| 7 | ADAPTER TAGGED COMPETITIVE POLYMERASE CHAIN REACTION | Author keyword | 1 | 50% | 2% | 1 |
| 8 | ADAPTOR TAGGED COMPETITIVE PCR | Author keyword | 1 | 50% | 2% | 1 |
| 9 | CCT5 | Author keyword | 1 | 50% | 2% | 1 |
| 10 | MULTI PHOTON DETECTION | Author keyword | 1 | 50% | 2% | 1 |
Web of Science journal categories |
Author Key Words |
Key Words Plus |
| Rank | Web of Science journal category | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
|---|---|---|---|---|---|
| 1 | SUBNUCLEI | 0 | 20% | 2% | 1 |
| 2 | PRINCIPAL IDEALS | 0 | 17% | 2% | 1 |
| 3 | SYNAPSIN III | 0 | 14% | 2% | 1 |
| 4 | MULTIPHOTON DETECTION | 0 | 13% | 2% | 1 |
| 5 | NEURONATIN GENE | 0 | 11% | 2% | 1 |
| 6 | BOOLEAN MATRICES | 0 | 10% | 2% | 1 |
| 7 | PTPH1 | 0 | 10% | 2% | 1 |
| 8 | TAGGED COMPETITIVE PCR | 0 | 100% | 2% | 1 |
Journals |
Reviews |
| Title | Publ. year | Cit. | Active references |
% act. ref. to same field |
|---|---|---|---|---|
| Promise of multiphoton detection in discovery and diagnostic proteomics | 2007 | 1 | 24 | 17% |
| PrimerSelect: A transcriptome-wide oligonucleotide primer pair design program for kinetic RT-PCR-based transcript profiling | 2005 | 1 | 7 | 14% |
Address terms |
| Rank | Address term | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
|---|---|---|---|---|---|
| 1 | TAISHO FUNCT GENOM | 5 | 35% | 20% | 11 |
| 2 | PATENT EXAMINAT 3 | 1 | 50% | 1.9% | 1 |
| 3 | UMR 918 | 1 | 50% | 1.9% | 1 |
| 4 | THEORET LIFE SCI | 0 | 17% | 3.7% | 2 |
| 5 | TAISHO FUNCT GENOMICS | 0 | 17% | 1.9% | 1 |
| 6 | AXIS TEAM PROJECT | 0 | 100% | 1.9% | 1 |
| 7 | NARA SCI TECHNOLCORE EVOLUT SCI TEC | 0 | 100% | 1.9% | 1 |
Related classes at same level (level 1) |
| Rank | Relatedness score | Related classes |
|---|---|---|
| 1 | 0.0000122063 | MISSING VALUE IMPUTATION//MISSING VALUE ESTIMATION//COMPUTAT MATH BIOL |
| 2 | 0.0000112077 | SERIAL ANALYSIS OF GENE EXPRESSION//SAGE//GLGI |
| 3 | 0.0000107513 | EXPONENTIAL INTEGRATORS//MATRIX PTH ROOT//MATRIX EXPONENTIAL |
| 4 | 0.0000099359 | PIP92//LYMPHOCYTE PROTEINS//PROTEINPAEDIA |
| 5 | 0.0000090431 | BICLUSTERING//GENE EXPRESSION DATA//GENE EXPRESSION DATA ANALYSIS |
| 6 | 0.0000088245 | MKIAA//SMART CDNA LIBRARY//CDNA SEQUENCING |
| 7 | 0.0000082504 | MAX ALGEBRA//MATEMAT BEPPO LEVI//MAX PLUS ALGEBRA |
| 8 | 0.0000079451 | NEURITIN//ARG31//ACTIVITY REGULATED CYTOSKELETON ASSOCIATED PROTEIN |
| 9 | 0.0000077868 | MICROARRAY IMAGE PROCESSING//GENE FILTERING//MAQC |
| 10 | 0.0000076396 | CHD1L//CODON 249//SUPPRESSION OF TUMORIGENICITY |