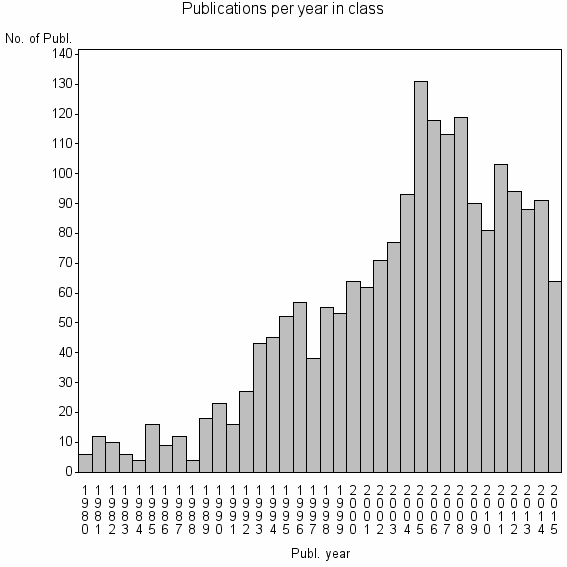
Class information for: |
Basic class information |
| ID | Publications | Average number of references |
Avg. shr. active ref. in WoS |
|---|---|---|---|
| 3189 | 1965 | 39.1 | 72% |
Classes in level above (level 2) |
| ID, lev. above |
Publications | Label for level above |
|---|---|---|
| 855 | 10973 | SYSTEMATIC BIOLOGY//SEKT ORNITHOL//FOSSIL BIRDS |
Terms with highest relevance score |
| Rank | Term | Type of term | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
|---|---|---|---|---|---|---|
| 1 | CODON MODEL | Author keyword | 37 | 81% | 1% | 22 |
| 2 | HETEROTACHY | Author keyword | 35 | 76% | 1% | 25 |
| 3 | PHYLOGENETIC INVARIANTS | Author keyword | 32 | 85% | 1% | 17 |
| 4 | SYSTEMATIC BIOLOGY | Journal | 30 | 15% | 9% | 186 |
| 5 | HADAMARD CONJUGATION | Author keyword | 24 | 91% | 1% | 10 |
| 6 | NEIGHBOR JOINING | Author keyword | 24 | 42% | 2% | 43 |
| 7 | PHYLOGENETIC INFERENCE | Author keyword | 22 | 33% | 3% | 55 |
| 8 | COVARION | Author keyword | 20 | 62% | 1% | 21 |
| 9 | SUBSTITUTION MODELS | Author keyword | 19 | 71% | 1% | 15 |
| 10 | MINIMUM EVOLUTION | Author keyword | 16 | 58% | 1% | 18 |
Web of Science journal categories |
Author Key Words |
| Rank | Web of Science journal category | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
LCSH search | Wikipedia search |
|---|---|---|---|---|---|---|---|
| 1 | CODON MODEL | 37 | 81% | 1% | 22 | Search CODON+MODEL | Search CODON+MODEL |
| 2 | HETEROTACHY | 35 | 76% | 1% | 25 | Search HETEROTACHY | Search HETEROTACHY |
| 3 | PHYLOGENETIC INVARIANTS | 32 | 85% | 1% | 17 | Search PHYLOGENETIC+INVARIANTS | Search PHYLOGENETIC+INVARIANTS |
| 4 | HADAMARD CONJUGATION | 24 | 91% | 1% | 10 | Search HADAMARD+CONJUGATION | Search HADAMARD+CONJUGATION |
| 5 | NEIGHBOR JOINING | 24 | 42% | 2% | 43 | Search NEIGHBOR+JOINING | Search NEIGHBOR+JOINING |
| 6 | PHYLOGENETIC INFERENCE | 22 | 33% | 3% | 55 | Search PHYLOGENETIC+INFERENCE | Search PHYLOGENETIC+INFERENCE |
| 7 | COVARION | 20 | 62% | 1% | 21 | Search COVARION | Search COVARION |
| 8 | SUBSTITUTION MODELS | 19 | 71% | 1% | 15 | Search SUBSTITUTION+MODELS | Search SUBSTITUTION+MODELS |
| 9 | MINIMUM EVOLUTION | 16 | 58% | 1% | 18 | Search MINIMUM+EVOLUTION | Search MINIMUM+EVOLUTION |
| 10 | CODON MODELS | 15 | 59% | 1% | 17 | Search CODON+MODELS | Search CODON+MODELS |
Key Words Plus |
| Rank | Web of Science journal category | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
|---|---|---|---|---|---|
| 1 | EVOLUTIONARY TREES | 73 | 26% | 12% | 242 |
| 2 | NUCLEOTIDE SUBSTITUTION | 52 | 17% | 14% | 270 |
| 3 | LINEAR INVARIANTS | 38 | 86% | 1% | 19 |
| 4 | AMINO ACID SITES | 32 | 16% | 9% | 178 |
| 5 | GENERAL MARKOV MODEL | 30 | 84% | 1% | 16 |
| 6 | CODON SUBSTITUTION MODELS | 28 | 28% | 4% | 85 |
| 7 | PARSIMONY | 28 | 15% | 9% | 177 |
