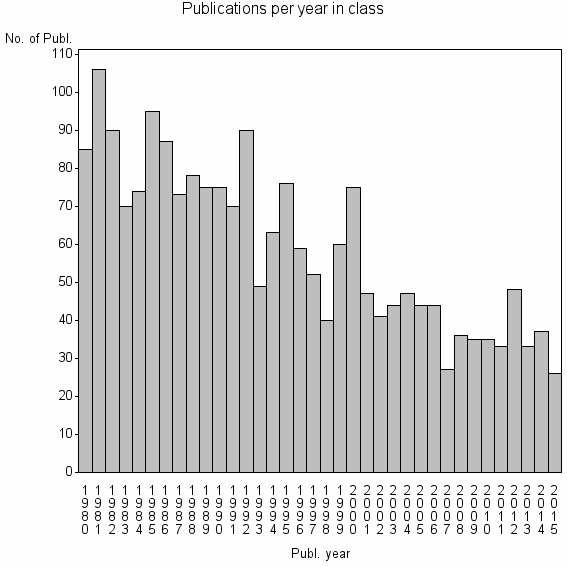
Class information for: |
Basic class information |
| ID | Publications | Average number of references |
Avg. shr. active ref. in WoS |
|---|---|---|---|
| 2618 | 2119 | 36.5 | 53% |
Classes in level above (level 2) |
| ID, lev. above |
Publications | Label for level above |
|---|---|---|
| 1363 | 7774 | PARAMECIUM AURELIA SPECIES COMPLEX//EUROPEAN JOURNAL OF PROTISTOLOGY//INFRACILIATURE |
Terms with highest relevance score |
| Rank | Term | Type of term | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
|---|---|---|---|---|---|---|
| 1 | MACRONUCLEUS | Author keyword | 52 | 61% | 3% | 55 |
| 2 | MACRONUCLEAR DEVELOPMENT | Author keyword | 45 | 94% | 1% | 16 |
| 3 | EPIPLASM | Author keyword | 23 | 86% | 1% | 12 |
| 4 | TETRAHYMENA | Author keyword | 19 | 20% | 4% | 87 |
| 5 | DNA ELIMINATION | Author keyword | 18 | 83% | 0% | 10 |
| 6 | ANTARCTIC BIOLOGY | Author keyword | 14 | 100% | 0% | 7 |
| 7 | HYPOTRICHOUS CILIATES | Author keyword | 14 | 100% | 0% | 7 |
| 8 | INTERNAL ELIMINATED SEQUENCES | Author keyword | 14 | 100% | 0% | 7 |
| 9 | HYPOTRICH | Author keyword | 13 | 69% | 1% | 11 |
| 10 | MACRONUCLEAR DNA | Author keyword | 11 | 67% | 0% | 10 |
Web of Science journal categories |
Author Key Words |
| Rank | Web of Science journal category | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
LCSH search | Wikipedia search |
|---|---|---|---|---|---|---|---|
| 1 | MACRONUCLEUS | 52 | 61% | 3% | 55 | Search MACRONUCLEUS | Search MACRONUCLEUS |
| 2 | MACRONUCLEAR DEVELOPMENT | 45 | 94% | 1% | 16 | Search MACRONUCLEAR+DEVELOPMENT | Search MACRONUCLEAR+DEVELOPMENT |
| 3 | EPIPLASM | 23 | 86% | 1% | 12 | Search EPIPLASM | Search EPIPLASM |
| 4 | TETRAHYMENA | 19 | 20% | 4% | 87 | Search TETRAHYMENA | Search TETRAHYMENA |
| 5 | DNA ELIMINATION | 18 | 83% | 0% | 10 | Search DNA+ELIMINATION | Search DNA+ELIMINATION |
| 6 | ANTARCTIC BIOLOGY | 14 | 100% | 0% | 7 | Search ANTARCTIC+BIOLOGY | Search ANTARCTIC+BIOLOGY |
| 7 | HYPOTRICHOUS CILIATES | 14 | 100% | 0% | 7 | Search HYPOTRICHOUS+CILIATES | Search HYPOTRICHOUS+CILIATES |
| 8 | INTERNAL ELIMINATED SEQUENCES | 14 | 100% | 0% | 7 | Search INTERNAL+ELIMINATED+SEQUENCES | Search INTERNAL+ELIMINATED+SEQUENCES |
| 9 | HYPOTRICH | 13 | 69% | 1% | 11 | Search HYPOTRICH | Search HYPOTRICH |
| 10 | MACRONUCLEAR DNA | 11 | 67% | 0% | 10 | Search MACRONUCLEAR+DNA | Search MACRONUCLEAR+DNA |
Key Words Plus |
| Rank | Web of Science journal category | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
|---|---|---|---|---|---|
| 1 | MACRONUCLEAR DEVELOPMENT | 349 | 95% | 5% | 116 |
| 2 | INTERNAL ELIMINATED SEQUENCES | 126 | 95% | 2% | 41 |
| 3 | NUCLEAR DIFFERENTIATION | 118 | 95% | 2% | 39 |
| 4 | DICER LIKE PROTEIN | 88 | 82% | 2% | 51 |
| 5 | EUPLOTES CRASSUS | 78 | 64% | 4% | 77 |
