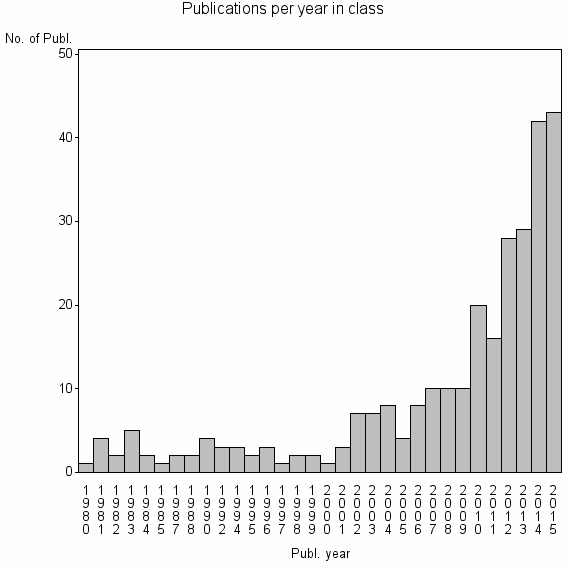
Class information for: |
Basic class information |
| ID | Publications | Average number of references |
Avg. shr. active ref. in WoS |
|---|---|---|---|
| 23630 | 285 | 44.3 | 82% |
Classes in level above (level 2) |
Terms with highest relevance score |
| Rank | Term | Type of term | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
|---|---|---|---|---|---|---|
| 1 | ALKB | Author keyword | 17 | 41% | 11% | 32 |
| 2 | METTL3 | Author keyword | 8 | 100% | 2% | 5 |
| 3 | ALKBH3 | Author keyword | 6 | 100% | 1% | 4 |
| 4 | N 6 METHYLADENOSINE | Author keyword | 5 | 50% | 2% | 7 |
| 5 | ALKBH2 | Author keyword | 4 | 75% | 1% | 3 |
| 6 | ALKBH5 | Author keyword | 4 | 75% | 1% | 3 |
| 7 | ALKBH7 | Author keyword | 3 | 100% | 1% | 3 |
| 8 | 1 N 2 ETHENOGUANINE | Author keyword | 2 | 67% | 1% | 2 |
| 9 | 3 METHYLCYTOSINE | Author keyword | 2 | 67% | 1% | 2 |
| 10 | YT521 | Author keyword | 2 | 67% | 1% | 2 |
Web of Science journal categories |
Author Key Words |
| Rank | Web of Science journal category | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
LCSH search | Wikipedia search |
|---|---|---|---|---|---|---|---|
| 1 | ALKB | 17 | 41% | 11% | 32 | Search ALKB | Search ALKB |
| 2 | METTL3 | 8 | 100% | 2% | 5 | Search METTL3 | Search METTL3 |
| 3 | ALKBH3 | 6 | 100% | 1% | 4 | Search ALKBH3 | Search ALKBH3 |
| 4 | N 6 METHYLADENOSINE | 5 | 50% | 2% | 7 | Search N+6+METHYLADENOSINE | Search N+6+METHYLADENOSINE |
| 5 | ALKBH2 | 4 | 75% | 1% | 3 | Search ALKBH2 | Search ALKBH2 |
| 6 | ALKBH5 | 4 | 75% | 1% | 3 | Search ALKBH5 | Search ALKBH5 |
| 7 | ALKBH7 | 3 | 100% | 1% | 3 | Search ALKBH7 | Search ALKBH7 |
| 8 | 1 N 2 ETHENOGUANINE | 2 | 67% | 1% | 2 | Search 1+N+2+ETHENOGUANINE | Search 1+N+2+ETHENOGUANINE |
| 9 | 3 METHYLCYTOSINE | 2 | 67% | 1% | 2 | Search 3+METHYLCYTOSINE | Search 3+METHYLCYTOSINE |
| 10 | YT521 | 2 | 67% | 1% | 2 | Search YT521 | Search YT521 |
Key Words Plus |
| Rank | Web of Science journal category | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
|---|---|---|---|---|---|
| 1 | OXIDATIVE DEMETHYLATION | 60 | 43% | 38% | 107 |
| 2 | ENZYME ALKB | 48 | 100% | 6% | 17 |
| 3 | OBESITY ASSOCIATED FTO | 42 | 64% | 14% | 41 |
| 4 | N6 METHYLADENOSINE | 41 | 81% | 9% | 25 |
| 5 | ADENOSINE RESIDUES | 30 | 100% | 4% | 12 |
| 6 | N 6 METHYLADENOSINE RESIDUES | 30 | 100% | 4% | 12 |
| 7 | ALKYLATION DAMAGE | 28 | 36% | 22% | 62 |
| 8 | 3 METHYLTHYMINE | 26 | 80% | 6% | 16 |
| 9 | POLYA RICH RNA | 18 | 89% | 3% | 8 |
| 10 | DNA ALKYLATION DAMAGE | 16 | 65% | 5% | 15 |
Journals |
Reviews |
