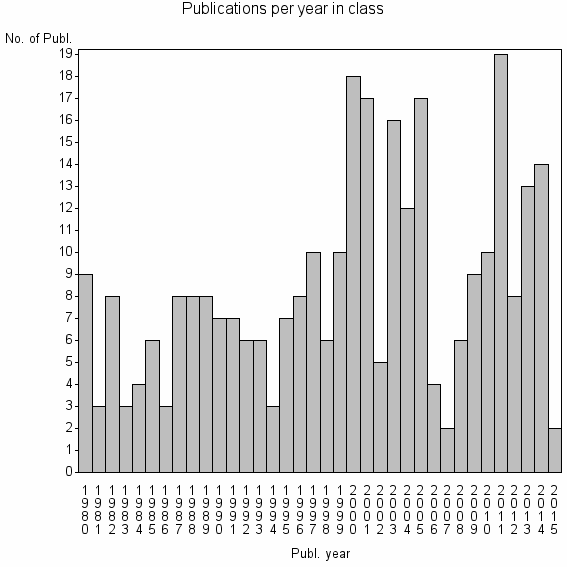
Class information for: |
Basic class information |
| ID | Publications | Average number of references |
Avg. shr. active ref. in WoS |
|---|---|---|---|
| 23104 | 302 | 33.8 | 66% |
Classes in level above (level 2) |
| ID, lev. above |
Publications | Label for level above |
|---|---|---|
| 2342 | 3993 | CORYNEBACTERIUM GLUTAMICUM//SHIKIMATE PATHWAY//SHIKIMATE KINASE |
Terms with highest relevance score |
| Rank | Term | Type of term | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
|---|---|---|---|---|---|---|
| 1 | KDO8P SYNTHASE | Author keyword | 20 | 100% | 3% | 9 |
| 2 | 3 DEOXY D ARABINO HEPTULOSONATE 7 PHOSPHATE SYNTHASE | Author keyword | 10 | 58% | 4% | 11 |
| 3 | KDO8PS | Author keyword | 6 | 100% | 1% | 4 |
| 4 | NMR INTERACTION STUDIES | Author keyword | 6 | 100% | 1% | 4 |
| 5 | DAHPS | Author keyword | 5 | 60% | 2% | 6 |
| 6 | KDO BIOSYNTHESIS | Author keyword | 4 | 75% | 1% | 3 |
| 7 | DAH7P SYNTHASE | Author keyword | 3 | 100% | 1% | 3 |
| 8 | DAH7PS | Author keyword | 3 | 100% | 1% | 3 |
| 9 | KDO8P | Author keyword | 3 | 100% | 1% | 3 |
| 10 | KDOP SYNTHASE | Author keyword | 3 | 100% | 1% | 3 |
Web of Science journal categories |
Author Key Words |
| Rank | Web of Science journal category | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
LCSH search | Wikipedia search |
|---|---|---|---|---|---|---|---|
| 1 | KDO8P SYNTHASE | 20 | 100% | 3% | 9 | Search KDO8P+SYNTHASE | Search KDO8P+SYNTHASE |
| 2 | 3 DEOXY D ARABINO HEPTULOSONATE 7 PHOSPHATE SYNTHASE | 10 | 58% | 4% | 11 | Search 3+DEOXY+D+ARABINO+HEPTULOSONATE+7+PHOSPHATE+SYNTHASE | Search 3+DEOXY+D+ARABINO+HEPTULOSONATE+7+PHOSPHATE+SYNTHASE |
| 3 | KDO8PS | 6 | 100% | 1% | 4 | Search KDO8PS | Search KDO8PS |
| 4 | NMR INTERACTION STUDIES | 6 | 100% | 1% | 4 | Search NMR+INTERACTION+STUDIES | Search NMR+INTERACTION+STUDIES |
| 5 | DAHPS | 5 | 60% | 2% | 6 | Search DAHPS | Search DAHPS |
| 6 | KDO BIOSYNTHESIS | 4 | 75% | 1% | 3 | Search KDO+BIOSYNTHESIS | Search KDO+BIOSYNTHESIS |
| 7 | DAH7P SYNTHASE | 3 | 100% | 1% | 3 | Search DAH7P+SYNTHASE | Search DAH7P+SYNTHASE |
| 8 | DAH7PS | 3 | 100% | 1% | 3 | Search DAH7PS | Search DAH7PS |
| 9 | KDO8P | 3 | 100% | 1% | 3 | Search KDO8P | Search KDO8P |
| 10 | KDOP SYNTHASE | 3 | 100% | 1% | 3 | Search KDOP+SYNTHASE | Search KDOP+SYNTHASE |
Key Words Plus |
| Rank | Web of Science journal category | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
|---|---|---|---|---|---|
| 1 | 3 DEOXY D MANNO OCTULOSONATE 8 PHOSPHATE SYNTHASE | 76 | 85% | 13% | 40 |
| 2 | KDO8P SYNTHASE | 59 | 84% | 11% | 32 |
| 3 | 3 DEOXY D MANNO 2 OCTULOSONATE 8 PHOSPHATE SYNTHASE | 56 | 100% | 6% | 19 |
| 4 | PUTATIVE REACTION INTERMEDIATE | 55 | 92% | 7% | 22 |
| 5 | 7 PHOSPHATE SYNTHASE | 30 | 84% | 5% | 16 |
| 6 | AEOLICUS | 24 | 82% | 5% | 14 |
