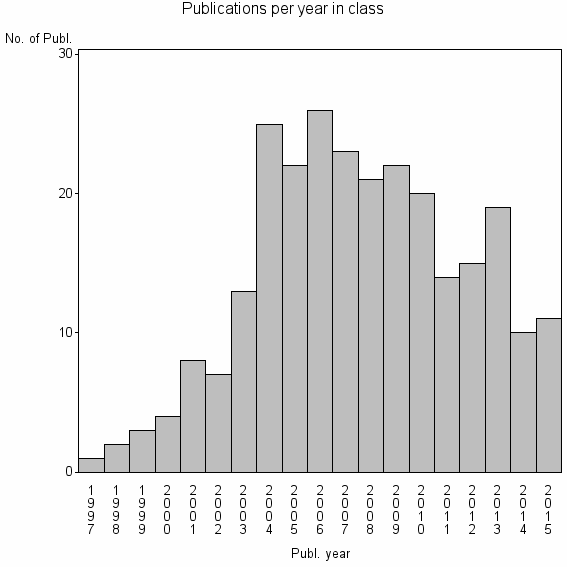
Class information for: |
Basic class information |
| ID | Publications | Average number of references |
Avg. shr. active ref. in WoS |
|---|---|---|---|
| 24275 | 266 | 42.8 | 85% |
Classes in level above (level 2) |
| ID, lev. above |
Publications | Label for level above |
|---|---|---|
| 238 | 19563 | BIOINFORMATICS//BMC BIOINFORMATICS//GENOME RESEARCH |
Terms with highest relevance score |
| Rank | Term | Type of term | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
|---|---|---|---|---|---|---|
| 1 | PROGRAM COMPUTAT GENOM | Address | 10 | 52% | 5% | 13 |
| 2 | CIFN | Address | 8 | 62% | 3% | 8 |
| 3 | PROGRAMA GENOM COMPUTAC | Address | 5 | 38% | 4% | 10 |
| 4 | PROGRAMA BIOL MOL COMPUTAC | Address | 4 | 67% | 2% | 4 |
| 5 | OPERON PREDICTION | Author keyword | 3 | 50% | 2% | 4 |
| 6 | CO REGULATION NETWORK | Author keyword | 1 | 50% | 1% | 2 |
| 7 | AA KHARKEVICH INFORMAT TRANSMISS PROBLEMS | Address | 1 | 13% | 3% | 9 |
| 8 | REGULATORY NETWORK INFERENCE | Author keyword | 1 | 40% | 1% | 2 |
| 9 | STREPTOCOCCACEAE | Author keyword | 1 | 33% | 1% | 2 |
| 10 | CNRSUMR 6241 | Address | 1 | 50% | 0% | 1 |
Web of Science journal categories |
Author Key Words |
| Rank | Web of Science journal category | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
LCSH search | Wikipedia search |
|---|---|---|---|---|---|---|---|
| 1 | OPERON PREDICTION | 3 | 50% | 2% | 4 | Search OPERON+PREDICTION | Search OPERON+PREDICTION |
| 2 | CO REGULATION NETWORK | 1 | 50% | 1% | 2 | Search CO+REGULATION+NETWORK | Search CO+REGULATION+NETWORK |
| 3 | REGULATORY NETWORK INFERENCE | 1 | 40% | 1% | 2 | Search REGULATORY+NETWORK+INFERENCE | Search REGULATORY+NETWORK+INFERENCE |
| 4 | STREPTOCOCCACEAE | 1 | 33% | 1% | 2 | Search STREPTOCOCCACEAE | Search STREPTOCOCCACEAE |
| 5 | COG FUNCTION | 1 | 50% | 0% | 1 | Search COG+FUNCTION | Search COG+FUNCTION |
| 6 | COLLABORATION STRUCTURE | 1 | 50% | 0% | 1 | Search COLLABORATION+STRUCTURE | Search COLLABORATION+STRUCTURE |
| 7 | CONSERVED GENE PAIR | 1 | 50% | 0% | 1 | Search CONSERVED+GENE+PAIR | Search CONSERVED+GENE+PAIR |
| 8 | EVALUATION OF PREDICTION | 1 | 50% | 0% | 1 | Search EVALUATION+OF+PREDICTION | Search EVALUATION+OF+PREDICTION |
| 9 | EXPRESSION CONSERVATION | 1 | 50% | 0% | 1 | Search EXPRESSION+CONSERVATION | Search EXPRESSION+CONSERVATION |
| 10 | GENOMIC NEIGHBORHOOD | 1 | 50% | 0% | 1 | Search GENOMIC+NEIGHBORHOOD | Search GENOMIC+NEIGHBORHOOD |
Key Words Plus |
| Rank | Web of Science journal category | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
|---|---|---|---|---|---|
| 1 | GUIDED GENETIC ALGORITHM | 6 | 80% | 2% | 4 |
| 2 | MICROBIAL GENOMES | 5 | 11% | 14% | 38 |
| 3 | SENSING MACHINERY | 4 | 75% | 1% | 3 |
| 4 | OPERON PREDICTION | 3 | 35% | 3% | 8 |
| 5 | COMPARATIVE GENOMIC RECONSTRUCTION | 2 | 67% | 1% | 2 |
| 6 | COMPARATIVE GENOMICS APPROACH | 2 | 67% | 1% | 2 |