| 6 | MICRONUCLEAR GENOME | 76 | 100% | 1% | 24 |
| 7 | PARENTAL EXPRESSION | 72 | 100% | 1% | 23 |
| 8 | TETRAHYMENA THERMOPHILA | 72 | 32% | 9% | 188 |
| 9 | THERMOPHILA | 56 | 37% | 6% | 123 |
| 10 | PROGRAMMED DNA ELIMINATION | 52 | 80% | 2% | 32 |
Journals |
Reviews |
| Title | Publ. year | Cit. | Active references | % act. ref. to same field |
|---|---|---|---|---|
| THE DNA OF CILIATED PROTOZOA | 1994 | 427 | 239 | 72% |
| DNA Elimination in Ciliates: Transposon Domestication and Genome Surveillance | 2011 | 44 | 93 | 75% |
| Macronuclear genome sequence of the ciliate Tetrahymena thermophila, a model eukaryote | 2006 | 262 | 161 | 27% |
| Genomes on the Edge: Programmed Genome Instability in Ciliates | 2013 | 13 | 94 | 78% |
| Biolistic transformation of macro- and micronuclei | 2000 | 51 | 9 | 100% |
| Tetrahymena genetics: Two nuclei are better than one | 2000 | 69 | 189 | 76% |
| Genome gymnastics: Unique modes of DNA evolution and processing in ciliates | 2000 | 80 | 26 | 65% |
| Nuclear Dualism | 2012 | 13 | 130 | 74% |
| RNA-guided DNA deletion in tetrahymena: An RNAi-based mechanism for programmed genome rearrangements | 2005 | 67 | 82 | 73% |
| Developmental progression of Tetrahymena through the cell cycle and conjugation | 2012 | 10 | 92 | 85% |
Address terms |
| Rank | Address term | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
|---|---|---|---|---|---|
| 1 | DIPARTIMENTO SCI AMBIENTALI NAT | 5 | 63% | 0.2% | 5 |
| 2 | DIPARTIMENTO BIOL MOL CELLULARE ANIM | 4 | 18% | 0.9% | 19 |
| 3 | EUKARYOT MICROBIOL ANIM BIOL | 3 | 57% | 0.2% | 4 |
| 4 | ALLGEMEINE ZOOL GENET | 2 | 16% | 0.6% | 13 |
| 5 | MOL AUTHENTICAT OURCE | 2 | 67% | 0.1% | 2 |
| 6 | EEC | 2 | 21% | 0.4% | 8 |
| 7 | BIOL CELLULAIRE 4 | 2 | 19% | 0.4% | 9 |
| 8 | UPR3404 | 2 | 16% | 0.4% | 9 |
| 9 | ABORAT PROTE MASS SPE OMETRY | 1 | 100% | 0.1% | 2 |
| 10 | COMPUTAT BIOMODELING | 1 | 50% | 0.1% | 2 |
Related classes at same level (level 1) |
| Rank | Relatedness score | Related classes |
|---|---|---|
| 1 | 0.0000221072 | PARAMECIUM//PARAFUSIN//TRICHOCYST |
| 2 | 0.0000151148 | INFRACILIATURE//PROTOZOOL//MARINE CILIATES |
| 3 | 0.0000116286 | PARAMECIUM AURELIA SPECIES COMPLEX//HOLOSPORA//HOLOSPORA OBTUSA |
| 4 | 0.0000092159 | CHROMATIN DIMINUTION//GERM LINE LIMITED CHROMOSOMES//PARASCARIS UNIVALENS |
| 5 | 0.0000066286 | HORMONAL IMPRINTING//GENET CELL IMMUNOBIOL//TETRAHYMENA |
| 6 | 0.0000046291 | SLUDGE BIOTIC INDEX//SPIROTOX//TETRAHYMENA PYRIFORMIS GL |
| 7 | 0.0000043239 | ICHTHYOPHTHIRIUS MULTIFILIIS//PHILASTERIDES DICENTRARCHI//SCUTICOCILIATOSIS |
| 8 | 0.0000041621 | HELIOZOA//ROTOSPHAERIDA//DESMOTHORACIDA |
| 9 | 0.0000039088 | TUBULIN ISOTYPES//TYROSINATION//DETYROSINATION |
| 10 | 0.0000037939 | EUGLENIDA//EUGLENOPHYTA//TRACHELOMONAS |