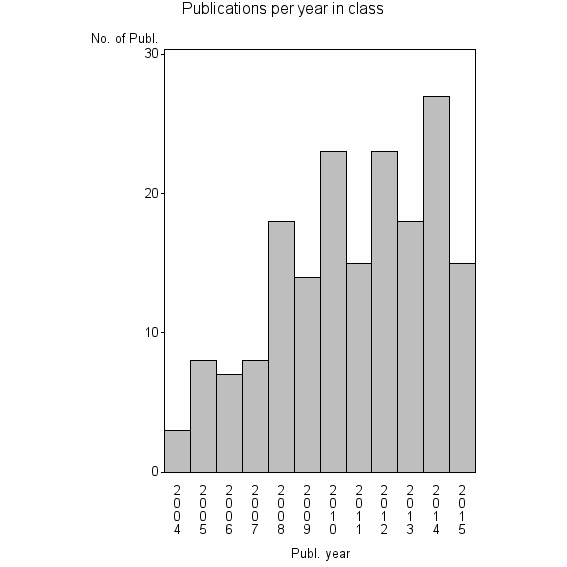
Class information for: |
Basic class information |
| ID | Publications | Average number of references |
Avg. shr. active ref. in WoS |
|---|---|---|---|
| 27902 | 179 | 47.1 | 94% |
Classes in level above (level 2) |
Terms with highest relevance score |
| Rank | Term | Type of term | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
|---|---|---|---|---|---|---|
| 1 | SMALL ACTIVATING RNA | Author keyword | 12 | 86% | 3% | 6 |
| 2 | SARNA | Author keyword | 9 | 83% | 3% | 5 |
| 3 | RNA ACTIVATION | Author keyword | 5 | 60% | 3% | 6 |
| 4 | RNA ACTIVATION RNAA | Author keyword | 4 | 75% | 2% | 3 |
| 5 | SMALL ACTIVATING RNA SARNA | Author keyword | 4 | 75% | 2% | 3 |
| 6 | TRANSCRIPTIONAL GENE SILENCING | Author keyword | 3 | 17% | 8% | 15 |
| 7 | TRANSCRIPTIONAL GENE SILENCING TGS | Author keyword | 2 | 50% | 2% | 3 |
| 8 | SENDAI VIROSOME | Author keyword | 1 | 50% | 1% | 2 |
| 9 | TRANSCRIPTIONAL GENE ACTIVATION | Author keyword | 1 | 100% | 1% | 2 |
| 10 | RNAA | Author keyword | 1 | 17% | 4% | 7 |
Web of Science journal categories |
Author Key Words |
| Rank | Web of Science journal category | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
LCSH search | Wikipedia search |
|---|---|---|---|---|---|---|---|
| 1 | SMALL ACTIVATING RNA | 12 | 86% | 3% | 6 | Search SMALL+ACTIVATING+RNA | Search SMALL+ACTIVATING+RNA |
| 2 | SARNA | 9 | 83% | 3% | 5 | Search SARNA | Search SARNA |
| 3 | RNA ACTIVATION | 5 | 60% | 3% | 6 | Search RNA+ACTIVATION | Search RNA+ACTIVATION |
| 4 | RNA ACTIVATION RNAA | 4 | 75% | 2% | 3 | Search RNA+ACTIVATION+RNAA | Search RNA+ACTIVATION+RNAA |
| 5 | SMALL ACTIVATING RNA SARNA | 4 | 75% | 2% | 3 | Search SMALL+ACTIVATING+RNA+SARNA | Search SMALL+ACTIVATING+RNA+SARNA |
| 6 | TRANSCRIPTIONAL GENE SILENCING | 3 | 17% | 8% | 15 | Search TRANSCRIPTIONAL+GENE+SILENCING | Search TRANSCRIPTIONAL+GENE+SILENCING |
| 7 | TRANSCRIPTIONAL GENE SILENCING TGS | 2 | 50% | 2% | 3 | Search TRANSCRIPTIONAL+GENE+SILENCING+TGS | Search TRANSCRIPTIONAL+GENE+SILENCING+TGS |
| 8 | SENDAI VIROSOME | 1 | 50% | 1% | 2 | Search SENDAI+VIROSOME | Search SENDAI+VIROSOME |
| 9 | TRANSCRIPTIONAL GENE ACTIVATION | 1 | 100% | 1% | 2 | Search TRANSCRIPTIONAL+GENE+ACTIVATION | Search TRANSCRIPTIONAL+GENE+ACTIVATION |
| 10 | RNAA | 1 | 17% | 4% | 7 | Search RNAA | Search RNAA |
Key Words Plus |
| Rank | Web of Science journal category | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
|---|---|---|---|---|---|
| 1 | INDUCE TRANSCRIPTIONAL ACTIVATION | 15 | 82% | 5% | 9 |
| 2 | NONCODING TRANSCRIPT | 11 | 100% | 3% | 6 |
| 3 | DUPLEX RNAS | 9 | 83% | 3% | 5 |
| 4 | ANTIGENE RNAS | 6 | 100% | 2% | 4 |