| 7 | GRAMMATICAL MODEL | 2 | 67% | 1% | 2 |
| 8 | GAMMA PROTEOBACTERIAL GENOMES | 1 | 50% | 1% | 2 |
| 9 | ODB | 1 | 100% | 1% | 2 |
| 10 | TRANSCRIPTIONAL UNITS | 1 | 40% | 1% | 2 |
Journals |
Reviews |
| Title | Publ. year | Cit. | Active references | % act. ref. to same field |
|---|---|---|---|---|
| Structure and evolution of transcriptional regulatory networks | 2004 | 320 | 40 | 23% |
| Regulation by transcription factors in bacteria: beyond description | 2009 | 44 | 189 | 13% |
| Informatics resources for tuberculosis - Towards drug discovery | 2012 | 6 | 37 | 19% |
| Comparative genomic reconstruction of transcriptional regulatory networks in bacteria | 2007 | 51 | 259 | 8% |
| Scaling relationship in the gene content of transcriptional machinery in bacteria | 2009 | 3 | 58 | 24% |
| New insights into the regulatory networks of paralogous genes in bacteria | 2010 | 14 | 58 | 14% |
| Bioinformatics Resources for the Study of Gene Regulation in Bacteria | 2009 | 9 | 106 | 16% |
| Identifying global regulators in transcriptional regulatory networks in bacteria | 2003 | 219 | 62 | 6% |
| Transcriptional regulatory networks in bacteria: from input signals to output responses | 2006 | 52 | 39 | 8% |
| Global analysis of gene transcription regulation in prokaryotes | 2006 | 18 | 258 | 12% |
Address terms |
| Rank | Address term | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
|---|---|---|---|---|---|
| 1 | PROGRAM COMPUTAT GENOM | 10 | 52% | 4.9% | 13 |
| 2 | CIFN | 8 | 62% | 3.0% | 8 |
| 3 | PROGRAMA GENOM COMPUTAC | 5 | 38% | 3.8% | 10 |
| 4 | PROGRAMA BIOL MOL COMPUTAC | 4 | 67% | 1.5% | 4 |
| 5 | AA KHARKEVICH INFORMAT TRANSMISS PROBLEMS | 1 | 13% | 3.4% | 9 |
| 6 | CNRSUMR 6241 | 1 | 50% | 0.4% | 1 |
| 7 | COMPUTAT MODELING SIMULAT | 1 | 50% | 0.4% | 1 |
| 8 | DOE BIOENERGY SCI BESC | 1 | 50% | 0.4% | 1 |
| 9 | GENET GENOMES BACTERIENS | 1 | 50% | 0.4% | 1 |
| 10 | INFORMAT TRANSMISS PROBLEMS RAS | 1 | 50% | 0.4% | 1 |
Related classes at same level (level 1) |
| Rank | Relatedness score | Related classes |
|---|---|---|
| 1 | 0.0000198284 | MOTIF DISCOVERY//CLOSEST STRING PROBLEM//MOTIF FINDING |
| 2 | 0.0000163566 | BIDIRECTIONAL PROMOTER//NEIGHBORING GENES//GENOME NETWORK PROJECT CORE GRPGENOM SCI |
| 3 | 0.0000163257 | NETWORK INFERENCE//GENE REGULATORY NETWORKS//GENE NETWORK INFERENCE |
| 4 | 0.0000137618 | CIRCOS//BIODADOS//ORTHOLOGY |
| 5 | 0.0000132322 | TREE OF LIFE//HORIZONTAL GENE TRANSFER//LATERAL GENE TRANSFER |
| 6 | 0.0000109833 | ESSENTIAL GENES//SUBTRACTIVE GENOMICS//ESSENTIAL GENE |
| 7 | 0.0000103572 | PROTEIN INTERACTION NETWORKS//PROTEIN PROTEIN INTERACTION NETWORKS//PROTEIN PROTEIN INTERACTION NETWORK |
| 8 | 0.0000101546 | CIS MOTIF//ARABIDOPSIS T87 CELL LINE//GENE ATLAS |
| 9 | 0.0000094909 | GENOME REARRANGEMENTS//SORTING BY REVERSALS//BREAKPOINT GRAPH |
| 10 | 0.0000090692 | FLUX BALANCE ANALYSIS//CONSTRAINT BASED MODELING//METABOLIC NETWORK |