| 8 | MAXIMUM LIKELIHOOD APPROACH | 26 | 29% | 4% | 76 |
| 9 | PHYLOGENETIC INVARIANTS | 26 | 87% | 1% | 13 |
| 10 | SEQUENCE EVOLUTION | 22 | 20% | 5% | 101 |
Journals |
| Rank | Web of Science journal category | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
|---|---|---|---|---|---|
| 1 | SYSTEMATIC BIOLOGY | 30 | 15% | 9% | 186 |
Reviews |
| Title | Publ. year | Cit. | Active references | % act. ref. to same field |
|---|---|---|---|---|
| Model selection and model averaging in phylogenetics: Advantages of akaike information criterion and Bayesian approaches over likelihood ratio tests | 2004 | 1753 | 97 | 43% |
| Phylogenomics: the beginning of incongruence? | 2006 | 231 | 23 | 48% |
| Molecular phylogenetics: principles and practice | 2012 | 78 | 110 | 39% |
| Investigating Protein-Coding Sequence Evolution with Probabilistic Codon Substitution Models | 2009 | 63 | 151 | 65% |
| Among-site rate variation and its impact on phylogenetic analyses | 1996 | 744 | 39 | 85% |
| Evolution - Bayesian inference of phylogeny and its impact on evolutionary biology | 2001 | 1377 | 28 | 36% |
| Neighbor-joining revealed | 2006 | 67 | 17 | 71% |
| Model selection in phylogenetics | 2005 | 183 | 63 | 68% |
| The posterior and the prior in Bayesian phylogenetics | 2006 | 69 | 65 | 66% |
| Models of coding sequence evolution | 2009 | 23 | 91 | 82% |
Address terms |
| Rank | Address term | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
|---|---|---|---|---|---|
| 1 | INFORMAT FONDAMENTALE PLICAT | 5 | 54% | 0.4% | 7 |
| 2 | URM 5506 | 3 | 100% | 0.2% | 3 |
| 3 | MARINE GENET BIOTECHNOL | 3 | 60% | 0.2% | 3 |
| 4 | ROBERT CEDERGREN | 3 | 19% | 0.7% | 13 |
| 5 | EHLTH INNOVAT PLATFORM | 2 | 67% | 0.1% | 2 |
| 6 | FUS DATA ANAL DEV TEAM | 2 | 67% | 0.1% | 2 |
| 7 | INFORMAT FONDAMENTALE PL | 2 | 67% | 0.1% | 2 |
| 8 | MUELLER 311 | 2 | 67% | 0.1% | 2 |
| 9 | UNIV VIENNA | 2 | 18% | 0.5% | 9 |
| 10 | DEV GRP NEXT GENERAT INTEGRATED LIVING MATT | 2 | 43% | 0.2% | 3 |
Related classes at same level (level 1) |
| Rank | Relatedness score | Related classes |
|---|---|---|
| 1 | 0.0000227804 | MOLECULAR CLOCK//RELAXED CLOCK//PHYLODYNAMICS |
| 2 | 0.0000214898 | PHYLOGENETIC NETWORK//PHYLOGENETIC NETWORKS//TIGHT SPAN |
| 3 | 0.0000100702 | MULTIPLE SEQUENCE ALIGNMENT//T COFFEE//STATISTICAL ALIGNMENT |
| 4 | 0.0000089207 | TREE OF LIFE//HORIZONTAL GENE TRANSFER//LATERAL GENE TRANSFER |
| 5 | 0.0000074949 | PROGRAM COMPUTAT SYST BIOL//PERIODIC SELECTION//CHURINCE |
| 6 | 0.0000072185 | ISOCHORES//EVOLUZ MOL//ISOCHORE |
| 7 | 0.0000068465 | MACROEVOLUTION//TREE BALANCE//YULE MODEL |
| 8 | 0.0000066171 | CIRCOS//BIODADOS//ORTHOLOGY |
| 9 | 0.0000065402 | ERYTHROCYTE POTASSIUM//BLOOD POLYMORPHISM//HB POLYMORPHISM |
| 10 | 0.0000065055 | BLAST SEARCH//GENES FOR BASIC HELIX LOOP HELIX PROTEINS//ORTHOLOGOUS FAMILY |