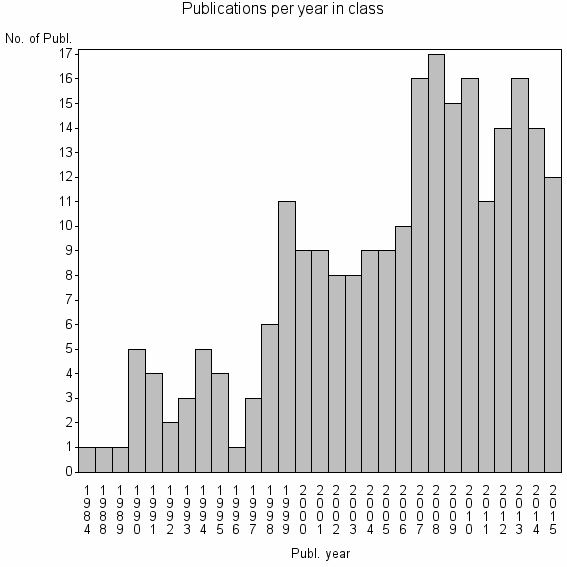
Class information for: |
Basic class information |
| ID | Publications | Average number of references |
Avg. shr. active ref. in WoS |
|---|---|---|---|
| 25198 | 240 | 50.7 | 79% |
Classes in level above (level 2) |
Terms with highest relevance score |
| Rank | Term | Type of term | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
|---|---|---|---|---|---|---|
| 1 | ZINC PROTEINS | Author keyword | 2 | 43% | 1% | 3 |
| 2 | DRUG DESIGN BIOTECHNOL | Address | 1 | 27% | 2% | 4 |
| 3 | METALLOPROTEOME | Author keyword | 1 | 40% | 1% | 2 |
| 4 | METAL BINDING SITES | Author keyword | 1 | 15% | 2% | 5 |
| 5 | BIOENGN GENET | Address | 1 | 50% | 0% | 1 |
| 6 | BRIDGE CONFORMATION | Author keyword | 1 | 50% | 0% | 1 |
| 7 | COMPRESSED ANGLE | Author keyword | 1 | 50% | 0% | 1 |
| 8 | DE NOVO STRUCTURE PREDICTION | Author keyword | 1 | 50% | 0% | 1 |
| 9 | ENVIRONM SYST BIOCHEM | Address | 1 | 50% | 0% | 1 |
| 10 | HIS TAG AFFINITY CHROMATOGRAPHY | Author keyword | 1 | 50% | 0% | 1 |
Web of Science journal categories |
Author Key Words |
Key Words Plus |
| Rank | Web of Science journal category | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
|---|---|---|---|---|---|
| 1 | AMINO ACID CHAINS | 6 | 71% | 2% | 5 |
| 2 | DFT CDM CALCULATIONS | 6 | 100% | 2% | 4 |
| 3 | ZN BINDING | 4 | 75% | 1% | 3 |
| 4 | ALUMINUMIII INTERACTIONS | 3 | 45% | 2% | 5 |
| 5 | CATALYTIC ZINC | 1 | 38% | 1% | 3 |
| 6 | INCLUDING CHARGE TRANSFER | 1 | 38% | 1% | 3 |
| 7 | ACETATE COMPLEXATION | 1 | 100% | 1% | 2 |
| 8 | ACIDIC NACL BRINES | 1 | 50% | 1% | 2 |
| 9 | PDB SURVEY | 1 | 100% | 1% | 2 |
| 10 | CARBOXYLATE INTERACTIONS | 1 | 27% | 2% | 4 |
Journals |
Reviews |
| Title | Publ. year | Cit. | Active references | % act. ref. to same field |
|---|---|---|---|---|
| Effect of carboxylate-binding mode on metal binding/selectivity and function in proteins | 2007 | 62 | 41 | 34% |
| Competition among Metal Ions for Protein Binding Sites: Determinants of Metal Ion Selectivity in Proteins | 2014 | 21 | 174 | 17% |
| Metalloproteomes: A Bioinformatic Approach | 2009 | 67 | 31 | 19% |
| Metal binding affinity and selectivity in metalloproteins: Insights from computational studies | 2008 | 82 | 52 | 31% |
| Web Tools for Predicting Metal Binding Sites in Proteins | 2013 | 0 | 36 | 64% |
| Metals in protein structures: a review of their principal features | 2010 | 41 | 119 | 18% |
| Calciomics: integrative studies of Ca2+-binding proteins and their interactomes in biological systems | 2013 | 7 | 87 | 16% |
| Principles governing Mg, Ca, and Zn binding and selectivity in proteins | 2003 | 232 | 155 | 12% |
| Tools for Predicting Metal Binding Sites in Protein: A Review | 2011 | 2 | 40 | 35% |
| Bioinformatics in bioinorganic chemistry | 2010 | 12 | 125 | 19% |
Address terms |
| Rank | Address term | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
|---|---|---|---|---|---|
| 1 | DRUG DESIGN BIOTECHNOL | 1 | 27% | 1.7% | 4 |
| 2 | BIOENGN GENET | 1 | 50% | 0.4% | 1 |
| 3 | ENVIRONM SYST BIOCHEM | 1 | 50% | 0.4% | 1 |
| 4 | NEQCNUCLEO ESTUDOS QUIM COMPUTAC | 1 | 50% | 0.4% | 1 |
| 5 | TUMOR BIOL PROGRAMMOL NEUROSCI PROGRAM | 1 | 50% | 0.4% | 1 |
| 6 | BIOMOL MODELING BIOENGN | 0 | 33% | 0.4% | 1 |
| 7 | ENZYME MICROBIAL TECHNOL EMTECH | 0 | 33% | 0.4% | 1 |
| 8 | DIPARTIMENTO SIST INFORMAT | 0 | 25% | 0.4% | 1 |
| 9 | STRUCT CHEM ARRHENIUS | 0 | 25% | 0.4% | 1 |
| 10 | CNRSUMR 7654 | 0 | 20% | 0.4% | 1 |
Related classes at same level (level 1) |
| Rank | Relatedness score | Related classes |
|---|---|---|
| 1 | 0.0000193875 | ARTIFICIAL TRANSCRIPTION FACTOR//ARTIFICIAL TRANSCRIPTION FACTORS//RECOGNITION CODE |
| 2 | 0.0000180181 | COMPLEX SCI SOFTWARE//3D ZERNIKE DESCRIPTOR//PROTEIN SURFACE SHAPE |
| 3 | 0.0000130286 | ONCOMODULIN//SARCOPLASMIC CALCIUM BINDING PROTEIN//EF HAND |
| 4 | 0.0000087958 | PROTEIN DESIGN//FOUR HELIX BUNDLE//TEMPLATE ASSEMBLED SYNTHETIC PROTEIN |
| 5 | 0.0000084830 | METALLOMICS//METALLOPROTEOMICS//SPE OMETRY SAMPLE PREPARAT MECHANIZAT GRP GEP |
| 6 | 0.0000078321 | METAL CATION MOLECULE COMPLEX//CONTINUAL EDUC//REFLECTRON TIME OF FLIGHT MASS SPECTROMETER |
| 7 | 0.0000076610 | FERRIC UPTAKE REGULATOR//DTXR//FUR BOX |
| 8 | 0.0000074466 | MINIQUAD75//COMPLEX EQUILIBRIA//DIOXAN |
| 9 | 0.0000069225 | AICARDI GOUTIERES SYNDROME//TREX1//RIBONUCLEASE H |
| 10 | 0.0000064355 | ZINC TRANSPORTERS//ZNT//ZINQUIN |