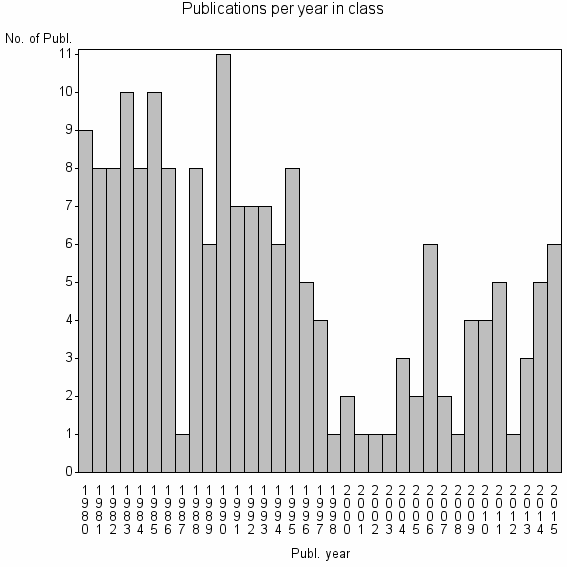
Class information for: |
Basic class information |
| ID | Publications | Average number of references |
Avg. shr. active ref. in WoS |
|---|---|---|---|
| 27893 | 179 | 38.1 | 50% |
Classes in level above (level 2) |
| ID, lev. above |
Publications | Label for level above |
|---|---|---|
| 42 | 31225 | PROTEINS-STRUCTURE FUNCTION AND BIOINFORMATICS//PROTEIN STRUCTURE PREDICTION//PROTEIN FOLDING |
Terms with highest relevance score |
| Rank | Term | Type of term | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
|---|---|---|---|---|---|---|
| 1 | BIOINFORMAT TELEMED | Address | 8 | 31% | 13% | 23 |
| 2 | HYDROPHOBICITY DEFICIENCY | Author keyword | 2 | 67% | 1% | 2 |
| 3 | HELIX INTERACTIONS | Author keyword | 1 | 38% | 2% | 3 |
| 4 | LATE STAGE FOLDING | Author keyword | 1 | 100% | 1% | 2 |
| 5 | HEPTAD MOTIF | Author keyword | 1 | 40% | 1% | 2 |
| 6 | ACTIVE SITE RECOGNITION | Author keyword | 1 | 50% | 1% | 1 |
| 7 | CONFORMATIONAL PROPENSITIES | Author keyword | 1 | 50% | 1% | 1 |
| 8 | DIVERGENCE ENTROPY | Author keyword | 1 | 50% | 1% | 1 |
| 9 | HELIX HELIX INTERFACE | Author keyword | 1 | 50% | 1% | 1 |
| 10 | INTERMEDIATES IN PROTEIN FOLDING | Author keyword | 1 | 50% | 1% | 1 |
Web of Science journal categories |
Author Key Words |
Key Words Plus |
| Rank | Web of Science journal category | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
|---|---|---|---|---|---|
| 1 | ENERGETIC APPROACH | 7 | 44% | 7% | 12 |
| 2 | SPATIAL GEOMETRIC ARRANGEMENTS | 3 | 57% | 2% | 4 |
| 3 | BETA PACKING | 3 | 100% | 2% | 3 |
| 4 | 4 ALPHA HELICAL PROTEINS | 3 | 45% | 3% | 5 |
| 5 | RIGHT HANDED TWIST | 3 | 24% | 6% | 11 |
| 6 | DISULFIDE CROSSLINKED LOOPS | 1 | 50% | 1% | 2 |
| 7 | ALPHA TURN TYPES | 1 | 50% | 1% | 1 |
| 8 | CROSSLINKED LOOPS | 1 | 50% | 1% | 1 |
| 9 | TO STRUCTURE RELATION | 1 | 50% | 1% | 1 |
| 10 | COMPLEX SALT BRIDGES | 0 | 33% | 1% | 1 |
Journals |
Reviews |
| Title | Publ. year | Cit. | Active references | % act. ref. to same field |
|---|---|---|---|---|
| Insight into a molecular interaction force supporting peptide backbones and its implication to protein loops and folding | 2015 | 0 | 96 | 25% |
