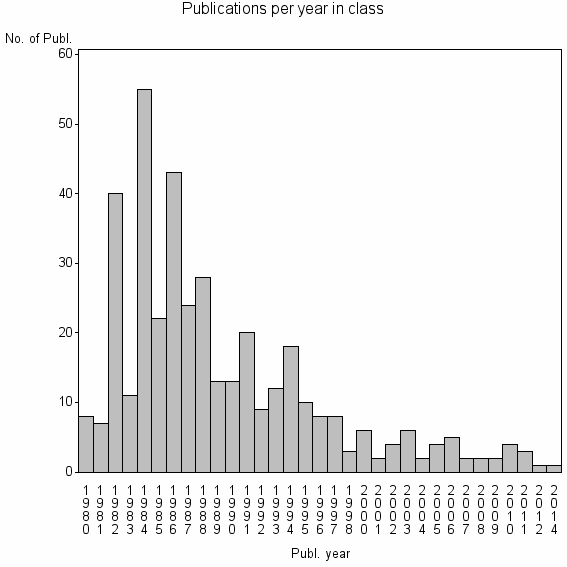
Class information for: |
Basic class information |
| ID | Publications | Average number of references |
Avg. shr. active ref. in WoS |
|---|---|---|---|
| 20492 | 396 | 15.8 | 51% |
Classes in level above (level 2) |
| ID, lev. above |
Publications | Label for level above |
|---|---|---|
| 2449 | 3598 | PREDICTION WITH EXPERT ADVICE//UNIVERSAL CODING//GRAPHICAL REPRESENTATION |
Terms with highest relevance score |
| Rank | Term | Type of term | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
|---|---|---|---|---|---|---|
| 1 | AMBIGRAM | Author keyword | 2 | 67% | 1% | 2 |
| 2 | DESK TOP MOLECULAR MODELING | Author keyword | 2 | 67% | 1% | 2 |
| 3 | PARTIAL DIGEST | Author keyword | 2 | 67% | 1% | 2 |
| 4 | ADAPTIVE WEIGHTED LEAST SQUARES | Author keyword | 1 | 100% | 1% | 2 |
| 5 | DNA PHYSICAL MAPPING | Author keyword | 1 | 50% | 1% | 2 |
| 6 | GRAPH THEORETIC TECHNIQUES | Author keyword | 1 | 40% | 1% | 2 |
| 7 | LEARNING BY QUERIES | Author keyword | 1 | 40% | 1% | 2 |
| 8 | BIOLOGICAL DATA BANKS | Author keyword | 1 | 50% | 0% | 1 |
| 9 | IUPAC NOTATION | Author keyword | 1 | 50% | 0% | 1 |
| 10 | NEUROGENIC LOCUS | Author keyword | 1 | 50% | 0% | 1 |
Web of Science journal categories |
Author Key Words |
| Rank | Web of Science journal category | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
LCSH search | Wikipedia search |
|---|---|---|---|---|---|---|---|
| 1 | AMBIGRAM | 2 | 67% | 1% | 2 | Search AMBIGRAM | Search AMBIGRAM |
| 2 | DESK TOP MOLECULAR MODELING | 2 | 67% | 1% | 2 | Search DESK+TOP+MOLECULAR+MODELING | Search DESK+TOP+MOLECULAR+MODELING |
| 3 | PARTIAL DIGEST | 2 | 67% | 1% | 2 | Search PARTIAL+DIGEST | Search PARTIAL+DIGEST |
| 4 | ADAPTIVE WEIGHTED LEAST SQUARES | 1 | 100% | 1% | 2 | Search ADAPTIVE+WEIGHTED+LEAST+SQUARES | Search ADAPTIVE+WEIGHTED+LEAST+SQUARES |
| 5 | DNA PHYSICAL MAPPING | 1 | 50% | 1% | 2 | Search DNA+PHYSICAL+MAPPING | Search DNA+PHYSICAL+MAPPING |
| 6 | GRAPH THEORETIC TECHNIQUES | 1 | 40% | 1% | 2 | Search GRAPH+THEORETIC+TECHNIQUES | Search GRAPH+THEORETIC+TECHNIQUES |
| 7 | LEARNING BY QUERIES | 1 | 40% | 1% | 2 | Search LEARNING+BY+QUERIES | Search LEARNING+BY+QUERIES |
| 8 | BIOLOGICAL DATA BANKS | 1 | 50% | 0% | 1 | Search BIOLOGICAL+DATA+BANKS | Search BIOLOGICAL+DATA+BANKS |
| 9 | IUPAC NOTATION | 1 | 50% | 0% | 1 | Search IUPAC+NOTATION | Search IUPAC+NOTATION |
| 10 | NEUROGENIC LOCUS | 1 | 50% | 0% | 1 | Search NEUROGENIC+LOCUS | Search NEUROGENIC+LOCUS |
Key Words Plus |
| Rank | Web of Science journal category | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
|---|---|---|---|---|---|
| 1 | MAPPING DNA | 2 | 67% | 1% | 2 |
