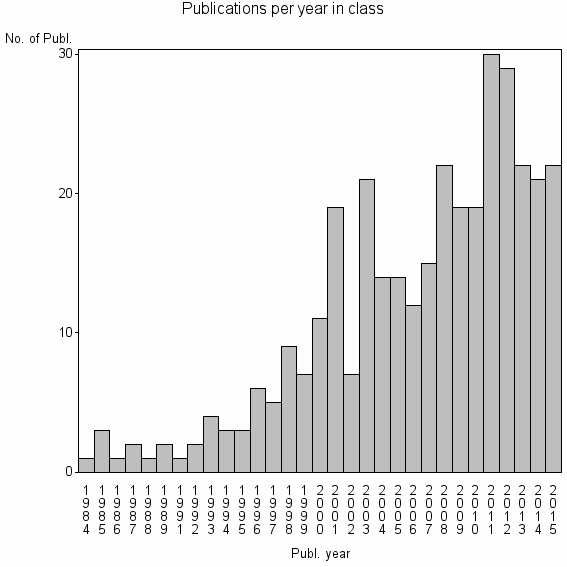
Class information for: |
Basic class information |
| ID | Publications | Average number of references |
Avg. shr. active ref. in WoS |
|---|---|---|---|
| 21821 | 347 | 34.9 | 85% |
Classes in level above (level 2) |
| ID, lev. above |
Publications | Label for level above |
|---|---|---|
| 1336 | 7916 | MARINER//SLEEPING BEAUTY//TRANSGENIC CHICKEN |
Terms with highest relevance score |
| Rank | Term | Type of term | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
|---|---|---|---|---|---|---|
| 1 | RECOMBINEERING | Author keyword | 21 | 43% | 11% | 38 |
| 2 | LAMBDA RED | Author keyword | 12 | 75% | 3% | 9 |
| 3 | LAMBDA RED RECOMBINATION | Author keyword | 6 | 71% | 1% | 5 |
| 4 | MOL CONTROL GENET SECT | Address | 4 | 40% | 2% | 8 |
| 5 | P1 TRANSDUCTION | Author keyword | 3 | 60% | 1% | 3 |
| 6 | GENE REGULAT CHROMOSOME BIOL | Address | 3 | 11% | 6% | 22 |
| 7 | BIOINNOVAT ZENTRUM | Address | 3 | 22% | 3% | 10 |
| 8 | AJINOMOTO GENET | Address | 2 | 67% | 1% | 2 |
| 9 | BAC ENGINEERING | Author keyword | 2 | 67% | 1% | 2 |
| 10 | REGENERAT THER Y D DEN | Address | 2 | 67% | 1% | 2 |
Web of Science journal categories |
Author Key Words |
| Rank | Web of Science journal category | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
LCSH search | Wikipedia search |
|---|---|---|---|---|---|---|---|
| 1 | RECOMBINEERING | 21 | 43% | 11% | 38 | Search RECOMBINEERING | Search RECOMBINEERING |
| 2 | LAMBDA RED | 12 | 75% | 3% | 9 | Search LAMBDA+RED | Search LAMBDA+RED |
| 3 | LAMBDA RED RECOMBINATION | 6 | 71% | 1% | 5 | Search LAMBDA+RED+RECOMBINATION | Search LAMBDA+RED+RECOMBINATION |
| 4 | P1 TRANSDUCTION | 3 | 60% | 1% | 3 | Search P1+TRANSDUCTION | Search P1+TRANSDUCTION |
| 5 | BAC ENGINEERING | 2 | 67% | 1% | 2 | Search BAC+ENGINEERING | Search BAC+ENGINEERING |
| 6 | RECT | 2 | 50% | 1% | 3 | Search RECT | Search RECT |
| 7 | RED RECOMBINASE | 2 | 50% | 1% | 3 | Search RED+RECOMBINASE | Search RED+RECOMBINASE |
| 8 | RED ET RECOMBINATION | 2 | 43% | 1% | 3 | Search RED+ET+RECOMBINATION | Search RED+ET+RECOMBINATION |
| 9 | OLIGO RECOMBINATION | 1 | 100% | 1% | 2 | Search OLIGO+RECOMBINATION | Search OLIGO+RECOMBINATION |
| 10 | MARKERLESS DELETION | 1 | 40% | 1% | 2 | Search MARKERLESS+DELETION | Search MARKERLESS+DELETION |
Key Words Plus |
| Rank | Web of Science journal category | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
|---|---|---|---|---|---|
| 1 | BACTERIAL ARTIFICIAL CHROMOSOMES | 9 | 17% | 13% | 45 |
| 2 | ESCHERICHIA COLI RECE | 8 | 75% | 2% | 6 |
| 3 | PROMOTES RENATURATION | 8 | 62% | 2% | 8 |
| 4 | ANNEALING PROTEINS | 4 | 67% | 1% | 4 |
| 5 | LAMBDA RED | 3 | 50% | 1% | 5 |
| 6 | GENE REPLACEMENT | 3 | 10% | 8% | 29 |
| 7 | RECT PROTEIN | 3 | 60% | 1% | 3 |
