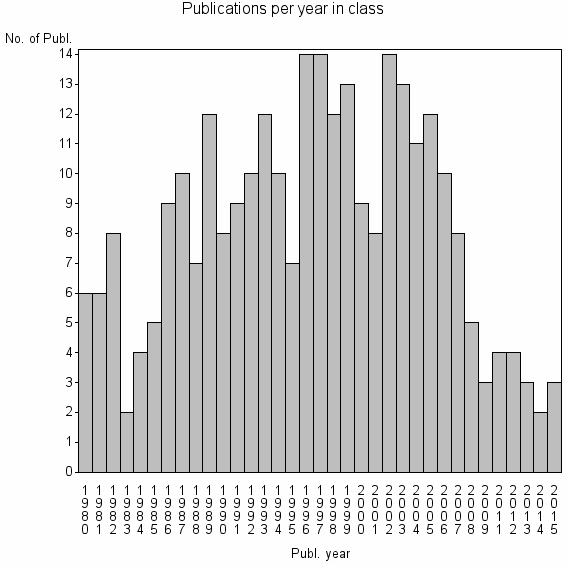
Class information for: |
Basic class information |
| ID | Publications | Average number of references |
Avg. shr. active ref. in WoS |
|---|---|---|---|
| 23550 | 287 | 33.2 | 55% |
Classes in level above (level 2) |
| ID, lev. above |
Publications | Label for level above |
|---|---|---|
| 663 | 12629 | CHLOROPEROXIDASE//CYTOCHROME C3//PEROXIDASE |
Terms with highest relevance score |
| Rank | Term | Type of term | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
|---|---|---|---|---|---|---|
| 1 | HEME PEPTIDE | Author keyword | 17 | 100% | 3% | 8 |
| 2 | MICROPEROXIDASE 11 | Author keyword | 15 | 45% | 9% | 25 |
| 3 | MICROPEROXIDASE 8 | Author keyword | 14 | 65% | 5% | 13 |
| 4 | MICROPEROXIDASE | Author keyword | 10 | 37% | 8% | 23 |
| 5 | MICROPEROXIDASES | Author keyword | 6 | 71% | 2% | 5 |
| 6 | HEME PEPTIDES | Author keyword | 2 | 67% | 1% | 2 |
| 7 | HEMEPEPTIDE | Author keyword | 2 | 67% | 1% | 2 |
| 8 | MITIGATIVE EFFECT | Author keyword | 2 | 67% | 1% | 2 |
| 9 | CYTOCHROME C HEME PEPTIDE | Author keyword | 1 | 100% | 1% | 2 |
| 10 | INT EXCELLENCE INTERDISCIPLINARY SCI | Address | 1 | 50% | 1% | 2 |
Web of Science journal categories |
Author Key Words |
| Rank | Web of Science journal category | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
LCSH search | Wikipedia search |
|---|---|---|---|---|---|---|---|
| 1 | HEME PEPTIDE | 17 | 100% | 3% | 8 | Search HEME+PEPTIDE | Search HEME+PEPTIDE |
| 2 | MICROPEROXIDASE 11 | 15 | 45% | 9% | 25 | Search MICROPEROXIDASE+11 | Search MICROPEROXIDASE+11 |
| 3 | MICROPEROXIDASE 8 | 14 | 65% | 5% | 13 | Search MICROPEROXIDASE+8 | Search MICROPEROXIDASE+8 |
| 4 | MICROPEROXIDASE | 10 | 37% | 8% | 23 | Search MICROPEROXIDASE | Search MICROPEROXIDASE |
| 5 | MICROPEROXIDASES | 6 | 71% | 2% | 5 | Search MICROPEROXIDASES | Search MICROPEROXIDASES |
| 6 | HEME PEPTIDES | 2 | 67% | 1% | 2 | Search HEME+PEPTIDES | Search HEME+PEPTIDES |
| 7 | HEMEPEPTIDE | 2 | 67% | 1% | 2 | Search HEMEPEPTIDE | Search HEMEPEPTIDE |
| 8 | MITIGATIVE EFFECT | 2 | 67% | 1% | 2 | Search MITIGATIVE+EFFECT | Search MITIGATIVE+EFFECT |
| 9 | CYTOCHROME C HEME PEPTIDE | 1 | 100% | 1% | 2 | Search CYTOCHROME+C+HEME+PEPTIDE | Search CYTOCHROME+C+HEME+PEPTIDE |
| 10 | MNIII MICROPEROXIDASE 8 | 1 | 100% | 1% | 2 | Search MNIII+MICROPEROXIDASE+8 | Search MNIII+MICROPEROXIDASE+8 |
Key Words Plus |
| Rank | Web of Science journal category | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
