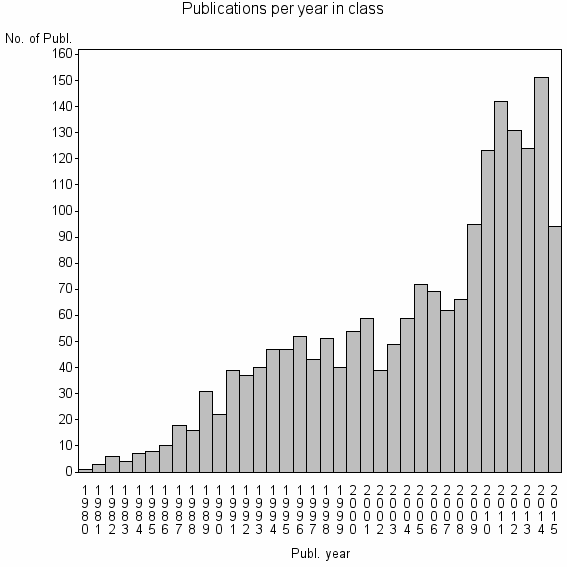
Class information for: |
Basic class information |
| ID | Publications | Average number of references |
Avg. shr. active ref. in WoS |
|---|---|---|---|
| 3431 | 1911 | 48.9 | 76% |
Classes in level above (level 2) |
| ID, lev. above |
Publications | Label for level above |
|---|---|---|
| 42 | 31225 | PROTEINS-STRUCTURE FUNCTION AND BIOINFORMATICS//PROTEIN STRUCTURE PREDICTION//PROTEIN FOLDING |
Terms with highest relevance score |
| Rank | Term | Type of term | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
|---|---|---|---|---|---|---|
| 1 | ENVIRONM IND CHEM | Address | 31 | 92% | 1% | 12 |
| 2 | ONE STEP PERTURBATION | Author keyword | 23 | 100% | 1% | 10 |
| 3 | SAMPL4 | Author keyword | 21 | 90% | 0% | 9 |
| 4 | THERMODYNAMIC INTEGRATION | Author keyword | 20 | 36% | 2% | 45 |
| 5 | FREE ENERGY PERTURBATION | Author keyword | 20 | 37% | 2% | 42 |
| 6 | LINEAR INTERACTION ENERGY | Author keyword | 17 | 63% | 1% | 17 |
| 7 | FREE ENERGY CALCULATIONS | Author keyword | 17 | 33% | 2% | 42 |
| 8 | FREE ENERGY CALCULATION | Author keyword | 11 | 29% | 2% | 32 |
| 9 | FREE ENERGY PERTURBATION METHOD | Author keyword | 11 | 100% | 0% | 6 |
| 10 | HYDRATION FREE ENERGIES | Author keyword | 8 | 37% | 1% | 18 |
Web of Science journal categories |
Author Key Words |
Key Words Plus |
| Rank | Web of Science journal category | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
|---|---|---|---|---|---|
| 1 | THERMODYNAMIC INTEGRATION | 74 | 47% | 6% | 115 |
| 2 | PERTURBATION CALCULATIONS | 57 | 54% | 4% | 74 |
| 3 | BINDING FREE ENERGIES | 50 | 36% | 6% | 113 |
| 4 | FREE ENERGY CALCULATIONS | 49 | 17% | 14% | 267 |
| 5 | PROTEIN LIGAND BINDING | 32 | 42% | 3% | 59 |
| 6 | HYDRATION FREE ENERGIES | 30 | 28% | 5% | 93 |
| 7 | RELATIVE FREE ENERGY | 27 | 92% | 1% | 11 |
| 8 | SINGLE SIMULATION | 26 | 87% | 1% | 13 |
| 9 | SPACE OVERLAP MEASURES | 24 | 91% | 1% | 10 |
| 10 | ABSOLUTE FREE ENERGIES | 22 | 63% | 1% | 22 |
Journals |
Reviews |
| Title | Publ. year | Cit. | Active references | % act. ref. to same field |
|---|---|---|---|---|
| CHARMM: The Biomolecular Simulation Program | 2009 | 2048 | 616 | 13% |
| Direct methods for computing single-molecule entropies from molecular simulations | 2015 | 2 | 90 | 63% |
| A 2ND GENERATION FORCE-FIELD FOR THE SIMULATION OF PROTEINS, NUCLEIC-ACIDS, AND ORGANIC-MOLECULES | 1995 | 7169 | 58 | 24% |
