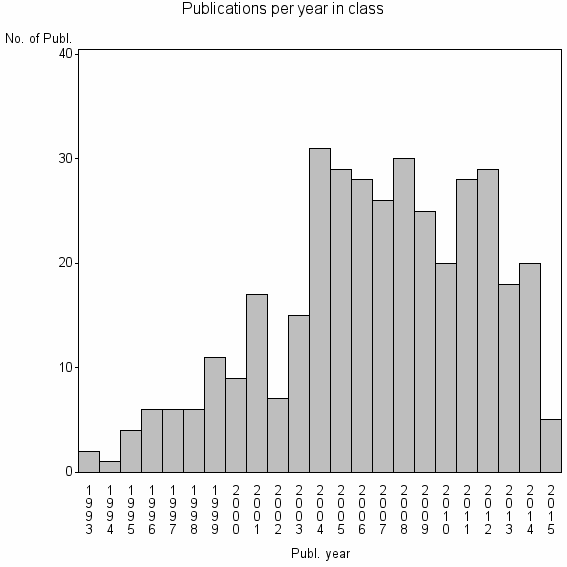
Class information for: |
Basic class information |
| ID | Publications | Average number of references |
Avg. shr. active ref. in WoS |
|---|---|---|---|
| 21081 | 373 | 46.5 | 92% |
Classes in level above (level 2) |
| ID, lev. above |
Publications | Label for level above |
|---|---|---|
| 238 | 19563 | BIOINFORMATICS//BMC BIOINFORMATICS//GENOME RESEARCH |
Terms with highest relevance score |
| Rank | Term | Type of term | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
|---|---|---|---|---|---|---|
| 1 | IROQUOIS | Author keyword | 5 | 24% | 5% | 17 |
| 2 | CONSERVED NONCODING ELEMENTS | Author keyword | 3 | 57% | 1% | 4 |
| 3 | ENERGY ENVIRONM BIOL COMP | Address | 3 | 100% | 1% | 3 |
| 4 | CONSERVED NON CODING ELEMENTS | Author keyword | 3 | 40% | 2% | 6 |
| 5 | SARS MARINE MOL BIOL | Address | 3 | 21% | 3% | 13 |
| 6 | CHORDO NEURAL HINGE | Author keyword | 2 | 67% | 1% | 2 |
| 7 | ULTRACONSERVED ELEMENTS | Author keyword | 2 | 30% | 2% | 6 |
| 8 | ULTRACONSERVATION | Author keyword | 1 | 50% | 1% | 2 |
| 9 | ULTRACONSERVED ELEMENTS UCES | Author keyword | 1 | 100% | 1% | 2 |
| 10 | ULTRACONSERVED REGIONS | Author keyword | 1 | 50% | 1% | 2 |
Web of Science journal categories |
Author Key Words |
| Rank | Web of Science journal category | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
LCSH search | Wikipedia search |
|---|---|---|---|---|---|---|---|
| 1 | IROQUOIS | 5 | 24% | 5% | 17 | Search IROQUOIS | Search IROQUOIS |
| 2 | CONSERVED NONCODING ELEMENTS | 3 | 57% | 1% | 4 | Search CONSERVED+NONCODING+ELEMENTS | Search CONSERVED+NONCODING+ELEMENTS |
| 3 | CONSERVED NON CODING ELEMENTS | 3 | 40% | 2% | 6 | Search CONSERVED+NON+CODING+ELEMENTS | Search CONSERVED+NON+CODING+ELEMENTS |
| 4 | CHORDO NEURAL HINGE | 2 | 67% | 1% | 2 | Search CHORDO+NEURAL+HINGE | Search CHORDO+NEURAL+HINGE |
| 5 | ULTRACONSERVED ELEMENTS | 2 | 30% | 2% | 6 | Search ULTRACONSERVED+ELEMENTS | Search ULTRACONSERVED+ELEMENTS |
| 6 | ULTRACONSERVATION | 1 | 50% | 1% | 2 | Search ULTRACONSERVATION | Search ULTRACONSERVATION |
| 7 | ULTRACONSERVED ELEMENTS UCES | 1 | 100% | 1% | 2 | Search ULTRACONSERVED+ELEMENTS+UCES | Search ULTRACONSERVED+ELEMENTS+UCES |
| 8 | ULTRACONSERVED REGIONS | 1 | 50% | 1% | 2 | Search ULTRACONSERVED+REGIONS | Search ULTRACONSERVED+REGIONS |
| 9 | IRX | 1 | 22% | 1% | 4 | Search IRX | Search IRX |
| 10 | SYNTENY CONSERVATION | 1 | 33% | 1% | 2 | Search SYNTENY+CONSERVATION | Search SYNTENY+CONSERVATION |
Key Words Plus |
| Rank | Web of Science journal category | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
|---|---|---|---|---|---|
| 1 | ULTRACONSERVED ELEMENTS | 20 | 30% | 15% | 55 |
| 2 | GENE DESERTS | 15 | 56% | 5% | 18 |
| 3 | CONSERVED NONCODING SEQUENCES | 13 | 26% | 12% | 44 |
| 4 | XENOPUS HOMOLOG | 8 | 36% | 5% | 19 |
| 5 | NONGENIC SEQUENCES | 8 | 100% | 1% | 5 |
