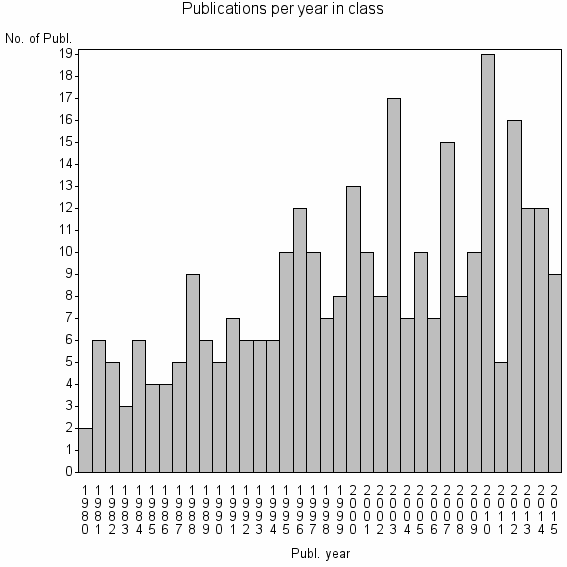
Class information for: |
Basic class information |
| ID | Publications | Average number of references |
Avg. shr. active ref. in WoS |
|---|---|---|---|
| 23004 | 305 | 35.8 | 69% |
Classes in level above (level 2) |
| ID, lev. above |
Publications | Label for level above |
|---|---|---|
| 134 | 22710 | SACCHAROMYCES CEREVISIAE//YEAST//SCHIZOSACCHAROMYCES POMBE |
Terms with highest relevance score |
| Rank | Term | Type of term | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
|---|---|---|---|---|---|---|
| 1 | PACC | Author keyword | 7 | 43% | 4% | 13 |
| 2 | PI SENSING | Author keyword | 4 | 75% | 1% | 3 |
| 3 | HUMICOLA LUTEA | Author keyword | 3 | 40% | 2% | 6 |
| 4 | PACC GENE | Author keyword | 2 | 67% | 1% | 2 |
| 5 | RIM101 PATHWAY | Author keyword | 2 | 67% | 1% | 2 |
| 6 | PHOSPHATE SENSING | Author keyword | 2 | 40% | 1% | 4 |
| 7 | ALKALINE PROTEASE ALP | Author keyword | 1 | 100% | 1% | 2 |
| 8 | ENZYME EXCRETION | Author keyword | 1 | 100% | 1% | 2 |
| 9 | PAL SIGNALING PATHWAY | Author keyword | 1 | 100% | 1% | 2 |
| 10 | RIM PATHWAY | Author keyword | 1 | 100% | 1% | 2 |
Web of Science journal categories |
Author Key Words |
| Rank | Web of Science journal category | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
LCSH search | Wikipedia search |
|---|---|---|---|---|---|---|---|
| 1 | PACC | 7 | 43% | 4% | 13 | Search PACC | Search PACC |
| 2 | PI SENSING | 4 | 75% | 1% | 3 | Search PI+SENSING | Search PI+SENSING |
| 3 | HUMICOLA LUTEA | 3 | 40% | 2% | 6 | Search HUMICOLA+LUTEA | Search HUMICOLA+LUTEA |
| 4 | PACC GENE | 2 | 67% | 1% | 2 | Search PACC+GENE | Search PACC+GENE |
| 5 | RIM101 PATHWAY | 2 | 67% | 1% | 2 | Search RIM101+PATHWAY | Search RIM101+PATHWAY |
| 6 | PHOSPHATE SENSING | 2 | 40% | 1% | 4 | Search PHOSPHATE+SENSING | Search PHOSPHATE+SENSING |
| 7 | ALKALINE PROTEASE ALP | 1 | 100% | 1% | 2 | Search ALKALINE+PROTEASE+ALP | Search ALKALINE+PROTEASE+ALP |
| 8 | ENZYME EXCRETION | 1 | 100% | 1% | 2 | Search ENZYME+EXCRETION | Search ENZYME+EXCRETION |
| 9 | PAL SIGNALING PATHWAY | 1 | 100% | 1% | 2 | Search PAL+SIGNALING+PATHWAY | Search PAL+SIGNALING+PATHWAY |
| 10 | RIM PATHWAY | 1 | 100% | 1% | 2 | Search RIM+PATHWAY | Search RIM+PATHWAY |
Key Words Plus |
| Rank | Web of Science journal category | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
|---|---|---|---|---|---|
| 1 | AMBIENT PH | 43 | 36% | 31% | 94 |
| 2 | PECTATE LYASE SECRETION | 15 | 73% | 4% | 11 |
| 3 | RIM101 PATHWAY | 12 | 47% | 6% | 18 |
| 4 | PACC TRANSCRIPTION FACTOR | 9 | 50% | 4% | 13 |
| 5 | ENVIRONMENTAL PH | 6 | 27% | 7% | 20 |