| 7 | SUBSTRATE AMBIGUITY | 24 | 91% | 3% | 10 |
| 8 | 8 PHOSPHATE SYNTHASE | 23 | 86% | 4% | 12 |
| 9 | ACID 8 PHOSPHATE SYNTHASE | 23 | 100% | 3% | 10 |
| 10 | 7 PHOSPHATE SYNTHETASE | 15 | 82% | 3% | 9 |
Journals |
Reviews |
| Title | Publ. year | Cit. | Active references | % act. ref. to same field |
|---|---|---|---|---|
| The diversity of allosteric controls at the gateway to aromatic amino acid biosynthesis | 2013 | 6 | 32 | 81% |
| New Targets for Antibacterial Design: Kdo Biosynthesis and LPS Machinery Transport to the Cell Surface | 2011 | 15 | 156 | 37% |
| Kdo: a critical monosaccharide for bacteria viability | 2010 | 18 | 138 | 41% |
| Recent Approaches to Novel Antibacterials Designed After LPS Structure and Biochemistry | 2012 | 5 | 136 | 22% |
| Re LPS biogenetic pathway: Enzyme characterisation and synthetic efforts towards inhibitors | 2008 | 1 | 143 | 40% |
| Conservation of the 2-keto-3-deoxymanno-octulosonic acid (Kdo) biosynthesis pathway between plants and bacteria | 2013 | 0 | 86 | 43% |
| SYNTHESIS AND USE OF 3-DEOXY-D-MANNO-2-OCTULOSONATE (KDO) IN ESCHERICHIA-COLI - POTENTIAL SITES OF INHIBITION | 1983 | 56 | 9 | 78% |
| Detection of novel enzyme intermediates in PEP-utilizing enzymes | 2005 | 5 | 63 | 22% |
| BIOGENESIS OF THE OUTER-MEMBRANE OF SALMONELLA | 1983 | 0 | 4 | 50% |
Address terms |
| Rank | Address term | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
|---|---|---|---|---|---|
| 1 | MED CHEM CHEM | 2 | 43% | 1.0% | 3 |
| 2 | INTERDEPARTMENTAL PROGRAM MED CHEM | 2 | 21% | 2.3% | 7 |
| 3 | INFECT DISCOVERY CANC INFECT AREA | 1 | 50% | 0.3% | 1 |
| 4 | GEN BOT BOT GARDEN | 0 | 33% | 0.3% | 1 |
| 5 | HEINRICH HEINE UNIV DUSSELDORF | 0 | 33% | 0.3% | 1 |
| 6 | CANC THER EUT CRC | 0 | 25% | 0.3% | 1 |
| 7 | INCM FR 1742 | 0 | 25% | 0.3% | 1 |
| 8 | INRA BORDEAUX AQUITAINE | 0 | 25% | 0.3% | 1 |
| 9 | PESQUISA DESENVOLVIMENTO BIOL MOL FUNC | 0 | 25% | 0.3% | 1 |
| 10 | UMR 7565GEVSM | 0 | 25% | 0.3% | 1 |
Related classes at same level (level 1) |
| Rank | Relatedness score | Related classes |
|---|---|---|
| 1 | 0.0000237768 | LPXC//LPXC INHIBITORS//LPXA |
| 2 | 0.0000219943 | SHIKIMATE KINASE//SHIKIMATE PATHWAY//CHORISMATE SYNTHASE |
| 3 | 0.0000219806 | ANTHRANILATE SYNTHASE//CHORISMATE MUTASE//PREPHENATE DEHYDROGENASE |
| 4 | 0.0000214176 | TYRR//SHIKIMIC ACID PRODUCTION//CIS CIS MUCONIC ACID |
| 5 | 0.0000183332 | N ACETYLNEURAMINATE LYASE//N ACETYLNEURAMINIC ACID//KDN |
| 6 | 0.0000176611 | SIALOBIOL//COLANIC ACID//CMP SIALIC ACID SYNTHETASE |
| 7 | 0.0000087200 | LIPID A ANALOG//LIPID A//LPS ANTAGONIST |
| 8 | 0.0000086395 | UDP SUGAR HYDROLASE//PHOSPHOHEXOMUTASE//HAD SUPERFAMILY |
| 9 | 0.0000060621 | GLYPHOSATE RESISTANCE//5 ENOLPYRUVYLSHIKIMATE 3 PHOSPHATE SYNTHASE//GLYPHOSATE |
| 10 | 0.0000059294 | ALANINE RACEMASE//MURD//GLUTAMATE RACEMASE |