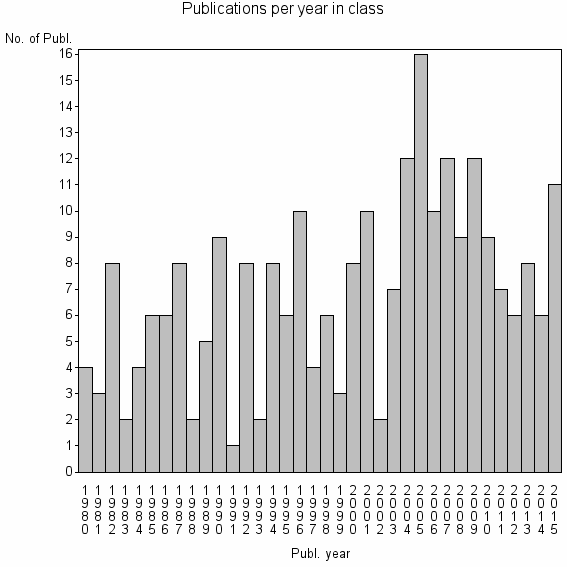
Class information for: |
Basic class information |
| ID | Publications | Average number of references |
Avg. shr. active ref. in WoS |
|---|---|---|---|
| 24848 | 250 | 32.8 | 61% |
Classes in level above (level 2) |
Terms with highest relevance score |
| Rank | Term | Type of term | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
|---|---|---|---|---|---|---|
| 1 | NAD KINASE | Author keyword | 23 | 51% | 13% | 33 |
| 2 | GRP BIOCHIM BIOL MOL VEGETALES | Address | 15 | 82% | 4% | 9 |
| 3 | NADH KINASE | Author keyword | 12 | 75% | 4% | 9 |
| 4 | EA 917 | Address | 11 | 100% | 2% | 6 |
| 5 | BASIC PL MOL BIOTECHNOL | Address | 3 | 17% | 6% | 16 |
| 6 | BIOTECNOL BIOCHIM | Address | 2 | 67% | 1% | 2 |
| 7 | FOOD BIOL SCI | Address | 2 | 22% | 2% | 6 |
| 8 | CAM DEPENDENT NAD KINASE | Author keyword | 1 | 100% | 1% | 2 |
| 9 | GBBMV EA917 | Address | 1 | 100% | 1% | 2 |
| 10 | NAD KINASE NADK | Author keyword | 1 | 100% | 1% | 2 |
Web of Science journal categories |
Author Key Words |
| Rank | Web of Science journal category | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
LCSH search | Wikipedia search |
|---|---|---|---|---|---|---|---|
| 1 | NAD KINASE | 23 | 51% | 13% | 33 | Search NAD+KINASE | Search NAD+KINASE |
| 2 | NADH KINASE | 12 | 75% | 4% | 9 | Search NADH+KINASE | Search NADH+KINASE |
| 3 | CAM DEPENDENT NAD KINASE | 1 | 100% | 1% | 2 | Search CAM+DEPENDENT+NAD+KINASE | Search CAM+DEPENDENT+NAD+KINASE |
| 4 | NAD KINASE NADK | 1 | 100% | 1% | 2 | Search NAD+KINASE+NADK | Search NAD+KINASE+NADK |
| 5 | NADP PHOSPHATASE | 1 | 100% | 1% | 2 | Search NADP+PHOSPHATASE | Search NADP+PHOSPHATASE |
| 6 | NICOTINAMIDE PYRIDINE NUCLEOTIDE | 1 | 100% | 1% | 2 | Search NICOTINAMIDE+PYRIDINE+NUCLEOTIDE | Search NICOTINAMIDE+PYRIDINE+NUCLEOTIDE |
| 7 | POS5P | 1 | 100% | 1% | 2 | Search POS5P | Search POS5P |
| 8 | ACTIVITY BANDS | 1 | 50% | 0% | 1 | Search ACTIVITY+BANDS | Search ACTIVITY+BANDS |
| 9 | APPARENT K M | 1 | 50% | 0% | 1 | Search APPARENT+K+M | Search APPARENT+K+M |
| 10 | CALCIUM CALMODULIN COMPLEX | 1 | 50% | 0% | 1 | Search CALCIUM+CALMODULIN+COMPLEX | Search CALCIUM+CALMODULIN+COMPLEX |
Key Words Plus |
| Rank | Web of Science journal category | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
|---|---|---|---|---|---|
| 1 | MICROCOCCUS FLAVUS | 17 | 100% | 3% | 8 |
| 2 | KLEBS STRAIN Z | 11 | 78% | 3% | 7 |
| 3 | NADP PHOSPHATASE | 6 | 100% | 2% | 4 |
| 4 | ACHLOROPHYLLOUS ZC MUTANT | 5 | 44% | 3% | 8 |
| 5 | MICROBIAL ADAPTATION | 3 | 57% | 2% | 4 |
| 6 | ATCC 13525 | 2 | 38% | 2% | 5 |
| 7 | TUBERCULOSIS NAD KINASE | 2 | 50% | 1% | 3 |
| 8 | NAD KINASE | 2 | 16% | 4% | 10 |
| 9 | CALMODULIN DEPENDENT ISOZYME | 1 | 100% | 1% | 2 |
| 10 | NADH KINASE | 1 | 30% | 1% | 3 |
Journals |
Reviews |
