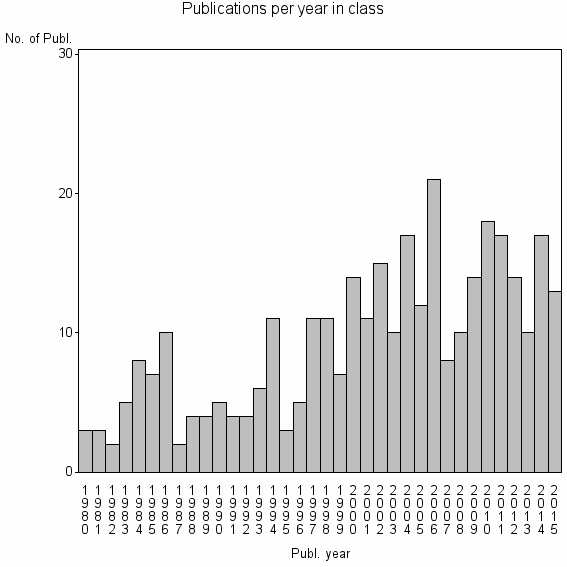
Class information for: |
Basic class information |
| ID | Publications | Average number of references |
Avg. shr. active ref. in WoS |
|---|---|---|---|
| 22111 | 336 | 34.8 | 70% |
Classes in level above (level 2) |
| ID, lev. above |
Publications | Label for level above |
|---|---|---|
| 2915 | 2265 | GALACTOSEMIA//GALACTOSE 1 PHOSPHATE URIDYLTRANSFERASE//GALACTOSE 1 PHOSPHATE |
Terms with highest relevance score |
| Rank | Term | Type of term | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
|---|---|---|---|---|---|---|
| 1 | UDP SUGAR HYDROLASE | Author keyword | 12 | 86% | 2% | 6 |
| 2 | PHOSPHOHEXOMUTASE | Author keyword | 9 | 83% | 1% | 5 |
| 3 | HAD SUPERFAMILY | Author keyword | 4 | 42% | 2% | 8 |
| 4 | BACTERIAL PHOSPHATASE | Author keyword | 4 | 75% | 1% | 3 |
| 5 | CLASS B PHOSPHATASE | Author keyword | 3 | 100% | 1% | 3 |
| 6 | CLASS C ACID PHOSPHATASE | Author keyword | 3 | 100% | 1% | 3 |
| 7 | PHOSPHOGLUCOMUTASE | Author keyword | 3 | 16% | 6% | 19 |
| 8 | PHOSPHOSERINE PHOSPHATASE | Author keyword | 2 | 33% | 2% | 6 |
| 9 | BIOL MOL CELULAR PLANTAS EDUARDO PRIMO YUF | Address | 2 | 67% | 1% | 2 |
| 10 | HAD FAMILY | Author keyword | 2 | 67% | 1% | 2 |
Web of Science journal categories |
Author Key Words |
| Rank | Web of Science journal category | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
LCSH search | Wikipedia search |
|---|---|---|---|---|---|---|---|
| 1 | UDP SUGAR HYDROLASE | 12 | 86% | 2% | 6 | Search UDP+SUGAR+HYDROLASE | Search UDP+SUGAR+HYDROLASE |
| 2 | PHOSPHOHEXOMUTASE | 9 | 83% | 1% | 5 | Search PHOSPHOHEXOMUTASE | Search PHOSPHOHEXOMUTASE |
| 3 | HAD SUPERFAMILY | 4 | 42% | 2% | 8 | Search HAD+SUPERFAMILY | Search HAD+SUPERFAMILY |
| 4 | BACTERIAL PHOSPHATASE | 4 | 75% | 1% | 3 | Search BACTERIAL+PHOSPHATASE | Search BACTERIAL+PHOSPHATASE |
| 5 | CLASS B PHOSPHATASE | 3 | 100% | 1% | 3 | Search CLASS+B+PHOSPHATASE | Search CLASS+B+PHOSPHATASE |
| 6 | CLASS C ACID PHOSPHATASE | 3 | 100% | 1% | 3 | Search CLASS+C+ACID+PHOSPHATASE | Search CLASS+C+ACID+PHOSPHATASE |
| 7 | PHOSPHOGLUCOMUTASE | 3 | 16% | 6% | 19 | Search PHOSPHOGLUCOMUTASE | Search PHOSPHOGLUCOMUTASE |
| 8 | PHOSPHOSERINE PHOSPHATASE | 2 | 33% | 2% | 6 | Search PHOSPHOSERINE+PHOSPHATASE | Search PHOSPHOSERINE+PHOSPHATASE |
| 9 | HAD FAMILY | 2 | 67% | 1% | 2 | Search HAD+FAMILY | Search HAD+FAMILY |
| 10 | PHOSPHOMANNOMUTASE | 2 | 19% | 3% | 10 | Search PHOSPHOMANNOMUTASE | Search PHOSPHOMANNOMUTASE |
Key Words Plus |
| Rank | Web of Science journal category | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
|---|---|---|---|---|---|
| 1 | PMM PGM | 26 | 100% | 3% | 11 |
| 2 | BETA PHOSPHOGLUCOMUTASE | 24 | 68% | 6% | 21 |
| 3 | E P4 | 18 | 83% | 3% | 10 |
| 4 | D PHOSPHOHEXOMUTASE SUPERFAMILY | 17 | 100% | 2% | 8 |
| 5 | PSEUDOMONAS AERUGINOSA PHOSPHOMANNOMUTASE PHOSPHOGLUCOMUTASE | 15 | 88% | 2% | 7 |
| 6 | PO3 TRANSFER | 14 | 100% | 2% | 7 |
| 7 | PHOSPHOMANNOMUTASE PHOSPHOGLUCOMUTASE | 11 | 78% | 2% | 7 |
