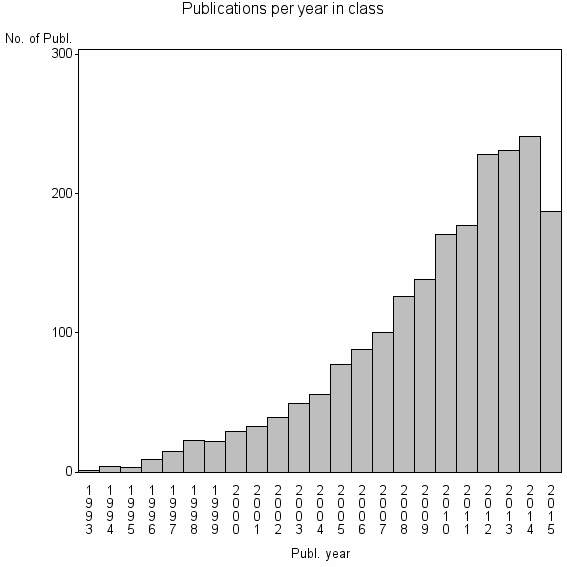
Class information for: |
Basic class information |
| ID | Publications | Average number of references |
Avg. shr. active ref. in WoS |
|---|---|---|---|
| 2886 | 2047 | 48.7 | 93% |
Classes in level above (level 2) |
Terms with highest relevance score |
| Rank | Term | Type of term | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
|---|---|---|---|---|---|---|
| 1 | P110 ALPHA | Author keyword | 70 | 89% | 2% | 32 |
| 2 | PIK3CA | Author keyword | 48 | 33% | 6% | 118 |
| 3 | PI3K ALPHA | Author keyword | 38 | 86% | 1% | 19 |
| 4 | PI3K GAMMA | Author keyword | 37 | 73% | 1% | 29 |
| 5 | PI3K | Author keyword | 33 | 12% | 12% | 250 |
| 6 | PI3K INHIBITOR | Author keyword | 24 | 52% | 2% | 33 |
| 7 | P110 DELTA | Author keyword | 24 | 82% | 1% | 14 |
| 8 | P110 BETA | Author keyword | 19 | 76% | 1% | 13 |
| 9 | ZSTK474 | Author keyword | 19 | 76% | 1% | 13 |
| 10 | PI3K INHIBITORS | Author keyword | 18 | 58% | 1% | 21 |
Web of Science journal categories |
Author Key Words |
| Rank | Web of Science journal category | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
LCSH search | Wikipedia search |
|---|---|---|---|---|---|---|---|
| 1 | P110 ALPHA | 70 | 89% | 2% | 32 | Search P110+ALPHA | Search P110+ALPHA |
| 2 | PIK3CA | 48 | 33% | 6% | 118 | Search PIK3CA | Search PIK3CA |
| 3 | PI3K ALPHA | 38 | 86% | 1% | 19 | Search PI3K+ALPHA | Search PI3K+ALPHA |
| 4 | PI3K GAMMA | 37 | 73% | 1% | 29 | Search PI3K+GAMMA | Search PI3K+GAMMA |
| 5 | PI3K | 33 | 12% | 12% | 250 | Search PI3K | Search PI3K |
| 6 | PI3K INHIBITOR | 24 | 52% | 2% | 33 | Search PI3K+INHIBITOR | Search PI3K+INHIBITOR |
| 7 | P110 DELTA | 24 | 82% | 1% | 14 | Search P110+DELTA | Search P110+DELTA |
| 8 | P110 BETA | 19 | 76% | 1% | 13 | Search P110+BETA | Search P110+BETA |
| 9 | ZSTK474 | 19 | 76% | 1% | 13 | Search ZSTK474 | Search ZSTK474 |
| 10 | PI3K INHIBITORS | 18 | 58% | 1% | 21 | Search PI3K+INHIBITORS | Search PI3K+INHIBITORS |
Key Words Plus |
| Rank | Web of Science journal category | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
|---|---|---|---|---|---|
| 1 | P110 ALPHA | 107 | 76% | 4% | 74 |
| 2 | P110 DELTA | 100 | 73% | 4% | 76 |
| 3 | PI3K | 86 | 44% | 7% | 147 |
| 4 | PIK3CA GENE | 80 | 45% | 6% | 132 |
| 5 | PI3K GAMMA | 66 | 55% | 4% | 82 |
| 6 | P110 GAMMA | 50 | 84% | 1% | 27 |
| 7 | PHOSPHOINOSITIDE 3 KINASE | 49 | 10% | 22% | 448 |
| 8 | PI3K GAMMA INHIBITION | 45 | 90% | 1% | 19 |
| 9 | PHOSPHOINOSITIDE 3 KINASE P110 DELTA | 44 | 77% | 1% | 30 |