| Title | Publ. year | Cit. | Active references | % act. ref. to same field |
|---|---|---|---|---|
| MNADK, a Long-Awaited Human Mitochondrion-Localized NAD Kinase | 2015 | 1 | 37 | 54% |
| The power to reduce: pyridine nucleotides - small molecules with a multitude of functions | 2007 | 226 | 143 | 25% |
| Structure and function of NAD kinase and NADP phosphatase: Key enzymes that regulate the intracellular balance of NAD(H) and NADP(H) | 2008 | 23 | 64 | 66% |
| Molecular properties, functions, and potential applications of NAD kinases | 2009 | 16 | 69 | 67% |
| Structural and functional properties of NAD kinase, a key enzyme in NADP biosynthesis | 2006 | 18 | 48 | 58% |
| Pseudomonas fluorescens orchestrates a fine metabolic-balancing act to counter aluminium toxicity | 2010 | 22 | 34 | 29% |
| Metabolic reengineering invoked by microbial systems to decontaminate aluminum: Implications for bioremediation technologies | 2013 | 5 | 70 | 23% |
| The phosphate makes a difference: cellular functions of NADP | 2010 | 10 | 64 | 28% |
| NAD - new roles in signalling and gene regulation in plants | 2004 | 53 | 88 | 10% |
| The unravelling of metabolic dysfunctions linked to metal-associated diseases by blue native polyacrylamide gel electrophoresis | 2013 | 0 | 56 | 18% |
Address terms |
| Rank | Address term | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
|---|---|---|---|---|---|
| 1 | GRP BIOCHIM BIOL MOL VEGETALES | 15 | 82% | 3.6% | 9 |
| 2 | EA 917 | 11 | 100% | 2.4% | 6 |
| 3 | BASIC PL MOL BIOTECHNOL | 3 | 17% | 6.4% | 16 |
| 4 | BIOTECNOL BIOCHIM | 2 | 67% | 0.8% | 2 |
| 5 | FOOD BIOL SCI | 2 | 22% | 2.4% | 6 |
| 6 | GBBMV EA917 | 1 | 100% | 0.8% | 2 |
| 7 | GUELPH WATERLOO WORK CHEM BIOCHEM GWC | 1 | 50% | 0.4% | 1 |
| 8 | BIOTECHNOL SECTOR | 1 | 20% | 1.2% | 3 |
| 9 | MICROBIOL INTERACT | 0 | 33% | 0.4% | 1 |
| 10 | IST BIOTECNOL BIOCHIM | 0 | 13% | 1.2% | 3 |
Related classes at same level (level 1) |
| Rank | Relatedness score | Related classes |
|---|---|---|
| 1 | 0.0000158109 | WLDS//NMNAT//NAD BIOSYNTHESIS |
| 2 | 0.0000116784 | TELLURITE RESISTANCE//TELLURITE REDUCTION//TELLURITE |
| 3 | 0.0000099628 | CALCIUM DEPENDENT PROTEIN KINASE//CDPK//CIPK |
| 4 | 0.0000076923 | C 13 AND N 15 LABELLING//GROWING LEAVES//HETEROTROPHY AUTOTROPHY |
| 5 | 0.0000076475 | TRANSHYDROGENASE//NICOTINAMIDE NUCLEOTIDE//NICOTINAMIDE NUCLEOTIDE TRANSHYDROGENASE |
| 6 | 0.0000070868 | CAFFEINE DEGRADATION//METAB BIOL GRPBUNKYO KU//CAFFEINE BIOSYNTHESIS |
| 7 | 0.0000070671 | CALERYTHRIN//CALMODULIN LIKE PROTEIN//DODECIN |
| 8 | 0.0000063770 | EXOPOLYPHOSPHATASE//INORGANIC POLYPHOSPHATES//POLYPHOSPHATASE |
| 9 | 0.0000061404 | 3 ISOPROPYLMALATE DEHYDROGENASE//ISOCITRATE DEHYDROGENASE//ISOPROPYLMALATE DEHYDROGENASE |
| 10 | 0.0000060795 | CADMIUM RESISTANT BACTERIA//CUPRIAVIDUS METALLIDURANS CH34//HEAVY METAL RESISTANCE |