| 8 | HAD SUPERFAMILY | 10 | 57% | 4% | 12 |
| 9 | BETA PHOSPHOGLUCOMUTASE CATALYSIS | 9 | 83% | 1% | 5 |
| 10 | PHOSPHOSERINE PHOSPHATASE | 8 | 31% | 7% | 22 |
Journals |
Reviews |
| Title | Publ. year | Cit. | Active references | % act. ref. to same field |
|---|---|---|---|---|
| Mutations in hereditary phosphoglucomutase 1 deficiency map to key regions of enzyme structure and function | 2015 | 1 | 30 | 50% |
| Enzyme Promiscuity: Engine of Evolutionary Innovation | 2014 | 10 | 66 | 23% |
| Phosphoryl group transfer: evolution of a catalytic scaffold | 2004 | 153 | 55 | 31% |
| Human HAD phosphatases: structure, mechanism, and roles in health and disease | 2013 | 24 | 110 | 14% |
| Evolutionary genomics of the HAD superfamily: Understanding the structural adaptations and catalytic diversity in a superfamily of phosphoesterases and allied enzymes | 2006 | 140 | 178 | 13% |
| Focus on phosphoaspartate and phosphoglutamate | 2011 | 6 | 56 | 18% |
| Why did Nature select phosphate for its dominant roles in biology? | 2010 | 40 | 30 | 7% |
| Reflections on biocatalysis involving phosphorus | 2012 | 1 | 57 | 19% |
| Evidence for phosphotransferases phosphorylated on aspartate residue in N-terminal DXDX(T/V) motif | 2002 | 12 | 28 | 21% |
| Metallic Fluoride Complexes as Phosphate Analogues for Structural and Mechanistic Studies of Phosphoryl Group Transfer Enzymes | 2010 | 3 | 118 | 15% |
Address terms |
| Rank | Address term | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
|---|---|---|---|---|---|
| 1 | BIOL MOL CELULAR PLANTAS EDUARDO PRIMO YUF | 2 | 67% | 0.6% | 2 |
| 2 | EP1088 | 1 | 50% | 0.6% | 2 |
| 3 | BIO MED RRC | 1 | 50% | 0.3% | 1 |
| 4 | COMP STRUCT SCI | 1 | 50% | 0.3% | 1 |
| 5 | HAS ELTE | 1 | 50% | 0.3% | 1 |
| 6 | LIFE SCI CLONING TEAM | 1 | 50% | 0.3% | 1 |
| 7 | ANALYT CHEM MED PHYSICOCHEM | 0 | 13% | 0.9% | 3 |
| 8 | ENVELOPPES BACTERIENNES | 0 | 33% | 0.3% | 1 |
| 9 | KAMAKURA | 0 | 18% | 0.6% | 2 |
| 10 | STRUCT BIOL CORE | 0 | 17% | 0.6% | 2 |
Related classes at same level (level 1) |
| Rank | Relatedness score | Related classes |
|---|---|---|
| 1 | 0.0000177718 | PHOSPHORYLCHOLINE PHOSPHATASE//SPHINGOMYELINASE C//SYMBIOT SYST SCI TECHNOL |
| 2 | 0.0000144609 | CELL MOL BIOL ICM//METAPHOSPHONATE//SULFATE TRANSFER |
| 3 | 0.0000134622 | SERINE BIOSYNTHESIS//PHOSPHORYLATED PATHWAY//3 PHOSPHOGLYCERATE DEHYDROGENASE |
| 4 | 0.0000134010 | PHOSPHOGLYCERATE MUTASE//BISPHOSPHOGLYCERATE MUTASE//2 3 BISPHOSPHOGLYCERATE MUTASE |
| 5 | 0.0000105083 | HISTIDINE BIOSYNTHESIS//IMIDAZOLEGLYCEROL PHOSPHATE DEHYDRATASE//HISTIDINOL DEHYDROGENASE |
| 6 | 0.0000100505 | CONGENITAL DISORDERS OF GLYCOSYLATION//CARBOHYDRATE DEFICIENT GLYCOPROTEIN SYNDROME//CDG |
| 7 | 0.0000096546 | PHO REGULON//C P LYASE//CARBON PHOSPHORUS BOND |
| 8 | 0.0000096455 | MANDELATE RACEMASE//ENOLASE SUPERFAMILY//MANDELATE RACEMASE EC 5122 |
| 9 | 0.0000090102 | HALOALKANE DEHALOGENASE//DEHALOGENASE//LO MIDT S |
| 10 | 0.0000087831 | CN II//PYRIMIDINE 5 NUCLEOTIDASE//5 NUCLEOTIDASES |