| STRUCTURAL-PROPERTIES OF PROTEIN BETA-SHEETS | 1983 | 165 | 17 | 47% |
| RATIONALIZATION OF MOLECULAR-MODELS | 1985 | 2 | 1 | 100% |
| PREDICTION OF PROTEIN-FOLDING TYPES FROM AMINO-ACID-COMPOSITION BY CORRELATION ANGLES | 1994 | 0 | 13 | 46% |
| Intrinsically Disordered Proteins-Relation to General Model Expressing the Active Role of the Water Environment | 2014 | 0 | 32 | 9% |
| ORIENTATION OF AMINO-ACID SIDE-CHAINS - INTRAPROTEIN AND SOLVENT INTERACTIONS | 1986 | 9 | 28 | 25% |
| THEORETICAL AND EMPIRICAL APPROACHES TO PROTEIN-STRUCTURE PREDICTION AND ANALYSIS | 1991 | 5 | 262 | 11% |
| AN EMPIRICAL POTENTIAL FUNCTION FOR THE INTERACTION BETWEEN UNIVALENT IONS IN WATER | 1982 | 6 | 16 | 13% |
Address terms |
| Rank | Address term | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
|---|---|---|---|---|---|
| 1 | BIOINFORMAT TELEMED | 8 | 31% | 13% | 23 |
| 2 | NANNING FERMENTAT ENZYME ENGN | 1 | 50% | 0.6% | 1 |
| 3 | CNRSUMR7590 | 0 | 25% | 0.6% | 1 |
| 4 | WOLFSON BIOINFORMAT | 0 | 25% | 0.6% | 1 |
| 5 | BIOTECHNOL CHEM ENVIRONM SCI | 0 | 20% | 0.6% | 1 |
| 6 | EQUIPE SYST MOL BIOL STRUCT | 0 | 14% | 0.6% | 1 |
| 7 | CHRISTIAN DOPPLER INNOVAT THER Y PROACHE S | 0 | 100% | 0.6% | 1 |
Related classes at same level (level 1) |
| Rank | Relatedness score | Related classes |
|---|---|---|
| 1 | 0.0000258553 | LOOP PREDICTION//LOOP MODELING//STEPWISE FOLDING |
| 2 | 0.0000192483 | SECONDARY STRUCTURE PREDICTION//PROTEIN SECONDARY STRUCTURE PREDICTION//COMPOUND PYRAMID MODEL |
| 3 | 0.0000163884 | BPTI//BAKER CHEM CHEM BIOL//SCRAMBLED PROTEINS |
| 4 | 0.0000150818 | HP MODEL//STATISTICAL POTENTIALS//PROTEIN STRUCTURE PREDICTION |
| 5 | 0.0000144525 | TIME DEPENDENT TRANSITION RATES//G MILL FLUID DYNAM NANOSCI IND MATH// |
| 6 | 0.0000138391 | BAKER CHEM CHEM BIOL//REPLICA EXCHANGE METHOD//GENERALIZED ENSEMBLE ALGORITHM |
| 7 | 0.0000132699 | PSEUDO AMINO ACID COMPOSITION//COMPUTAT MUTAT PROJECT//JACKKNIFE TEST |
| 8 | 0.0000111082 | JOURNAL OF MOLECULAR GRAPHICS//SPACE FILLING MOLECULAR MODELS//CPK MODELING |
| 9 | 0.0000107232 | BETA HAIRPIN//SIDE CHAIN INTERACTIONS//TRP CAGE |
| 10 | 0.0000098866 | CYTOCHROME P460//N N BIS CARBOXYMETHYL N N DINITROSO 1 4 PHENYLENEDIAMINE//NITROSYLHEME |