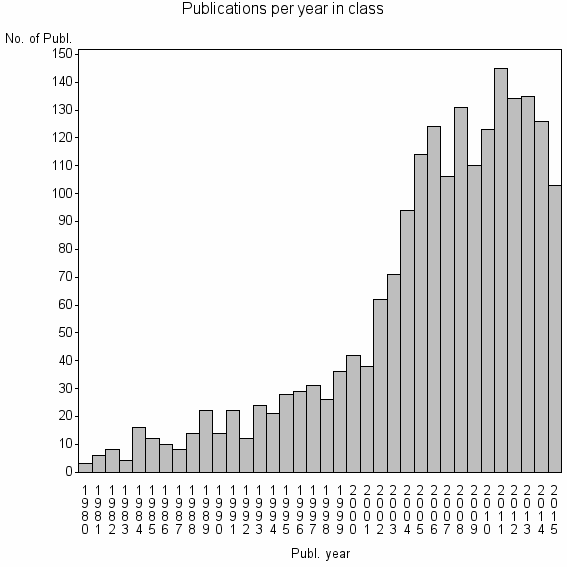
Class information for: |
Basic class information |
| ID | Publications | Average number of references |
Avg. shr. active ref. in WoS |
|---|---|---|---|
| 3040 | 2004 | 40.0 | 77% |
Classes in level above (level 2) |
Terms with highest relevance score |
| Rank | Term | Type of term | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
|---|---|---|---|---|---|---|
| 1 | RNA SECONDARY STRUCTURE | Author keyword | 45 | 29% | 7% | 132 |
| 2 | RNA SECONDARY STRUCTURE PREDICTION | Author keyword | 29 | 65% | 1% | 28 |
| 3 | RNA STRUCTURE PREDICTION | Author keyword | 20 | 67% | 1% | 18 |
| 4 | PSEUDOKNOTS | Author keyword | 19 | 56% | 1% | 23 |
| 5 | PSEUDOKNOT | Author keyword | 11 | 24% | 2% | 41 |
| 6 | NONCODING RNA TECHNOL HLTH | Address | 9 | 28% | 1% | 28 |
| 7 | RNA STRUCTURAL MOTIF | Author keyword | 9 | 83% | 0% | 5 |
| 8 | K NONCROSSING RNA STRUCTURE | Author keyword | 8 | 100% | 0% | 5 |
| 9 | RNA FOLDING | Author keyword | 7 | 14% | 3% | 51 |
| 10 | RNA MOTIF | Author keyword | 6 | 38% | 1% | 13 |
Web of Science journal categories |
Author Key Words |
| Rank | Web of Science journal category | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
LCSH search | Wikipedia search |
|---|---|---|---|---|---|---|---|
| 1 | RNA SECONDARY STRUCTURE | 45 | 29% | 7% | 132 | Search RNA+SECONDARY+STRUCTURE | Search RNA+SECONDARY+STRUCTURE |
| 2 | RNA SECONDARY STRUCTURE PREDICTION | 29 | 65% | 1% | 28 | Search RNA+SECONDARY+STRUCTURE+PREDICTION | Search RNA+SECONDARY+STRUCTURE+PREDICTION |
| 3 | RNA STRUCTURE PREDICTION | 20 | 67% | 1% | 18 | Search RNA+STRUCTURE+PREDICTION | Search RNA+STRUCTURE+PREDICTION |
| 4 | PSEUDOKNOTS | 19 | 56% | 1% | 23 | Search PSEUDOKNOTS | Search PSEUDOKNOTS |
| 5 | PSEUDOKNOT | 11 | 24% | 2% | 41 | Search PSEUDOKNOT | Search PSEUDOKNOT |
| 6 | RNA STRUCTURAL MOTIF | 9 | 83% | 0% | 5 | Search RNA+STRUCTURAL+MOTIF | Search RNA+STRUCTURAL+MOTIF |
| 7 | K NONCROSSING RNA STRUCTURE | 8 | 100% | 0% | 5 | Search K+NONCROSSING+RNA+STRUCTURE | Search K+NONCROSSING+RNA+STRUCTURE |
| 8 | RNA FOLDING | 7 | 14% | 3% | 51 | Search RNA+FOLDING | Search RNA+FOLDING |
| 9 | RNA MOTIF | 6 | 38% | 1% | 13 | Search RNA+MOTIF | Search RNA+MOTIF |
| 10 | NEAREST NEIGHBOR PARAMETERS | 6 | 100% | 0% | 4 | Search NEAREST+NEIGHBOR+PARAMETERS | Search NEAREST+NEIGHBOR+PARAMETERS |
Key Words Plus |
| Rank | Web of Science journal category | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
|---|---|---|---|---|---|
| 1 | INCLUDING PSEUDOKNOTS | 152 | 74% | 6% | 113 |
| 2 | PSEUDOKNOTS | 71 | 56% | 4% | 86 |
| 3 | CONTEXT FREE GRAMMARS | 52 | 42% | 5% | 95 |
| 4 | CONSENSUS STRUCTURE PREDICTION | 50 | 88% | 1% | 23 |
| 5 | DYNAMIC PROGRAMMING ALGORITHM | 44 | 52% | 3% | 60 |