| 5 | SMALL ACTIVATING RNA | 4 | 75% | 2% | 3 |
| 6 | MEDIATED OVEREXPRESSION | 1 | 24% | 2% | 4 |
| 7 | NATURAL ANTISENSE | 1 | 25% | 2% | 3 |
| 8 | TARGET CHROMOSOMAL DNA | 1 | 33% | 1% | 2 |
| 9 | DIRECTED DE NOVO | 1 | 50% | 1% | 1 |
| 10 | HETEROCHROMATIZATION | 1 | 50% | 1% | 1 |
Journals |
Reviews |
| Title | Publ. year | Cit. | Active references |
% act. ref. to same field |
|---|---|---|---|---|
| Chromatin remodeling by the small RNA machinery in mammalian cells | 2014 | 10 | 77 | 43% |
| Small RNA and transcriptional upregulation | 2011 | 37 | 76 | 33% |
| RNA activation: Promise as a new weapon against cancer | 2014 | 2 | 50 | 66% |
| Direct transcriptional regulation by nuclear microRNAs | 2014 | 7 | 86 | 37% |
| RNA-Directed Transcriptional Gene Silencing and Activation in Human Cells | 2009 | 27 | 48 | 50% |
| RNA-mediated gene activation | 2014 | 7 | 84 | 24% |
| Transcriptional gene silencing through epigenetic changes mediated by non-coding RNAs | 2010 | 37 | 96 | 22% |
| Controlling transcription with noncoding RNAs in mammalian cells | 2010 | 14 | 58 | 34% |
| Promoter-associated RNAs and promoter-targeted RNAs | 2012 | 8 | 78 | 31% |
| Nuclear microRNAs and their unconventional role in regulating non-coding RNAs | 2013 | 10 | 25 | 20% |
Address terms |
| Rank | Address term | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
|---|---|---|---|---|---|
| 1 | KELLOGG SCI TECHNOL | 1 | 12% | 3.9% | 7 |
| 2 | BIOTECHNOL BIOMOL MED | 1 | 50% | 0.6% | 1 |
| 3 | SENTERET | 1 | 50% | 0.6% | 1 |
| 4 | CHEM BIOCHEM HEMATOL | 0 | 33% | 0.6% | 1 |
| 5 | HEPAT DIFFERENTIAT | 0 | 25% | 0.6% | 1 |
| 6 | HIV PATHOGENESIS UNIT | 0 | 20% | 0.6% | 1 |
| 7 | CLIN DRUG DEV UNIT | 0 | 17% | 0.6% | 1 |
| 8 | PFIZER OLIGONUCLEOTIDE THER EUT UNIT | 0 | 17% | 0.6% | 1 |
| 9 | BIOMED BIOTECHNOL UNIT BIOBRU | 0 | 13% | 0.6% | 1 |
| 10 | FRONTIER SCIMINATO KU | 0 | 10% | 0.6% | 1 |
Related classes at same level (level 1) |
| Rank | Relatedness score | Related classes |
|---|---|---|
| 1 | 0.0000208113 | LONG NONCODING RNAS//LNCRNA//LONG NON CODING RNA |
| 2 | 0.0000192782 | PIRNA//PIWI//PIRNAS |
| 3 | 0.0000192525 | WRAP53//OVERLAPPING GENES//CUSTOM BIOTECHNOL SERV GRP |
| 4 | 0.0000166716 | RNA INTERFERENCE//SIRNA//RNAI |
| 5 | 0.0000143934 | GW182//DROSHA//ARGONAUTE |
| 6 | 0.0000126743 | HP1//POSITION EFFECT VARIEGATION//HETEROCHROMATIN |
| 7 | 0.0000123026 | DICER//DROSHA//MICROPROCESSOR COMPLEX |
| 8 | 0.0000084719 | O I//VIRAL MICRORNAS//VIRAL MIRNA |
| 9 | 0.0000073188 | P21//P21WAF1 CIP1//WAF1 |
| 10 | 0.0000051488 | RECK//REVERSION INDUCING CYSTEINE RICH PROTEIN WITH KAZAL MOTIFS//KINOME PROFILING UNIT |