| 6 | AMBIENT PH REGULATION | 6 | 100% | 1% | 4 |
| 7 | PH SIGNAL TRANSDUCTION | 5 | 63% | 2% | 5 |
| 8 | PACC | 5 | 50% | 2% | 7 |
| 9 | TRANSCRIPTION FACTOR PACC | 5 | 44% | 3% | 8 |
| 10 | RIM20 LOCALIZATION | 4 | 75% | 1% | 3 |
Journals |
Reviews |
| Title | Publ. year | Cit. | Active references | % act. ref. to same field |
|---|---|---|---|---|
| Ambient pH gene regulation in fungi: making connections | 2008 | 118 | 71 | 41% |
| The signaling mechanism of ambient pH sensing and adaptation in yeast and fungi | 2012 | 13 | 39 | 54% |
| PH regulation of gene expression in fungi | 2000 | 101 | 47 | 45% |
| How human pathogenic fungi sense and adapt to pH: the link to virulence | 2009 | 44 | 48 | 29% |
| MEASUREMENT OF ENZYME ACTIVITY | 2009 | 1 | 1 | 100% |
| Liaison alcaline: Pals entice non-endosomal ESCRTs to the plasma membrane for pH signaling | 2014 | 1 | 66 | 48% |
| Recent advances in the characterization of ambient pH regulation of gene expression in filamentour fungi and yeasts | 2004 | 98 | 106 | 33% |
| Quiescent and Necrotrophic Lifestyle Choice During Postharvest Disease Development | 2013 | 9 | 147 | 21% |
| Virulence Regulation of Phytopathogenic Fungi by pH | 2013 | 5 | 144 | 31% |
| Regulation of gene expression by ambient pH in filamentous fungi and yeasts | 2002 | 150 | 121 | 26% |
Address terms |
| Rank | Address term | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
|---|---|---|---|---|---|
| 1 | PL BIOL ENGN | 1 | 33% | 0.7% | 2 |
| 2 | CBAL | 1 | 50% | 0.3% | 1 |
| 3 | CNRSUMR2585 | 1 | 50% | 0.3% | 1 |
| 4 | INTEGRAT SYST BIOL CISBIC | 1 | 50% | 0.3% | 1 |
| 5 | POSTHARVEST SCI F H FRUIT | 1 | 50% | 0.3% | 1 |
| 6 | UNIDAD MANIPULAC GENET | 1 | 50% | 0.3% | 1 |
| 7 | EGIUM NAT SCI | 1 | 29% | 0.7% | 2 |
| 8 | CNRS URA 1925 | 0 | 25% | 0.3% | 1 |
| 9 | CNRS URA1925 | 0 | 25% | 0.3% | 1 |
| 10 | LNVEST BIOL | 0 | 25% | 0.3% | 1 |
Related classes at same level (level 1) |
| Rank | Relatedness score | Related classes |
|---|---|---|
| 1 | 0.0000282796 | P I TRANSPORTER//PHO81//PHO89 |
| 2 | 0.0000187811 | ESCRT//ALG 2//TSG101 |
| 3 | 0.0000156268 | NITROGEN METABOLITE REPRESSION//NMRA//FACB |
| 4 | 0.0000137931 | GAMMA DECALACTONE//YARROWIA LIPOLYTICA//ALKALINE EXTRACELLULAR PROTEASE |
| 5 | 0.0000129468 | CCH1//NALCN//CRZ1 |
| 6 | 0.0000097279 | XLNR//AMYR//TRICHODERMA REESEI |
| 7 | 0.0000086256 | CANDIDA ALBICANS//ABERDEEN FUNGAL GRP//USC2019 |
| 8 | 0.0000065631 | IME1//IME2//PROSPORE MEMBRANE |
| 9 | 0.0000064206 | POLYGALACTURONASE//EXO POLYGALACTURONASE//PECTIN LYASE |
| 10 | 0.0000063614 | THERMOSTABLE ALKALINE PHOSPHATASE//CALF INTESTINAL ALKALINE PHOSPHATASE//E COLI ALKALINE PHOSPHATASE |