|---|---|---|---|---|---|
| 1 | HEME OCTAPEPTIDE MICROPEROXIDASE 8 | 55 | 92% | 8% | 22 |
| 2 | OCTAPEPTIDE MICROPEROXIDASE 8 | 30 | 79% | 7% | 19 |
| 3 | PEROXIDASE MODEL ELECTRODES | 13 | 80% | 3% | 8 |
| 4 | MP 8 | 11 | 100% | 2% | 6 |
| 5 | CYTOCHROME C MICROPEROXIDASE | 9 | 83% | 2% | 5 |
| 6 | UNDECAPEPTIDE | 9 | 59% | 3% | 10 |
| 7 | HEMOPROTEINS | 9 | 15% | 18% | 51 |
| 8 | PROTON COUPLED REDUCTION | 8 | 100% | 2% | 5 |
| 9 | OCTAPEPTIDE MICROPEROXIDASE 8 MP 8 | 6 | 71% | 2% | 5 |
| 10 | HEME UNDECAPEPTIDE | 5 | 63% | 2% | 5 |
Journals |
Reviews |
| Title | Publ. year | Cit. | Active references | % act. ref. to same field |
|---|---|---|---|---|
| Insights into porphyrin chemistry provided by the microperoxidases, the haempeptides derived from cytochrome c | 2007 | 32 | 122 | 61% |
| Heme-peptide models for hemoproteins .1. Solution chemistry of N-acetylmicroperoxidase-8 | 1996 | 66 | 64 | 42% |
Address terms |
| Rank | Address term | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
|---|---|---|---|---|---|
| 1 | INT EXCELLENCE INTERDISCIPLINARY SCI | 1 | 50% | 0.7% | 2 |
| 2 | CNRS CHIM MOL MAT ORSAY | 1 | 50% | 0.3% | 1 |
| 3 | CNRSFRE 2127 | 1 | 50% | 0.3% | 1 |
| 4 | CRISTALLOCHEM CHIM STRUCT | 1 | 50% | 0.3% | 1 |
| 5 | FRE 2127 CHIM BIOORGAN BIOINORGAN | 1 | 50% | 0.3% | 1 |
| 6 | TECH ENGN CHEM | 1 | 50% | 0.3% | 1 |
| 7 | CNRSUMR 8124 | 0 | 33% | 0.3% | 1 |
| 8 | GUIDO DONEGANI | 0 | 25% | 0.3% | 1 |
| 9 | URA 2096SERV BIOENERGET | 0 | 25% | 0.3% | 1 |
| 10 | ECBB | 0 | 11% | 0.3% | 1 |
Related classes at same level (level 1) |
| Rank | Relatedness score | Related classes |
|---|---|---|
| 1 | 0.0000206617 | ALKALINE TRANSITION//ALKALINE ISOMERIZATION//CYTOCHROME C2 |
| 2 | 0.0000139485 | CYTOCHROME C PEROXIDASE//DEHALOPEROXIDASE//ARTHROMYCES RAMOSUS |
| 3 | 0.0000138709 | MAT THEORY COMPUTAT//NON PLANAR PORPHYRINS//UA 040804 |
| 4 | 0.0000126821 | PL BIOENGN PROGRAM//SURFACE ENHANCED RESONANCE RAMAN SPECTROSCOPY//AZURIN |
| 5 | 0.0000096559 | SCI RADIAT MED BURNS//SCI RADIAT BURNS MED//355 NM PULSE LASER |
| 6 | 0.0000088751 | PROTEIN DESIGN//FOUR HELIX BUNDLE//TEMPLATE ASSEMBLED SYNTHETIC PROTEIN |
| 7 | 0.0000084639 | INVEST MOL STRUCT//ANIONOTROPIC REARRANGEMENTS//RD ORGAN PHARMACEUT CHEM |
| 8 | 0.0000079907 | DICRANOPTERIS DICHOTOMA//RARE EARTH ELEMENTS REES//SOIL AGRICHEM |
| 9 | 0.0000079260 | ALBUMIN HEME//POLYMER CHEM SHINJUKU KU//CHIRAL ZINC PORPHYRIN |
| 10 | 0.0000072784 | COMPOUND I//LISE MEITNER MINERVA COMPUTAT QUANTUM CHEM//CPD I |