| 2 | PARTIAL DIGEST PROBLEM | 1 | 50% | 0% | 1 |
| 3 | RELATING MOBILITY | 1 | 50% | 0% | 1 |
| 4 | PROBLEM NP HARD | 0 | 33% | 0% | 1 |
| 5 | HYPERBOLIC REGRESSION | 0 | 25% | 0% | 1 |
| 6 | COMBINING ABILITY ANALYSIS | 0 | 17% | 0% | 1 |
| 7 | MANURE STORAGE PITS | 0 | 13% | 0% | 1 |
| 8 | DIGEST PROBLEM | 0 | 100% | 0% | 1 |
Journals |
Reviews |
| Title | Publ. year | Cit. | Active references | % act. ref. to same field |
|---|---|---|---|---|
| COMPUTER-PROGRAMS FOR ANALYZING DNA AND PROTEIN SEQUENCES | 1987 | 0 | 5 | 100% |
| THE GENBANK NUCLEIC-ACID SEQUENCE DATABASE | 1985 | 55 | 12 | 33% |
| QUANTITATIVE-ANALYSIS OF ENZYMATIC DIGESTS OF DNA USING GEL-ELECTROPHORESIS | 1993 | 4 | 24 | 17% |
| SEQUENCE-ANALYSIS ON MICROCOMPUTERS | 1987 | 10 | 10 | 30% |
| The Internet and molecular biology | 1997 | 4 | 7 | 14% |
| ANALYSIS OF BIOLOGICAL SEQUENCES ON SMALL COMPUTERS | 1984 | 30 | 88 | 56% |
| PROTEIN AND NUCLEIC-ACID SEQUENCE DATABASE-SYSTEMS | 1983 | 40 | 12 | 42% |
| NUCLEIC-ACID AND PROTEIN-SEQUENCE DATABASES | 1985 | 11 | 30 | 40% |
| COMPUTER-APPLICATIONS IN APPLIED GENETIC-ENGINEERING | 1984 | 4 | 8 | 38% |
| COMPUTERS IN BIOCHEMICAL-ANALYSIS | 1985 | 3 | 134 | 29% |
Address terms |
| Rank | Address term | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
|---|---|---|---|---|---|
| 1 | ELECT ENGN COMP SCI MC154 | 0 | 33% | 0.3% | 1 |
| 2 | IST GENET MOL EVOLUTIONARY GENET | 0 | 33% | 0.3% | 1 |
| 3 | 110 AVE PINS OUEST | 0 | 25% | 0.3% | 1 |
| 4 | DIPARTIMENTO INFORMAT PLICAZ RENATO M C OCEL | 0 | 25% | 0.3% | 1 |
| 5 | VISUAL STUDIES | 0 | 13% | 0.5% | 2 |
| 6 | EBUSINESS TECHNOL | 0 | 13% | 0.3% | 1 |
| 7 | DISCRETE SIMULAT SCI CCS5 | 0 | 100% | 0.3% | 1 |
| 8 | GRASCE | 0 | 100% | 0.3% | 1 |
Related classes at same level (level 1) |
| Rank | Relatedness score | Related classes |
|---|---|---|
| 1 | 0.0000121096 | ORDERED FACTORIZATIONS//FACTORISATIO NUMERORUM//IMPROPER EDGE |
| 2 | 0.0000092117 | MULTIPLE SEQUENCE ALIGNMENT//T COFFEE//STATISTICAL ALIGNMENT |
| 3 | 0.0000081099 | EMBL OUTSTN//PROT INFORMAT OURCE//HLTH SCI TECHNOL ICIR |
| 4 | 0.0000080009 | DYE TERMINATOR//ENERGY TRANSFER DYES//GENOMAT |
| 5 | 0.0000059848 | LONGEST COMMON SUBSEQUENCE//CONSTRAINED LONGEST COMMON SUBSEQUENCE//SHORTEST COMMON SUPERSTRING |
| 6 | 0.0000054905 | SET COVARIANCE//SAS//BOUNDED CONVEX DOMAIN |
| 7 | 0.0000048700 | THEORY OF ORGANISM//ARTIFICIAL CREATU//GUZEL SA AR TASARIM FAK |
| 8 | 0.0000045708 | MEGAPRIMER//OVERLAP EXTENSION//MULTIPLE SITE DIRECTED MUTAGENESIS |
| 9 | 0.0000043738 | ZINNIA VIOLACEA//XANTHOMONAS CAMPESTRIS PV ZINNIAE//ZINNIA MARYLANDICA |
| 10 | 0.0000043544 | LATENT PERIODICITY//3 BASE PERIODICITY//DNA WALK |