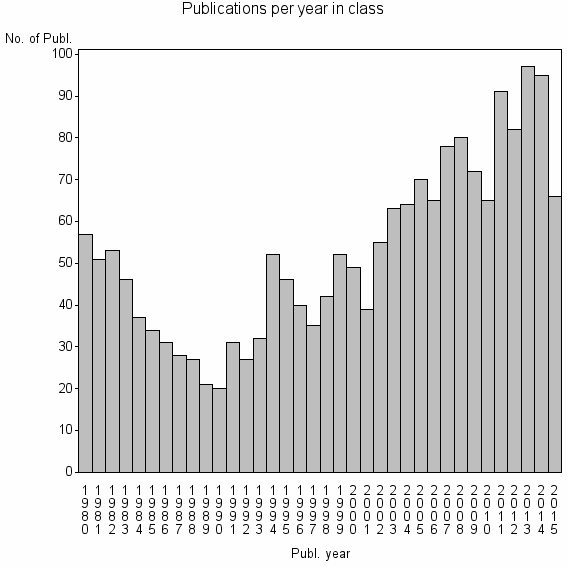
Class information for: |
Basic class information |
| ID | Publications | Average number of references |
Avg. shr. active ref. in WoS |
|---|---|---|---|
| 3487 | 1893 | 42.2 | 70% |
Classes in level above (level 2) |
Terms with highest relevance score |
| Rank | Term | Type of term | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
|---|---|---|---|---|---|---|
| 1 | QUEUINE | Author keyword | 75 | 91% | 2% | 31 |
| 2 | TRNA MODIFICATION | Author keyword | 35 | 55% | 2% | 44 |
| 3 | RNA MODIFICATION | Author keyword | 32 | 42% | 3% | 59 |
| 4 | PSEUDOURIDINE SYNTHASE | Author keyword | 31 | 73% | 1% | 24 |
| 5 | QUEUOSINE | Author keyword | 25 | 70% | 1% | 21 |
| 6 | PSEUDOURIDINE | Author keyword | 21 | 38% | 2% | 44 |
| 7 | TRNA | Author keyword | 18 | 13% | 7% | 130 |
| 8 | TRANSFER RNA MODIFICATION | Author keyword | 18 | 83% | 1% | 10 |
| 9 | MNME | Author keyword | 15 | 77% | 1% | 10 |
| 10 | TRM4 | Author keyword | 14 | 100% | 0% | 7 |
Web of Science journal categories |
Author Key Words |
| Rank | Web of Science journal category | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
LCSH search | Wikipedia search |
|---|---|---|---|---|---|---|---|
| 1 | QUEUINE | 75 | 91% | 2% | 31 | Search QUEUINE | Search QUEUINE |
| 2 | TRNA MODIFICATION | 35 | 55% | 2% | 44 | Search TRNA+MODIFICATION | Search TRNA+MODIFICATION |
| 3 | RNA MODIFICATION | 32 | 42% | 3% | 59 | Search RNA+MODIFICATION | Search RNA+MODIFICATION |
| 4 | PSEUDOURIDINE SYNTHASE | 31 | 73% | 1% | 24 | Search PSEUDOURIDINE+SYNTHASE | Search PSEUDOURIDINE+SYNTHASE |
| 5 | QUEUOSINE | 25 | 70% | 1% | 21 | Search QUEUOSINE | Search QUEUOSINE |
| 6 | PSEUDOURIDINE | 21 | 38% | 2% | 44 | Search PSEUDOURIDINE | Search PSEUDOURIDINE |
| 7 | TRNA | 18 | 13% | 7% | 130 | Search TRNA | Search TRNA |
| 8 | TRANSFER RNA MODIFICATION | 18 | 83% | 1% | 10 | Search TRANSFER+RNA+MODIFICATION | Search TRANSFER+RNA+MODIFICATION |
| 9 | MNME | 15 | 77% | 1% | 10 | Search MNME | Search MNME |
| 10 | TRM4 | 14 | 100% | 0% | 7 | Search TRM4 | Search TRM4 |
Key Words Plus |
| Rank | Web of Science journal category | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
|---|---|---|---|---|---|
| 1 | QUEUINE | 110 | 95% | 2% | 37 |
| 2 | MODIFIED NUCLEOSIDES | 83 | 46% | 7% | 135 |
| 3 | QUEUOSINE | 80 | 80% | 3% | 49 |
| 4 | MODIFIED NUCLEOSIDE | 54 | 83% | 2% | 30 |
| 5 | ANTICODON | 51 | 33% | 7% | 130 |
| 6 | M5C967 METHYLTRANSFERASE | 49 | 94% | 1% | 17 |
| 7 | AMINO ACID SPECIFICITIES | 47 | 84% | 1% | 26 |
| 8 | RNA GUANINE TRANSGLYCOSYLASE | 47 | 69% | 2% | 40 |
