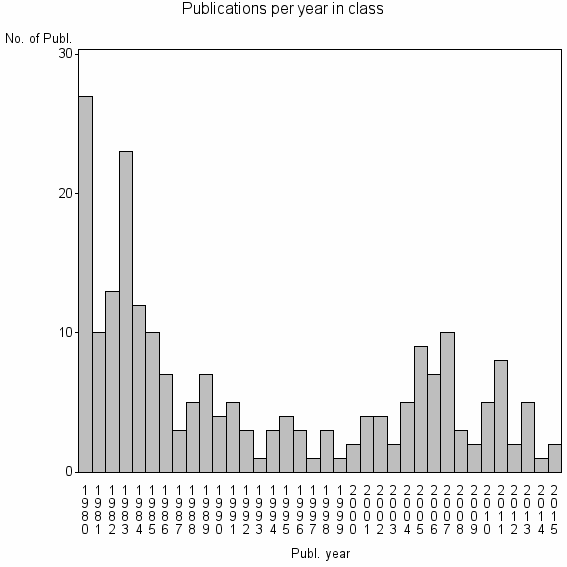
Class information for: |
Basic class information |
| ID | Publications | Average number of references |
Avg. shr. active ref. in WoS |
|---|---|---|---|
| 26138 | 216 | 28.4 | 43% |
Classes in level above (level 2) |
Terms with highest relevance score |
| Rank | Term | Type of term | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
|---|---|---|---|---|---|---|
| 1 | PHAGE TYPING BACTERIAL GENET | Address | 6 | 100% | 2% | 4 |
| 2 | VTB ENZYMTECHNOL | Address | 2 | 50% | 1% | 3 |
| 3 | DEGSU | Author keyword | 1 | 100% | 1% | 2 |
| 4 | TU BCE | Address | 1 | 33% | 1% | 2 |
| 5 | FRUCTOSYLOLIGOSACCHARIDE | Author keyword | 1 | 50% | 0% | 1 |
| 6 | GBF TU BCE | Address | 1 | 50% | 0% | 1 |
| 7 | GRP TU BCE | Address | 1 | 50% | 0% | 1 |
| 8 | HZI HELMHOLTZ INFECT | Address | 1 | 50% | 0% | 1 |
| 9 | IMMUNE INHIBITOR A | Author keyword | 1 | 50% | 0% | 1 |
| 10 | MICROBIOL PATHOGEN | Address | 1 | 50% | 0% | 1 |
Web of Science journal categories |
Author Key Words |
| Rank | Web of Science journal category | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
LCSH search | Wikipedia search |
|---|---|---|---|---|---|---|---|
| 1 | DEGSU | 1 | 100% | 1% | 2 | Search DEGSU | Search DEGSU |
| 2 | FRUCTOSYLOLIGOSACCHARIDE | 1 | 50% | 0% | 1 | Search FRUCTOSYLOLIGOSACCHARIDE | Search FRUCTOSYLOLIGOSACCHARIDE |
| 3 | IMMUNE INHIBITOR A | 1 | 50% | 0% | 1 | Search IMMUNE+INHIBITOR+A | Search IMMUNE+INHIBITOR+A |
| 4 | RESISTANCE TO GLYCOPEPTIDE ANTIBIOTICS | 1 | 50% | 0% | 1 | Search RESISTANCE+TO+GLYCOPEPTIDE+ANTIBIOTICS | Search RESISTANCE+TO+GLYCOPEPTIDE+ANTIBIOTICS |
| 5 | SPORULATION MUTANT | 1 | 50% | 0% | 1 | Search SPORULATION+MUTANT | Search SPORULATION+MUTANT |
| 6 | SUCROSE INDUCIBLE PROMOTER | 1 | 50% | 0% | 1 | Search SUCROSE+INDUCIBLE+PROMOTER | Search SUCROSE+INDUCIBLE+PROMOTER |
| 7 | XYLOSE OPERON | 1 | 50% | 0% | 1 | Search XYLOSE+OPERON | Search XYLOSE+OPERON |
| 8 | HETEROSPECIFIC MATINGS | 0 | 33% | 0% | 1 | Search HETEROSPECIFIC+MATINGS | Search HETEROSPECIFIC+MATINGS |
| 9 | LOW COVERAGE | 0 | 33% | 0% | 1 | Search LOW+COVERAGE | Search LOW+COVERAGE |
| 10 | SUBSTRATE GRADIENTS | 0 | 33% | 0% | 1 | Search SUBSTRATE+GRADIENTS | Search SUBSTRATE+GRADIENTS |
Key Words Plus |
| Rank | Web of Science journal category | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
|---|---|---|---|---|---|
| 1 | SPOIV GENE | 8 | 100% | 2% | 5 |
| 2 | CHROMOSOME EXPRESSION | 6 | 100% | 2% | 4 |
| 3 | SIGNAL PEPTIDASE GENE | 6 | 100% | 2% | 4 |