| 10 | P85 REGULATORY SUBUNIT | 42 | 69% | 2% | 36 |
Journals |
Reviews |
| Title | Publ. year | Cit. | Active references | % act. ref. to same field |
|---|---|---|---|---|
| PI3K in cancer: divergent roles of isoforms, modes of activation and therapeutic targeting | 2015 | 9 | 195 | 68% |
| PI3K and cancer: lessons, challenges and opportunities | 2014 | 149 | 215 | 29% |
| Targeting PI3 kinase in cancer | 2015 | 9 | 56 | 52% |
| Development of PI3K inhibitors: lessons learned from early clinical trials | 2013 | 151 | 72 | 43% |
| The PI3K Pathway As Drug Target in Human Cancer | 2010 | 435 | 98 | 51% |
| Targeting PI3K signalling in cancer: opportunities, challenges and limitations | 2009 | 875 | 150 | 41% |
| Targeting the phosphoinositide 3-kinase pathway in cancer | 2009 | 779 | 195 | 45% |
| The emerging mechanisms of isoform-specific PI3K signalling | 2010 | 377 | 166 | 50% |
| Picking the Point of Inhibition: A Comparative Review of PI3K/AKT/mTOR Pathway Inhibitors | 2014 | 35 | 51 | 59% |
| PI3K pathway alterations in cancer: variations on a theme | 2008 | 643 | 83 | 47% |
Address terms |
| Rank | Address term | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
|---|---|---|---|---|---|
| 1 | HIGH THROUGHPUT SCREENING MOL PHARMACOL | 11 | 78% | 0.3% | 7 |
| 2 | LYMPHOCYTE SIGNALLING DEV | 7 | 21% | 1.4% | 29 |
| 3 | CANC METAB BIOL | 6 | 100% | 0.2% | 4 |
| 4 | CANC METAB CHEM | 6 | 100% | 0.2% | 4 |
| 5 | CANC METAB DMPK | 6 | 100% | 0.2% | 4 |
| 6 | STRUCT BIOL MOL MODELING | 4 | 75% | 0.1% | 3 |
| 7 | ONCOL DIS AREA | 4 | 32% | 0.5% | 10 |
| 8 | ORIGINAL NEW DRUGS DESIGN DISCOVERY | 3 | 39% | 0.3% | 7 |
| 9 | SMALL MOL TRANSLAT ONCOL | 3 | 100% | 0.1% | 3 |
| 10 | INOSITIDE SIGNALLING GRP | 3 | 40% | 0.3% | 6 |
Related classes at same level (level 1) |
| Rank | Relatedness score | Related classes |
|---|---|---|
| 1 | 0.0000257388 | BIOL CHEMBUNKYO KU//DEMETHOXYVIRIDIN//2 4 MORPHOLINYL 8 PHENYLCHROMONE |
| 2 | 0.0000164790 | PDK1//AKT ISOFORMS//AKT2 |
| 3 | 0.0000118986 | MTOR//RAPAMYCIN//CCI 779 |
| 4 | 0.0000090450 | PTEN//LHERMITTE DUCLOS DISEASE//COWDEN SYNDROME |
| 5 | 0.0000084694 | SHIP2//INOSITOL 5 PHOSPHATASE//SHIP1 |
| 6 | 0.0000067862 | PIKFYVE//PTDINS5P//GOLPH3 |
| 7 | 0.0000054946 | CHEMOTAXIS SIGNAL SECT//GRADIENT SENSING//DIRECTIONAL SENSING |
| 8 | 0.0000046202 | C PEPTICLE//CLIN PHYSIOL METAB UNIT//HENRY WELLCOME S EGRATED CELL SIGNALLING |
| 9 | 0.0000042086 | HALENAQUINONE//PERHYDROPHENANTHRENES//XESTOQUINONE |
| 10 | 0.0000041923 | INSULIN RECEPTOR SUBSTRATE//INSULIN RECEPTOR SUBSTRATE 1//IRS 1 |