| 6 | NON GENIC SEQUENCES | 7 | 53% | 2% | 9 |
| 7 | VIABLE MICE | 5 | 63% | 1% | 5 |
| 8 | NONCODING SEQUENCES | 5 | 13% | 10% | 36 |
| 9 | MULTISPECIES CONSERVATION | 4 | 67% | 1% | 4 |
| 10 | KEY DEVELOPMENTAL GENES | 4 | 42% | 2% | 8 |
Journals |
Reviews |
| Title | Publ. year | Cit. | Active references | % act. ref. to same field |
|---|---|---|---|---|
| The mystery of extreme non-coding conservation | 2013 | 8 | 116 | 53% |
| Conserved non-coding elements and cis regulation: actions speak louder than words | 2013 | 16 | 93 | 27% |
| Tuning in to the signals: noncoding sequence conservation in vertebrate genornes | 2008 | 51 | 59 | 34% |
| Organization of Conserved Elements Near Key Developmental Regulators in Vertebrate Genomes | 2008 | 20 | 84 | 43% |
| Fugu: a compact vertebrate reference genome | 2000 | 81 | 36 | 44% |
| Deep conservation of cis-regulatory elements in metazoans | 2013 | 4 | 119 | 31% |
| Iroquois genes: genomic organization and function in vertebrate neural development | 2002 | 84 | 44 | 32% |
| Conserved non-genic sequences - an unexpected feature of mammalian genomes | 2005 | 159 | 53 | 23% |
| When needles look like hay: How to find tissue-specific enhancers in model organism genomes | 2011 | 15 | 281 | 19% |
| Comparative genomics at the vertebrate extremes | 2004 | 152 | 79 | 20% |
Address terms |
| Rank | Address term | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
|---|---|---|---|---|---|
| 1 | ENERGY ENVIRONM BIOL COMP | 3 | 100% | 0.8% | 3 |
| 2 | SARS MARINE MOL BIOL | 3 | 21% | 3.5% | 13 |
| 3 | ROSALIND FRANKLIN GENOM | 1 | 16% | 1.3% | 5 |
| 4 | NED UPR3294 | 1 | 33% | 0.5% | 2 |
| 5 | BORDET CANC | 1 | 50% | 0.3% | 1 |
| 6 | COMPUTAT CANC GENOM GRP | 1 | 50% | 0.3% | 1 |
| 7 | HEART STROKE RICHARD LEWAR CARDIOVASC | 1 | 50% | 0.3% | 1 |
| 8 | MARINE FUNCT GENOM STUDIES | 1 | 50% | 0.3% | 1 |
| 9 | PROGRAM NEUROSCI DEV BIOL BIOMED SCI | 1 | 50% | 0.3% | 1 |
| 10 | TROP AQUACULTURE PROGRAM FISHERIES AQUAT SC | 1 | 50% | 0.3% | 1 |
Related classes at same level (level 1) |
| Rank | Relatedness score | Related classes |
|---|---|---|
| 1 | 0.0000190556 | CHIP SEQ//PEAK CALLING//DNASE SEQ |
| 2 | 0.0000169140 | WHOLE GENOME DUPLICATION//GENET MOL CELLULAIRE ECT LEVU INTERE//GENOME DUPLICATION |
| 3 | 0.0000148554 | MOTIF DISCOVERY//CLOSEST STRING PROBLEM//MOTIF FINDING |
| 4 | 0.0000136034 | BIDIRECTIONAL PROMOTER//NEIGHBORING GENES//GENOME NETWORK PROJECT CORE GRPGENOM SCI |
| 5 | 0.0000119848 | ALFONSINO//BERYX SPLENDENS//GIRELLA LEONINA |
| 6 | 0.0000111350 | MULTIPLE SEQUENCE ALIGNMENT//T COFFEE//STATISTICAL ALIGNMENT |
| 7 | 0.0000096850 | ISOCHORES//EVOLUZ MOL//ISOCHORE |
| 8 | 0.0000080494 | ISTHMIC ORGANIZER//GBX2//RHOMBOMERE 1 |
| 9 | 0.0000079117 | LIMB DEVELOPMENT//APICAL ECTODERMAL RIDGE//LIMB |
| 10 | 0.0000066554 | LONG NONCODING RNAS//LNCRNA//LONG NON CODING RNA |