| 4 | XYLOSE UTILIZATION | 5 | 31% | 6% | 14 |
| 5 | NON COMPLEMENTING DIPLOIDS | 3 | 57% | 2% | 4 |
| 6 | PLASMID SYSTEM | 2 | 38% | 2% | 5 |
| 7 | SUBTILIS DEGU | 2 | 67% | 1% | 2 |
| 8 | CENTRAL METABOLIC FLUXES | 2 | 50% | 1% | 3 |
| 9 | QM B1551 | 2 | 50% | 1% | 3 |
| 10 | UV SENSITIVE MUTANTS | 2 | 40% | 2% | 4 |
Journals |
Reviews |
| Title | Publ. year | Cit. | Active references |
% act. ref. to same field |
|---|---|---|---|---|
| Bacillus megaterium from simple soil bacterium to industrial protein production host | 2007 | 48 | 83 | 41% |
| SYSTEMS BIOLOGY OF RECOMBINANT PROTEIN PRODUCTION USING BACILLUS MEGATERIUM | 2011 | 6 | 56 | 55% |
| PRIME-TIME FOR BACILLUS-MEGATERIUM | 1994 | 93 | 72 | 24% |
| FUSION OF STREPTOCOCCUS-LACTIS PROTOPLASTS | 1987 | 0 | 13 | 23% |
| LYSINE PRODUCTION BY YEASTS AND ITS MECHANISM | 1981 | 0 | 1 | 100% |
| IMPACT OF RECOMBINANT DNA TECHNOLOGY IN PHARMACOLOGY | 1985 | 1 | 19 | 5% |
| SEQUENTIAL EXPLOSIONS - THE MAD YEARS, 1980-1990 | 1983 | 1 | 20 | 5% |
Address terms |
| Rank | Address term | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
|---|---|---|---|---|---|
| 1 | PHAGE TYPING BACTERIAL GENET | 6 | 100% | 1.9% | 4 |
| 2 | VTB ENZYMTECHNOL | 2 | 50% | 1.4% | 3 |
| 3 | TU BCE | 1 | 33% | 0.9% | 2 |
| 4 | GBF TU BCE | 1 | 50% | 0.5% | 1 |
| 5 | GRP TU BCE | 1 | 50% | 0.5% | 1 |
| 6 | HZI HELMHOLTZ INFECT | 1 | 50% | 0.5% | 1 |
| 7 | MICROBIOL PATHOGEN | 1 | 50% | 0.5% | 1 |
| 8 | BCE | 0 | 17% | 0.9% | 2 |
| 9 | BRAUN WEIG INTEGRATED SYST BIOL BRICS | 0 | 17% | 0.5% | 1 |
| 10 | CNRSURA 2225 | 0 | 14% | 0.5% | 1 |
Related classes at same level (level 1) |
| Rank | Relatedness score | Related classes |
|---|---|---|
| 1 | 0.0000223423 | GENOME VECTOR//NIPPONKO KU//LONG ACCURATE PCR |
| 2 | 0.0000177076 | ROLLING CIRCLE REPLICATION//PLASMID REPLICATION CONTROL//CRYPTIC PLASMID |
| 3 | 0.0000114934 | ABRB//SPO0A//TRANSITION STATE REGULATOR |
| 4 | 0.0000089318 | PHARM YAMASHIRO CHO//TRYPSIN LIKE PROTEINASE//INTRACELLULAR SERINE PROTEASE |
| 5 | 0.0000081081 | ALPHA KETO ALPHA HYDROXY ACIDS//PLANT BLEEDING//H ANNUUS |
| 6 | 0.0000074193 | SHANNONS MUTUAL INFORMATION//FLORIDA RACING//COMPLETE DATA |
| 7 | 0.0000071493 | L FORMS//BACTERIAL L FORMS//L FORM CONVERSION |
| 8 | 0.0000064483 | MOL MICROBIAL STRUCT BIOL//SPORE RESISTANCE//SPORE KILLING |
| 9 | 0.0000061377 | NONLINEAR EXPERIMENTAL DESIGN//ALCOHOL TECHNOL//BIOMAT PHENOMENES TRANSPORT LBMPT |
| 10 | 0.0000060851 | LINEAR PLASMID//LINEAR CHROMOSOME//PUR CLUSTER |