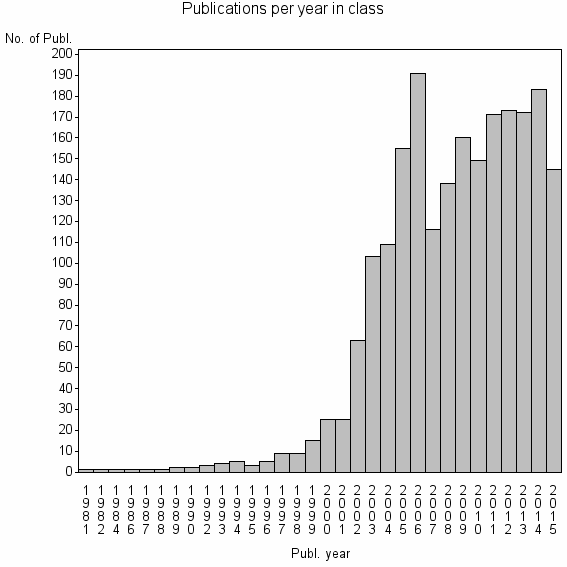
Class information for: |
Basic class information |
| ID | Publications | Average number of references |
Avg. shr. active ref. in WoS |
|---|---|---|---|
| 2550 | 2141 | 32.5 | 58% |
Classes in level above (level 2) |
| ID, lev. above |
Publications | Label for level above |
|---|---|---|
| 396 | 16220 | CREDIT SCORING//MACHINE LEARNING//FEATURE SELECTION |
Terms with highest relevance score |
| Rank | Term | Type of term | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
|---|---|---|---|---|---|---|
| 1 | GENE SELECTION | Author keyword | 82 | 46% | 6% | 134 |
| 2 | FEATURE SELECTION | Author keyword | 66 | 11% | 27% | 585 |
| 3 | CANCER CLASSIFICATION | Author keyword | 49 | 46% | 4% | 81 |
| 4 | MICROARRAY DATA | Author keyword | 14 | 19% | 3% | 66 |
| 5 | MICROARRAY DATA CLASSIFICATION | Author keyword | 13 | 71% | 0% | 10 |
| 6 | GENE EXPRESSION DATA | Author keyword | 10 | 16% | 3% | 57 |
| 7 | BIOINFORMAT GENOM SYST ENGN | Address | 9 | 83% | 0% | 5 |
| 8 | FEATURE GENE SELECTION | Author keyword | 8 | 75% | 0% | 6 |
| 9 | UNSUPERVISED FEATURE SELECTION | Author keyword | 8 | 38% | 1% | 17 |
| 10 | MICROARRAY CLASSIFICATION | Author keyword | 7 | 46% | 1% | 11 |
Web of Science journal categories |
Author Key Words |
| Rank | Web of Science journal category | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
LCSH search | Wikipedia search |
|---|---|---|---|---|---|---|---|
| 1 | GENE SELECTION | 82 | 46% | 6% | 134 | Search GENE+SELECTION | Search GENE+SELECTION |
| 2 | FEATURE SELECTION | 66 | 11% | 27% | 585 | Search FEATURE+SELECTION | Search FEATURE+SELECTION |
| 3 | CANCER CLASSIFICATION | 49 | 46% | 4% | 81 | Search CANCER+CLASSIFICATION | Search CANCER+CLASSIFICATION |
| 4 | MICROARRAY DATA | 14 | 19% | 3% | 66 | Search MICROARRAY+DATA | Search MICROARRAY+DATA |
| 5 | MICROARRAY DATA CLASSIFICATION | 13 | 71% | 0% | 10 | Search MICROARRAY+DATA+CLASSIFICATION | Search MICROARRAY+DATA+CLASSIFICATION |
| 6 | GENE EXPRESSION DATA | 10 | 16% | 3% | 57 | Search GENE+EXPRESSION+DATA | Search GENE+EXPRESSION+DATA |
| 7 | FEATURE GENE SELECTION | 8 | 75% | 0% | 6 | Search FEATURE+GENE+SELECTION | Search FEATURE+GENE+SELECTION |
| 8 | UNSUPERVISED FEATURE SELECTION | 8 | 38% | 1% | 17 | Search UNSUPERVISED+FEATURE+SELECTION | Search UNSUPERVISED+FEATURE+SELECTION |
| 9 | MICROARRAY CLASSIFICATION | 7 | 46% | 1% | 11 | Search MICROARRAY+CLASSIFICATION | Search MICROARRAY+CLASSIFICATION |
| 10 | FEATURE RANKING | 7 | 21% | 1% | 28 | Search FEATURE+RANKING | Search FEATURE+RANKING |
Key Words Plus |
| Rank | Web of Science journal category | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
|---|---|---|---|---|---|
| 1 | CANCER CLASSIFICATION | 91 | 49% | 6% | 135 |
| 2 | FEATURE SUBSET SELECTION | 88 | 40% | 8% | 170 |
| 3 | TUMOR CLASSIFICATION | 67 | 47% | 5% | 107 |
| 4 | CLASS PREDICTION | 43 | 46% | 3% | 70 |
