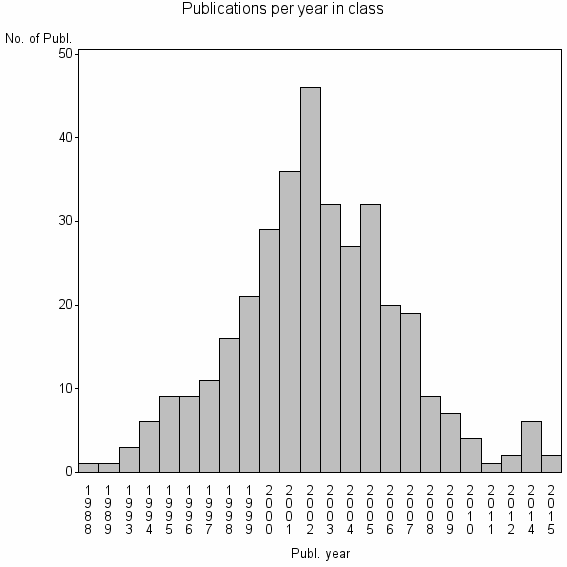
Class information for: |
Basic class information |
| ID | Publications | Average number of references |
Avg. shr. active ref. in WoS |
|---|---|---|---|
| 21755 | 349 | 40.1 | 81% |
Classes in level above (level 2) |
| ID, lev. above |
Publications | Label for level above |
|---|---|---|
| 777 | 11593 | MADS BOX GENES//MADS BOX//FLOWER DEVELOPMENT |
Terms with highest relevance score |
| Rank | Term | Type of term | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
|---|---|---|---|---|---|---|
| 1 | RICE GENOME PROGRAM | Address | 6 | 33% | 4% | 14 |
| 2 | CROP DESIGN | Address | 2 | 67% | 1% | 2 |
| 3 | GENOME IL | Address | 2 | 29% | 1% | 5 |
| 4 | RICE GENOME | Author keyword | 2 | 23% | 2% | 6 |
| 5 | PROGRAMA POS BIOTECNOL GENOM | Address | 1 | 100% | 1% | 2 |
| 6 | LITA ANNENBERG HAZEN GENOME | Address | 1 | 40% | 1% | 2 |
| 7 | PEKING YALE JOINT PLANT MOL GENET AGROB | Address | 1 | 17% | 1% | 5 |
| 8 | IRDUMR 5096 | Address | 1 | 33% | 1% | 2 |
| 9 | HANGZHOU GENOM | Address | 1 | 18% | 1% | 4 |
| 10 | ANCHOR PROBES | Author keyword | 1 | 50% | 0% | 1 |
Web of Science journal categories |
Author Key Words |
| Rank | Web of Science journal category | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
LCSH search | Wikipedia search |
|---|---|---|---|---|---|---|---|
| 1 | RICE GENOME | 2 | 23% | 2% | 6 | Search RICE+GENOME | Search RICE+GENOME |
| 2 | ANCHOR PROBES | 1 | 50% | 0% | 1 | Search ANCHOR+PROBES | Search ANCHOR+PROBES |
| 3 | ARABIDOPSIS THALIANA CHROMOSOME 3 | 1 | 50% | 0% | 1 | Search ARABIDOPSIS+THALIANA+CHROMOSOME+3 | Search ARABIDOPSIS+THALIANA+CHROMOSOME+3 |
| 4 | CPG DEPLETION | 1 | 50% | 0% | 1 | Search CPG+DEPLETION | Search CPG+DEPLETION |
| 5 | DOUBLE COLOR FISH | 1 | 50% | 0% | 1 | Search DOUBLE+COLOR+FISH | Search DOUBLE+COLOR+FISH |
| 6 | DROUGHT ENVIRONMENTS | 1 | 50% | 0% | 1 | Search DROUGHT+ENVIRONMENTS | Search DROUGHT+ENVIRONMENTS |
| 7 | GENE FUNCTION IDENTIFICATION | 1 | 50% | 0% | 1 | Search GENE+FUNCTION+IDENTIFICATION | Search GENE+FUNCTION+IDENTIFICATION |
| 8 | GENOMIC DNA ARRAY | 1 | 50% | 0% | 1 | Search GENOMIC+DNA+ARRAY | Search GENOMIC+DNA+ARRAY |
| 9 | GERMINATION DORMANCY | 1 | 50% | 0% | 1 | Search GERMINATION+DORMANCY | Search GERMINATION+DORMANCY |
| 10 | MAP BASED SEQUENCE | 1 | 50% | 0% | 1 | Search MAP+BASED+SEQUENCE | Search MAP+BASED+SEQUENCE |
Key Words Plus |
| Rank | Web of Science journal category | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
|---|---|---|---|---|---|
| 1 | DRAFT SEQUENCE | 9 | 10% | 24% | 84 |
| 2 | PLANT NUCLEAR GENOMES | 3 | 50% | 1% | 5 |
| 3 | DUPLICATED SEGMENTS | 3 | 40% | 2% | 6 |
| 4 | GRASS GENOMES | 2 | 14% | 4% | 14 |