| 6 | ABSTRACT SHAPES | 36 | 83% | 1% | 20 |
| 7 | INTERNAL LOOPS | 35 | 57% | 2% | 41 |
| 8 | SECONDARY STRUCTURE PREDICTION | 32 | 15% | 10% | 203 |
| 9 | STRUCTURE DATABASE ANALYSIS | 31 | 92% | 1% | 12 |
| 10 | SELECTIVE 2 HYDROXYL ACYLATION | 26 | 41% | 2% | 49 |
Journals |
Reviews |
| Title | Publ. year | Cit. | Active references | % act. ref. to same field |
|---|---|---|---|---|
| Expanded sequence dependence of thermodynamic parameters improves prediction of RNA secondary structure | 1999 | 2499 | 93 | 54% |
| Prediction of RNA secondary structure by free energy minimization | 2006 | 155 | 50 | 82% |
| The building blocks and motifs of RNA architecture | 2006 | 137 | 58 | 71% |
| Understanding the transcriptome through RNA structure | 2011 | 90 | 143 | 36% |
| Thermodynamic parameters for an expanded nearest-neighbor model for formation of RNA duplexes with Watson-Crick base pairs | 1998 | 540 | 72 | 47% |
| SHAPE-directed RNA secondary structure prediction | 2010 | 72 | 72 | 47% |
| Insights into RNA structure and function from genome-wide studies | 2014 | 22 | 95 | 32% |
| Advances in RNA structure analysis by chemical probing | 2010 | 67 | 52 | 44% |
| Computational approaches to 3D modeling of RNA | 2010 | 44 | 128 | 56% |
| Predicting thermodynamic properties of RNA | 1995 | 207 | 18 | 78% |
Address terms |
| Rank | Address term | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
|---|---|---|---|---|---|
| 1 | NONCODING RNA TECHNOL HLTH | 9 | 28% | 1.4% | 28 |
| 2 | INTERDISCIPLINARY BIOINFORMAT | 6 | 13% | 2.1% | 42 |
| 3 | CCNSB | 3 | 50% | 0.2% | 5 |
| 4 | IBHV | 3 | 22% | 0.7% | 14 |
| 5 | BIOINFORMAT COMPUTAT BIOL GRP | 3 | 45% | 0.2% | 5 |
| 6 | GRP THEORET BIOL PHYLOGENET | 3 | 60% | 0.1% | 3 |
| 7 | HUMAN GENET MOL PEDIAT DIS | 2 | 24% | 0.4% | 8 |
| 8 | COMP CONTROL MANAGEMENT ENGN DIAG | 2 | 67% | 0.1% | 2 |
| 9 | INFORMAT LIX | 2 | 17% | 0.5% | 11 |
| 10 | RNOM GRP | 2 | 18% | 0.4% | 9 |
Related classes at same level (level 1) |
| Rank | Relatedness score | Related classes |
|---|---|---|
| 1 | 0.0000174545 | BASE PAIR KINETICS//GNRA TETRALOOPS//RAAA HAIRPIN MOTIF |
| 2 | 0.0000164894 | RIBOSWITCH//RIBOSWITCHES//STREPTOMYCES DAVAWENSIS |
| 3 | 0.0000133754 | GROUP I INTRON//SELF SPLICING//RIBOZYME |
| 4 | 0.0000106866 | Q BETA REPLICASE//POLONIES//RNA PHAGE |
| 5 | 0.0000100779 | TREE EDIT DISTANCE//LARGEST COMMON SUBTREE//UNORDERED TREES |
| 6 | 0.0000076519 | FRAMESHIFTING//RIBOSOMAL FRAMESHIFTING//RNA PSEUDOKNOT |
| 7 | 0.0000069732 | MULTIPLE SEQUENCE ALIGNMENT//T COFFEE//STATISTICAL ALIGNMENT |
| 8 | 0.0000069429 | HFQ//S RNA//RYHB |
| 9 | 0.0000063729 | INTERACTION PROPENSITY//RNA BINDING RESIDUES//COMPUTAT INTELLIGENCE LEARNING DISCOVERY |
| 10 | 0.0000059782 | HAMMERHEAD RIBOZYME//HAIRPIN RIBOZYME//RIBOZYME |