| 5 | EXPRESSION DATA | 30 | 13% | 10% | 216 |
| 6 | GENE EXPRESSION DATA | 22 | 10% | 10% | 204 |
| 7 | MOLECULAR CLASSIFICATION | 18 | 12% | 6% | 137 |
| 8 | INPUT FEATURE SELECTION | 18 | 57% | 1% | 21 |
| 9 | GENE SELECTION | 17 | 25% | 3% | 59 |
| 10 | SVM RFE | 16 | 58% | 1% | 18 |
Journals |
Reviews |
| Title | Publ. year | Cit. | Active references | % act. ref. to same field |
|---|---|---|---|---|
| A review of feature selection techniques in bioinformatics | 2007 | 791 | 98 | 41% |
| A review of feature selection methods based on mutual information | 2014 | 8 | 22 | 86% |
| A review of feature selection methods on synthetic data | 2013 | 20 | 26 | 62% |
| A review of microarray datasets and applied feature selection methods | 2014 | 5 | 86 | 62% |
| Critical review of published microarray studies for cancer outcome and guidelines on statistical analysis and reporting | 2007 | 333 | 18 | 28% |
| Evolutionary computation for feature selection in classification problems | 2013 | 4 | 37 | 59% |
| Stable feature selection for biomarker discovery | 2010 | 39 | 58 | 31% |
| Learning Curves in Classification With Microarray Data | 2010 | 1 | 2 | 100% |
| A Review of Ensemble Methods in Bioinformatics | 2010 | 38 | 95 | 22% |
| Machine learning in bioinformatics | 2006 | 117 | 138 | 14% |
Address terms |
| Rank | Address term | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
|---|---|---|---|---|---|
| 1 | BIOINFORMAT GENOM SYST ENGN | 9 | 83% | 0.2% | 5 |
| 2 | INTELLIGENT COMPUTAT | 5 | 63% | 0.2% | 5 |
| 3 | IND SYST CONTROL | 4 | 50% | 0.3% | 6 |
| 4 | OERSTED DTU AUTOMAT | 4 | 75% | 0.1% | 3 |
| 5 | DEV ARTIFICIAL INTELLIGENCE LIDIA | 3 | 35% | 0.3% | 6 |
| 6 | INTELLIGENT COMP | 2 | 12% | 0.8% | 17 |
| 7 | BIOINFORMAT CELL | 2 | 67% | 0.1% | 2 |
| 8 | HEFEI INTELLIGENT MACHINES | 2 | 10% | 0.9% | 20 |
| 9 | SECT CLIN CANC GENET | 2 | 36% | 0.2% | 4 |
| 10 | INTELLIGENT SYS SCI | 1 | 100% | 0.1% | 2 |
Related classes at same level (level 1) |
| Rank | Relatedness score | Related classes |
|---|---|---|
| 1 | 0.0000145598 | BICLUSTERING//GENE EXPRESSION DATA//GENE EXPRESSION DATA ANALYSIS |
| 2 | 0.0000098243 | ACTUAL ERROR RATE//PRINCIPAL VARIABLES//HIGH DIMENSIONAL MULTIVARIATE BINARY DATA |
| 3 | 0.0000095173 | MICROARRAY IMAGE PROCESSING//GENE FILTERING//MAQC |
| 4 | 0.0000091762 | PHILIP MORRIS INT RD//COMPUTAT PL GENOM PROGRAM//SECT COMPUTAT BIOMED |
| 5 | 0.0000090986 | ERYTHEMATO SQUAMOUS//ERYTHEMATO SQUAMOUS DISEASES//FUZZY WEIGHTED PRE PROCESSING |
| 6 | 0.0000079992 | BOOSTING//BAGGING//ENSEMBLE PRUNING |
| 7 | 0.0000078466 | LASSO//ORACLE PROPERTY//VARIABLE SELECTION |
| 8 | 0.0000076497 | PROTOTYPE SELECTION//INSTANCE SELECTION//PROTOTYPE REDUCTION SCHEMES PRS |
| 9 | 0.0000070399 | MISSING VALUE IMPUTATION//MISSING VALUE ESTIMATION//COMPUTAT MATH BIOL |
| 10 | 0.0000068277 | GENE SET ANALYSIS//ENRICHMENT ANALYSIS//FUNCT GENOM NODE |