| Calculation of protein-ligand binding affinities | 2007 | 388 | 126 | 35% |
| Statistical mechanics and molecular dynamics in evaluating thermodynamic properties of biomolecular recognition | 2012 | 45 | 108 | 63% |
| Calculating structures and free energies of complex molecules: Combining molecular mechanics and continuum models | 2000 | 1536 | 44 | 34% |
| Computations of Standard Binding Free Energies with Molecular Dynamics Simulations | 2009 | 181 | 116 | 63% |
| Molecular Recognition and Ligand Association | 2013 | 33 | 131 | 62% |
| Molecular dynamics simulations and drug discovery | 2011 | 105 | 56 | 30% |
| All-atom empirical potential for molecular modeling and dynamics studies of proteins | 1998 | 6458 | 78 | 10% |
Address terms |
| Rank | Address term | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
|---|---|---|---|---|---|
| 1 | ENVIRONM IND CHEM | 31 | 92% | 0.6% | 12 |
| 2 | MOL MODELING SIMULAT | 7 | 29% | 1.0% | 20 |
| 3 | CAMBRIDGE MOL THER EUT PROGRAMME | 7 | 67% | 0.3% | 6 |
| 4 | CHEM PHYS NIH | 2 | 67% | 0.1% | 2 |
| 5 | CHEM PL BIOSCI COMPUTAT SCI | 2 | 67% | 0.1% | 2 |
| 6 | COMPUTAT CHEM LEAD IDENTIFICAT OPTIMIZAT SUPPOR | 2 | 67% | 0.1% | 2 |
| 7 | ENVIRONM FINE CHEM | 2 | 67% | 0.1% | 2 |
| 8 | ENVIRONM LIVING CHEM | 2 | 67% | 0.1% | 2 |
| 9 | NANO SOFT MAT FUNSOM | 2 | 67% | 0.1% | 2 |
| 10 | PHYS CHEM INSIGHTS GRP | 2 | 43% | 0.2% | 3 |
Related classes at same level (level 1) |
| Rank | Relatedness score | Related classes |
|---|---|---|
| 1 | 0.0000195421 | POISSON BOLTZMANN EQUATION//POISSON BOLTZMANN//BIOMOLECULAR ELECTROSTATICS |
| 2 | 0.0000150089 | EWALD METHODS//EWALD SUM//CUTOFF METHOD |
| 3 | 0.0000149348 | PROTEIN HYDRATION//INTERNAL SOLVENT//SOLVENT MODELING |
| 4 | 0.0000142003 | MARKOV STATE MODELS//TRANSITION PATH SAMPLING//TRANSITION PATH THEORY |
| 5 | 0.0000132360 | POLARIZABLE FORCE FIELD//HYBRIDIZATION DISPLACEMENT CHARGE//EXCHANGE REPULSION |
| 6 | 0.0000128225 | JOURNAL OF CHEMICAL INFORMATION AND MODELING//VIRTUAL SCREENING//JOURNAL OF COMPUTER-AIDED MOLECULAR DESIGN |
| 7 | 0.0000128065 | DIPARTIMENTO SCI BIOL AMBIENTALI//CAVITY CREATION//WORK OF CAVITY CREATION |
| 8 | 0.0000124089 | BAKER CHEM CHEM BIOL//REPLICA EXCHANGE METHOD//GENERALIZED ENSEMBLE ALGORITHM |
| 9 | 0.0000112581 | EXTREMELY LOCALIZED MOLECULAR ORBITALS//ASEP MD//LINK ATOM |
| 10 | 0.0000110637 | BIOPHYS MODELING SIMULAT//ALL ATOM RECONSTRUCTION//BIOL MODELING SIMULAT |