| 8 | LINEAR DNA FRAGMENTS | 2 | 50% | 1% | 3 |
| 9 | BACTERIOPHAGE LAMBDA RECOMBINATION | 2 | 36% | 1% | 4 |
| 10 | RED PATHWAY | 2 | 29% | 1% | 5 |
Journals |
Reviews |
| Title | Publ. year | Cit. | Active references | % act. ref. to same field |
|---|---|---|---|---|
| MULTIPLEXED GENOME ENGINEERING AND GENOTYPING METHODS: APPLICATIONS FOR SYNTHETIC BIOLOGY AND METABOLIC ENGINEERING | 2011 | 26 | 29 | 55% |
| Recombineering: In vivo genetic engineering in E-coli, S-enterica, and beyond | 2007 | 79 | 59 | 53% |
| Recombineering BAC transgenes for protein tagging | 2011 | 11 | 28 | 61% |
| Genome engineering and gene expression control for bacterial strain development | 2015 | 3 | 92 | 18% |
| Recombineering to homogeneity: extension of multiplex recombineering to large-scale genome editing | 2013 | 5 | 49 | 59% |
| lambda-red genetic engineering in Salmonella enterica serovar typhimurium | 2007 | 42 | 23 | 52% |
| Recombineering: Highly Efficient in vivo Genetic Engineering using Single-strand Oligos | 2013 | 1 | 11 | 100% |
| A RECOMBINEERING PIPELINE TO MAKE CONDITIONAL TARGETING CONSTRUCTS | 2010 | 15 | 31 | 52% |
| Phage Recombinases and Their Applications | 2012 | 19 | 153 | 31% |
| Genome-scale genetic engineering in Escherichia coli | 2013 | 5 | 68 | 38% |
Address terms |
| Rank | Address term | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
|---|---|---|---|---|---|
| 1 | MOL CONTROL GENET SECT | 4 | 40% | 2.3% | 8 |
| 2 | GENE REGULAT CHROMOSOME BIOL | 3 | 11% | 6.3% | 22 |
| 3 | BIOINNOVAT ZENTRUM | 3 | 22% | 2.9% | 10 |
| 4 | AJINOMOTO GENET | 2 | 67% | 0.6% | 2 |
| 5 | REGENERAT THER Y D DEN | 2 | 67% | 0.6% | 2 |
| 6 | GENE EXP S PROGRAM | 1 | 17% | 2.3% | 8 |
| 7 | BIOINNOVAT ZENTRUM D DEN | 1 | 100% | 0.6% | 2 |
| 8 | BIOINNOVATIONSZENTRUM | 1 | 100% | 0.6% | 2 |
| 9 | ES CELL IL | 1 | 100% | 0.6% | 2 |
| 10 | GENE BRIDGES GMBH | 1 | 33% | 0.6% | 2 |
Related classes at same level (level 1) |
| Rank | Relatedness score | Related classes |
|---|---|---|
| 1 | 0.0000255101 | SINGLE STRANDED OLIGONUCLEOTIDE//GENE REPAIR//SINGLE STRANDED OLIGONUCLEOTIDES |
| 2 | 0.0000186089 | HOLLIDAY JUNCTION//RECBCD ENZYME//RECBCD |
| 3 | 0.0000173388 | SYNTHETIC BIOLOGY//ACS SYNTHETIC BIOLOGY//CALIF QUANTITAT BIOL QB3 |
| 4 | 0.0000172391 | CRE RECOMBINASE//LOXP//ROSA26 |
| 5 | 0.0000138668 | T VECTOR//HIGH THROUGHPUT CLONING//TA CLONING |
| 6 | 0.0000130566 | ESSENTIAL GENES//SUBTRACTIVE GENOMICS//ESSENTIAL GENE |
| 7 | 0.0000123466 | BLOOMINGTON DROSOPHILA STOCK//ENDS OUT TARGETING//ENDS OUT |
| 8 | 0.0000107188 | BIBAC//TAR CLONING//BIBAC LIBRARY |
| 9 | 0.0000090800 | POLYLINKER//STREPTOMYCIN SENSITIVITY//INSERTIONAL INACTIVATION |
| 10 | 0.0000086969 | TYRR//SHIKIMIC ACID PRODUCTION//CIS CIS MUCONIC ACID |