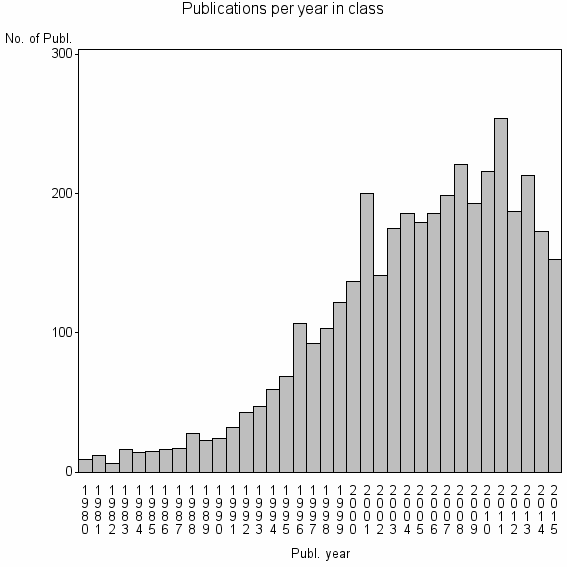
Class information for: |
Basic class information |
| ID | Publications | Average number of references |
Avg. shr. active ref. in WoS |
|---|---|---|---|
| 211 | 3867 | 41.0 | 71% |
Classes in level above (level 2) |
Terms with highest relevance score |
| Rank | Term | Type of term | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
|---|---|---|---|---|---|---|
| 1 | Y CHROMOSOME | Author keyword | 227 | 34% | 14% | 553 |
| 2 | Y STR | Author keyword | 114 | 66% | 3% | 106 |
| 3 | Y STRS | Author keyword | 95 | 78% | 2% | 62 |
| 4 | HAPLOGROUP | Author keyword | 87 | 49% | 3% | 129 |
| 5 | DYS19 | Author keyword | 77 | 73% | 1% | 58 |
| 6 | DYS391 | Author keyword | 71 | 75% | 1% | 51 |
| 7 | DYS392 | Author keyword | 68 | 80% | 1% | 43 |
| 8 | Y SNPS | Author keyword | 61 | 86% | 1% | 31 |
| 9 | FORENSIC SCIENCE INTERNATIONAL-GENETICS | Journal | 61 | 25% | 6% | 213 |
| 10 | DYS393 | Author keyword | 60 | 72% | 1% | 47 |
Web of Science journal categories |
Author Key Words |
| Rank | Web of Science journal category | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
LCSH search | Wikipedia search |
|---|---|---|---|---|---|---|---|
| 1 | Y CHROMOSOME | 227 | 34% | 14% | 553 | Search Y+CHROMOSOME | Search Y+CHROMOSOME |
| 2 | Y STR | 114 | 66% | 3% | 106 | Search Y+STR | Search Y+STR |
| 3 | Y STRS | 95 | 78% | 2% | 62 | Search Y+STRS | Search Y+STRS |
| 4 | HAPLOGROUP | 87 | 49% | 3% | 129 | Search HAPLOGROUP | Search HAPLOGROUP |
| 5 | DYS19 | 77 | 73% | 1% | 58 | Search DYS19 | Search DYS19 |
| 6 | DYS391 | 71 | 75% | 1% | 51 | Search DYS391 | Search DYS391 |
| 7 | DYS392 | 68 | 80% | 1% | 43 | Search DYS392 | Search DYS392 |
| 8 | Y SNPS | 61 | 86% | 1% | 31 | Search Y+SNPS | Search Y+SNPS |
| 9 | DYS393 | 60 | 72% | 1% | 47 | Search DYS393 | Search DYS393 |
| 10 | DYS390 | 52 | 68% | 1% | 45 | Search DYS390 | Search DYS390 |
Key Words Plus |
| Rank | Web of Science journal category | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
|---|---|---|---|---|---|
| 1 | NEAR EASTERN | 228 | 90% | 3% | 99 |
| 2 | MTDNA VARIATION | 207 | 59% | 6% | 234 |
| 3 | HUMAN MITOCHONDRIAL DNA | 186 | 40% | 9% | 360 |
| 4 | MIGRATORY EVENTS | 163 | 90% | 2% | 71 |
| 5 | STR HAPLOTYPES | 144 | 91% | 2% | 60 |
| 6 | MTDNA HAPLOGROUP H | 103 | 91% | 1% | 42 |
| 7 | 9 BP DELETION | 92 | 82% | 1% | 54 |
| 8 | HAPLOGROUPS | 89 | 51% | 3% | 126 |
| 9 | MTDNA | 86 | 20% | 10% | 378 |
| 10 | HAPLOGROUP H | 85 | 92% | 1% | 34 |
Journals |
| Rank | Web of Science journal category | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
|---|---|---|---|---|---|
| 1 | FORENSIC SCIENCE INTERNATIONAL-GENETICS | 61 | 25% | 6% | 213 |
