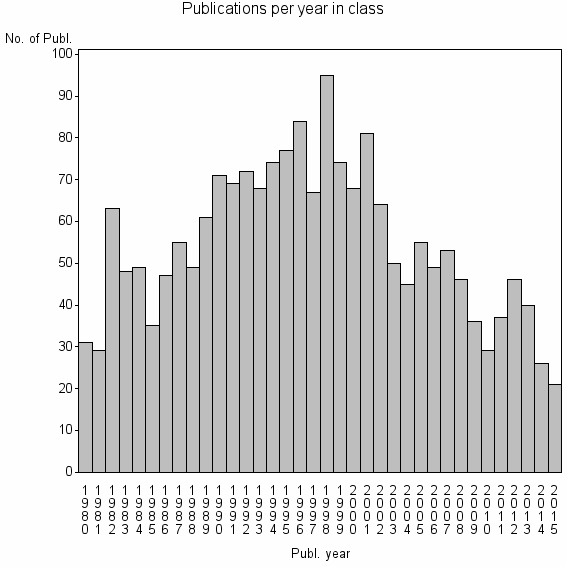
Class information for: |
Basic class information |
| ID | Publications | Average number of references |
Avg. shr. active ref. in WoS |
|---|---|---|---|
| 3191 | 1964 | 39.0 | 70% |
Classes in level above (level 2) |
| ID, lev. above |
Publications | Label for level above |
|---|---|---|
| 2019 | 4960 | PHOSPHOTRANSFERASE SYSTEM//LACTOSE PERMEASE//BIOL BAKTERIENPHYSIOL |
Terms with highest relevance score |
| Rank | Term | Type of term | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
|---|---|---|---|---|---|---|
| 1 | PHOSPHOTRANSFERASE SYSTEM | Author keyword | 127 | 68% | 6% | 112 |
| 2 | URA 1925 | Address | 27 | 74% | 1% | 20 |
| 3 | INDUCER EXCLUSION | Author keyword | 27 | 83% | 1% | 15 |
| 4 | HPR | Author keyword | 26 | 51% | 2% | 36 |
| 5 | ENZYME I | Author keyword | 23 | 79% | 1% | 15 |
| 6 | MICROBIAL BIOCHEM GENET UNIT | Address | 23 | 76% | 1% | 16 |
| 7 | PHOSPHOENOLPYRUVATE SUGAR PHOSPHOTRANSFERASE SYSTEM | Author keyword | 21 | 85% | 1% | 11 |
| 8 | MIKROBIOL BIOCHEM GENET | Address | 18 | 40% | 2% | 35 |
| 9 | SUGAR PHOSPHOTRANSFERASE SYSTEM | Author keyword | 13 | 80% | 0% | 8 |
| 10 | ENZYME II | Author keyword | 12 | 75% | 0% | 9 |
Web of Science journal categories |
Author Key Words |
Key Words Plus |
| Rank | Web of Science journal category | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
|---|---|---|---|---|---|
| 1 | PHOSPHOCARRIER PROTEIN | 165 | 81% | 5% | 99 |
| 2 | PHOSPHOENOLPYRUVATE | 163 | 44% | 14% | 278 |
| 3 | SUGAR PHOSPHOTRANSFERASE SYSTEM | 151 | 55% | 10% | 187 |
| 4 | HPR | 145 | 70% | 6% | 121 |
| 5 | PHOSPHOTRANSFERASE SYSTEM | 123 | 33% | 16% | 309 |
| 6 | BACTERIAL PHOSPHOENOLPYRUVATE | 104 | 75% | 4% | 75 |
| 7 | DEPENDENT PHOSPHOTRANSFERASE SYSTEM | 98 | 73% | 4% | 75 |
| 8 | ENZYME I | 83 | 63% | 4% | 84 |
| 9 | BETA GLUCOSIDE UTILIZATION | 79 | 87% | 2% | 39 |
| 10 | SIGNAL TRANSDUCING PROTEIN | 77 | 89% | 2% | 34 |
Journals |
Reviews |
| Title | Publ. year | Cit. | Active references | % act. ref. to same field |
|---|---|---|---|---|
| Structural insight into the PTS sugar transporter EIIC | 2015 | 3 | 68 | 87% |
| How phosphotransferase system-related protein phosphorylation regulates carbohydrate metabolism in bacteria | 2006 | 447 | 882 | 63% |
| The mechanisms of carbon catabolite repression in bacteria | 2008 | 175 | 50 | 66% |