| 9 | GUANINE TRANSGLYCOSYLASE | 44 | 77% | 2% | 30 |
| 10 | TRANSFER RNA TRANSGLYCOSYLASE | 37 | 100% | 1% | 14 |
Journals |
Reviews |
| Title | Publ. year | Cit. | Active references | % act. ref. to same field |
|---|---|---|---|---|
| tRNA's wobble decoding of the genome: 40 years of modification | 2007 | 203 | 80 | 53% |
| Biosynthesis and Function of Posttranscriptional Modifications of Transfer RNAs | 2012 | 62 | 129 | 71% |
| Posttranscriptional RNA Modifications: Playing Metabolic Games in a Cell's Chemical Legoland | 2014 | 12 | 50 | 68% |
| RNA nucleotide methylation | 2011 | 67 | 106 | 48% |
| Role of tRNA modifications in human diseases | 2014 | 21 | 69 | 38% |
| tRNA's modifications bring order to gene expression | 2008 | 92 | 45 | 56% |
| Human Mitochondrial tRNAs: Biogenesis, Function, Structural Aspects, and Diseases | 2011 | 73 | 180 | 39% |
| Decoding the genome: a modified view | 2004 | 166 | 131 | 56% |
| THE MODIFIED NUCLEOSIDES OF RNA - SUMMARY | 1994 | 344 | 47 | 45% |
| Pseudouridine in RNA: What, where, how, and why | 2000 | 160 | 72 | 57% |
Address terms |
| Rank | Address term | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
|---|---|---|---|---|---|
| 1 | IFR115UMR8621 | 6 | 80% | 0.2% | 4 |
| 2 | RNA MODIFICAT MITOCHONDRIAL DIS | 6 | 100% | 0.2% | 4 |
| 3 | RIEVE L S MASS SPE OMETRY | 5 | 26% | 0.8% | 16 |
| 4 | RECH MICROBIOL JEAN MARIE WIAME | 4 | 67% | 0.2% | 4 |
| 5 | PHARM INTER PROGRAM MED CHEM | 3 | 100% | 0.2% | 3 |
| 6 | MATURAT ARN ENZYMOL MOL | 2 | 21% | 0.5% | 10 |
| 7 | BIOINFORMAT PROT ENGN | 2 | 12% | 1.0% | 19 |
| 8 | CNRSUMR 7567 | 2 | 44% | 0.2% | 4 |
| 9 | CR UK CAMBRIDGE | 2 | 44% | 0.2% | 4 |
| 10 | BIOL MOL CELL BIOL ANIM BIOL | 2 | 67% | 0.1% | 2 |
Related classes at same level (level 1) |
| Rank | Relatedness score | Related classes |
|---|---|---|
| 1 | 0.0000156887 | ALKB//METTL3//ALKBH3 |
| 2 | 0.0000145203 | TRNA NUCLEOTIDYLTRANSFERASE//CCA ADDING ENZYME//TRNAHIS GUANYLYLTRANSFERASE |
| 3 | 0.0000127789 | FRAMESHIFTING//RIBOSOMAL FRAMESHIFTING//RNA PSEUDOKNOT |
| 4 | 0.0000117557 | TRNA SPLICING//POLYNUCLEOTIDE KINASE//SPLICING ENDONUCLEASE |
| 5 | 0.0000104286 | SNORNA//SMALL NUCLEOLAR RNAS//PRE RRNA PROCESSING |
| 6 | 0.0000093662 | PUPYLATION//PROKARYOTIC UBIQUITIN LIKE PROTEIN//URM1 |
| 7 | 0.0000082073 | URINARY NUCLEOSIDES//URINARY MODIFIED NUCLEOSIDES//MODIFIED NUCLEOSIDES |
| 8 | 0.0000078415 | DAP3//ICT1//MITOCHONDRIAL RIBOSOMAL PROTEINS |
| 9 | 0.0000076305 | OBG//DRG2//BACTERIAL GTPASE |
| 10 | 0.0000075126 | DISACCHARIDE NUCLEOSIDES//DISACCHARIDE NUCLEOSIDE//2 O MODIFIED NUCLEOSIDES |