| Title | Publ. year | Cit. | Active references | % act. ref. to same field |
|---|---|---|---|---|
| The dynamic epitranscriptome: N-6-methyladenosine and gene expression control | 2014 | 36 | 90 | 51% |
| The AlkB Family of Fe(II)/alpha-Ketoglutarate-dependent Dioxygenases: Repairing Nucleic Acid Alkylation Damage and Beyond | 2015 | 1 | 90 | 73% |
| A Non-Heme Iron-Mediated Chemical Demethylation in DNA and RNA | 2009 | 35 | 45 | 53% |
| Gene expression regulation mediated through reversible m(6)A RNA methylation | 2014 | 28 | 106 | 32% |
| N6-methyl-adenosine modification in messenger and long non-coding RNA | 2013 | 20 | 37 | 54% |
| Repair of alkylated DNA: Recent advances | 2007 | 129 | 93 | 28% |
| RNA N-6-methyladenosine methylation in post-transcriptional gene expression regulation | 2015 | 1 | 95 | 48% |
| Reversible RNA adenosine methylation in biological regulation | 2013 | 32 | 63 | 32% |
| Emerging Roles of RNA Modification: m(6)A and U-Tail | 2014 | 9 | 58 | 47% |
| Direct removal of alkylation damage from DNA by AlkB and related DNA dioxygenases | 2006 | 13 | 18 | 83% |
Address terms |
| Rank | Address term | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
|---|---|---|---|---|---|
| 1 | UBIQUITIN SIGNALLING GRP | 2 | 50% | 1.1% | 3 |
| 2 | CLIN DIAGNOST INTERVENT | 1 | 31% | 1.4% | 4 |
| 3 | BIG CAS OSLO GENOME COOPERAT | 1 | 100% | 0.7% | 2 |
| 4 | MOL CELLULAR BIOCHEM BBSRB | 1 | 100% | 0.7% | 2 |
| 5 | GENOME STRUCT STABIL GRP | 1 | 40% | 0.7% | 2 |
| 6 | GENET NETWORK BIOL | 1 | 33% | 0.7% | 2 |
| 7 | BEIJING MOL SCIMINIST EDUC CHEM | 1 | 50% | 0.4% | 1 |
| 8 | MOUSE PHENOTYPING IL | 1 | 50% | 0.4% | 1 |
| 9 | SYNTHET FUNCT BIOMOL MINIST EDUC | 1 | 50% | 0.4% | 1 |
| 10 | GENOM PRECIS MED | 1 | 20% | 1.1% | 3 |
Related classes at same level (level 1) |
| Rank | Relatedness score | Related classes |
|---|---|---|
| 1 | 0.0000230430 | FTO//FTO GENE//FAT MASS AND OBESITY ASSOCIATED GENE |
| 2 | 0.0000156887 | QUEUINE//TRNA MODIFICATION//RNA MODIFICATION |
| 3 | 0.0000134740 | 5 HYDROXYMETHYLCYTOSINE//5HMC//5 CARBOXYLCYTOSINE |
| 4 | 0.0000133888 | O 6 ALKYLGUANINE DNA ALKYLTRANSFERASE//O 6 BENZYLGUANINE//ALKYLTRANSFERASE |
| 5 | 0.0000108671 | MET BIOCATALYSIS//BIOMIMET SYST//NONHEME IRON ENZYMES |
| 6 | 0.0000098384 | ACROLEIN//ETHENO DNA ADDUCTS//ETHENO ADDUCT |
| 7 | 0.0000073524 | DNA GLYCOSYLASE//8 OXOGUANINE//BASE EXCISION REPAIR |
| 8 | 0.0000069755 | INTERACTION PROPENSITY//RNA BINDING RESIDUES//COMPUTAT INTELLIGENCE LEARNING DISCOVERY |
| 9 | 0.0000043485 | CANCER AND AIDS//CPG ISLETS//DNASE SENSITIVE CHROMATIN |
| 10 | 0.0000040880 | IME1//IME2//PROSPORE MEMBRANE |