| PHOSPHOENOLPYRUVATE - CARBOHYDRATE PHOSPHOTRANSFERASE SYSTEMS OF BACTERIA | 1993 | 1054 | 430 | 86% |
| The Bacterial Phosphoenolpyruvate:Carbohydrate Phosphotransferase System: Regulation by Protein Phosphorylation and Phosphorylation-Dependent Protein-Protein Interactions | 2014 | 19 | 201 | 78% |
| Carbon Catabolite Control of the Metabolic Network in Bacillus subtilis | 2009 | 94 | 145 | 64% |
| Carbon catabolite repression in bacteria: choice of the carbon source and autoregulatory limitation of sugar utilization | 2002 | 195 | 47 | 79% |
| CcpA-dependent carbon catabolite repression in bacteria | 2003 | 123 | 41 | 80% |
| Transcription regulators controlled by interaction with enzyme IIB components of the phosphoenolpyruvate:sugar phosphotransferase system | 2013 | 9 | 81 | 89% |
| Regulation of carbon catabolism in Bacillus species | 2000 | 195 | 182 | 75% |
Address terms |
| Rank | Address term | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
|---|---|---|---|---|---|
| 1 | URA 1925 | 27 | 74% | 1.0% | 20 |
| 2 | MICROBIAL BIOCHEM GENET UNIT | 23 | 76% | 0.8% | 16 |
| 3 | MIKROBIOL BIOCHEM GENET | 18 | 40% | 1.8% | 35 |
| 4 | GENET MICROORGANISMES | 12 | 36% | 1.4% | 27 |
| 5 | MICROBIOL ALIMENTAT SERV SANTE HUMAINE MICALIS | 6 | 71% | 0.3% | 5 |
| 6 | MC UM PRATT | 5 | 29% | 0.7% | 14 |
| 7 | EXP S GENET MICROBIENNE FRE3630 | 4 | 75% | 0.2% | 3 |
| 8 | UMR2585 | 4 | 44% | 0.4% | 7 |
| 9 | HGEBIET SYST BIOTECHNOL | 3 | 57% | 0.2% | 4 |
| 10 | AG PHYSIOL MICROORGANISMS | 3 | 100% | 0.2% | 3 |
Related classes at same level (level 1) |
| Rank | Relatedness score | Related classes |
|---|---|---|
| 1 | 0.0000126266 | INOSITOL DEHYDROGENASE//INOSITOL CATABOLISM//MYO INOSITOL DEHYDROGENASE |
| 2 | 0.0000108967 | CYCLIC AMP RECEPTOR PROTEIN//CATABOLITE ACTIVATOR PROTEIN CAP//CAMP RECEPTOR PROTEIN |
| 3 | 0.0000071993 | LACTOSE PERMEASE//PHYSIOL MICROBIOL//MELIBIOSE CARRIER |
| 4 | 0.0000071164 | CITRATE PERMEASE//DIACETYL//ALPHA ACETOLACTATE |
| 5 | 0.0000062700 | ABRB//SPO0A//TRANSITION STATE REGULATOR |
| 6 | 0.0000056593 | CARAB//PYRIMIDINE REGULATION//PYRR |
| 7 | 0.0000055553 | PHAGE TYPING BACTERIAL GENET//VTB ENZYMTECHNOL//DEGSU |
| 8 | 0.0000051986 | TERMINATION OF REPLICATION//REPLICATION TERMINATOR PROTEIN//FORK ARREST |
| 9 | 0.0000049387 | GLUCOSAMINE 6 PHOSPHATE SYNTHASE//GLUCOSAMINE 6P SYNTHASE//GLUCOSAMINE 6 PHOSPHATE DEAMINASE |
| 10 | 0.0000049255 | TRANSCRIPTION FACTOR TNRA//GLNR//BECKMAN COULTER |