| 2 | HUMAN BIOLOGY | 29 | 12% | 6% | 223 |
| 3 | JOURNAL OF ANTHROPOLOGICAL SCIENCES | 1 | 11% | 0% | 10 |
Reviews |
| Title | Publ. year | Cit. | Active references | % act. ref. to same field |
|---|---|---|---|---|
| Correcting for Purifying Selection: An Improved Human Mitochondrial Molecular Clock | 2009 | 216 | 104 | 56% |
| Harvesting the fruit of the human mtDNA tree | 2006 | 210 | 49 | 84% |
| The Archaeogenetics of Europe | 2010 | 74 | 97 | 67% |
| The Human Genetic History of the Americas: The Final Frontier | 2010 | 57 | 37 | 84% |
| The impact of whole-genome sequencing on the reconstruction of human population history APPLICATIONS OF NEXT-GENERATION SEQUENCING | 2014 | 21 | 157 | 29% |
| Use of Y chromosome and mitochondrial DNA population structure in tracing human migrations | 2007 | 138 | 116 | 65% |
| The human Y chromosome: An evolutionary marker comes of age | 2003 | 414 | 114 | 51% |
| Genetic variation and population structure in Native Americans | 2007 | 178 | 101 | 47% |
| The Human Genetic History of East Asia: Weaving a Complex Tapestry | 2010 | 31 | 62 | 89% |
| Farmers and their languages: The first expansions | 2003 | 248 | 17 | 65% |
Address terms |
| Rank | Address term | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
|---|---|---|---|---|---|
| 1 | GENET MOL POBLAC | 45 | 90% | 0.5% | 19 |
| 2 | UNIDADE XENET | 30 | 56% | 0.9% | 36 |
| 3 | ARCHAEOGENET | 26 | 80% | 0.4% | 16 |
| 4 | FORENS MOL BIOL | 25 | 36% | 1.5% | 57 |
| 5 | MED LEGAL | 25 | 35% | 1.5% | 59 |
| 6 | FORENS GENET | 24 | 30% | 1.7% | 66 |
| 7 | ARMED FORCES DNA IDENTIFICAT | 21 | 64% | 0.5% | 21 |
| 8 | UNIDAD XENET | 21 | 90% | 0.2% | 9 |
| 9 | BIOL PROBLEMS N | 20 | 25% | 1.8% | 70 |
| 10 | CHINESE FRONTIER ARCHAEOL | 20 | 54% | 0.7% | 26 |
Related classes at same level (level 1) |
| Rank | Relatedness score | Related classes |
|---|---|---|
| 1 | 0.0000133056 | ANCESTRY INFORMATIVE MARKERS//POPULATION STRATIFICATION//GENOMIC CONTROL |
| 2 | 0.0000125424 | ANCIENT DNA//GEOGENET//SKELETAL PREPARATION |
| 3 | 0.0000106099 | DNA TYPING//FORENSIC SCIENCE INTERNATIONAL-GENETICS//SHORT TANDEM REPEATS |
| 4 | 0.0000083398 | CULTURAL PHYLOGENETICS//LEXICOSTATISTICS//HUMAN EVOLUTIONARY ECOL GRP |
| 5 | 0.0000081296 | D310//MITOCHONDRIAL MICROSATELLITE INSTABILITY//MITOCHONDRIAL HAPLOGROUPS |
| 6 | 0.0000075616 | TRANSFERRIN C SUBTYPES//BLOOD GENETIC MARKERS//GENE GEOGRAPHY |
| 7 | 0.0000072937 | ANTHROPOLOGICAL SCIENCE//BIODISTANCE//INTERPARIETAL BONE |
| 8 | 0.0000065508 | ISONYMY//SURNAMES//SURNAME DISTRIBUTION |
| 9 | 0.0000063079 | CIENCIAS NAT ARQUEOL//NEW WORLD COLONIZATION//FIRST AMERICANS |
| 10 | 0.0000058811 | BRITONS//BELGAE//BIOL S 4100 |