| 5 | MONOCOT DICOT DIVERGENCE | 2 | 20% | 3% | 9 |
| 6 | COMPARATIVE GENOMICS STRATEGY | 1 | 100% | 1% | 2 |
| 7 | GRASS GENOME | 1 | 50% | 1% | 2 |
| 8 | TAGGED CONNECTORS | 1 | 50% | 1% | 2 |
| 9 | COMPARATIVE GENETICS | 1 | 11% | 3% | 9 |
| 10 | DEVELOPING ARABIDOPSIS | 1 | 40% | 1% | 2 |
Journals |
Reviews |
| Title | Publ. year | Cit. | Active references |
% act. ref. to same field |
|---|---|---|---|---|
| Updating the 'Crop circle' | 2005 | 150 | 55 | 25% |
| The genetic colinearity of rice and other cereals on the basis of genomic sequence analysis | 2003 | 74 | 42 | 55% |
| Hot paper in genomics - History in a grain of rice | 2007 | 0 | 1 | 100% |
| Decoding the rice genome | 2006 | 21 | 86 | 44% |
| From mapping to sequencing, post-sequencing and beyond | 2005 | 34 | 33 | 42% |
| Re-evaluating the relevance of ancestral shared synteny as a tool for crop improvement | 2004 | 25 | 44 | 45% |
| From databases to modelling of functional pathways | 2004 | 2 | 6 | 83% |
| Duplicate and diverge: the evolution of plant genome microstructure | 2001 | 36 | 30 | 37% |
| Towards an accurate sequence of the rice genome | 2003 | 23 | 23 | 57% |
| Rice as a model for cereal genomics | 1999 | 53 | 27 | 52% |
Address terms |
| Rank | Address term | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
|---|---|---|---|---|---|
| 1 | RICE GENOME PROGRAM | 6 | 33% | 4.0% | 14 |
| 2 | CROP DESIGN | 2 | 67% | 0.6% | 2 |
| 3 | GENOME IL | 2 | 29% | 1.4% | 5 |
| 4 | PROGRAMA POS BIOTECNOL GENOM | 1 | 100% | 0.6% | 2 |
| 5 | LITA ANNENBERG HAZEN GENOME | 1 | 40% | 0.6% | 2 |
| 6 | PEKING YALE JOINT PLANT MOL GENET AGROB | 1 | 17% | 1.4% | 5 |
| 7 | IRDUMR 5096 | 1 | 33% | 0.6% | 2 |
| 8 | HANGZHOU GENOM | 1 | 18% | 1.1% | 4 |
| 9 | BIOINFORMAT COMPARAT GENOM | 1 | 50% | 0.3% | 1 |
| 10 | COLD SPRING HARBOR PLANT BIOL GRP | 1 | 50% | 0.3% | 1 |
Related classes at same level (level 1) |
| Rank | Relatedness score | Related classes |
|---|---|---|
| 1 | 0.0000251755 | BRACHYPODIUM DISTACHYON//BRACHYPODIUM//BRACHYPODIUM HYBRIDUM |
| 2 | 0.0000249310 | RETROTRANSPOSONS//TY1 COPIA//TRANSPOSABLE ELEMENTS |
| 3 | 0.0000204754 | LONG GLUME//TRITICUM POLONICUM//TRITICUM PETROPAVLOVSKYI |
| 4 | 0.0000193437 | CIS MOTIF//ARABIDOPSIS T87 CELL LINE//GENE ATLAS |
| 5 | 0.0000186557 | ORYZA RUFIPOGON//WIDE COMPATIBILITY//COMMON WILD RICE |
| 6 | 0.0000156400 | WHOLE GENOME DUPLICATION//GENET MOL CELLULAIRE ECT LEVU INTERE//GENOME DUPLICATION |
| 7 | 0.0000155713 | AC DS//TRANSPOSON TAGGING//AC TRANSPOSASE |
| 8 | 0.0000150501 | MKIAA//SMART CDNA LIBRARY//CDNA SEQUENCING |
| 9 | 0.0000137435 | PLANT MICROBIAL GENOM//LRT1//NONADDITIVE PROTEIN |
| 10 | 0.0000131068 | BIBAC//TAR CLONING//BIBAC LIBRARY |