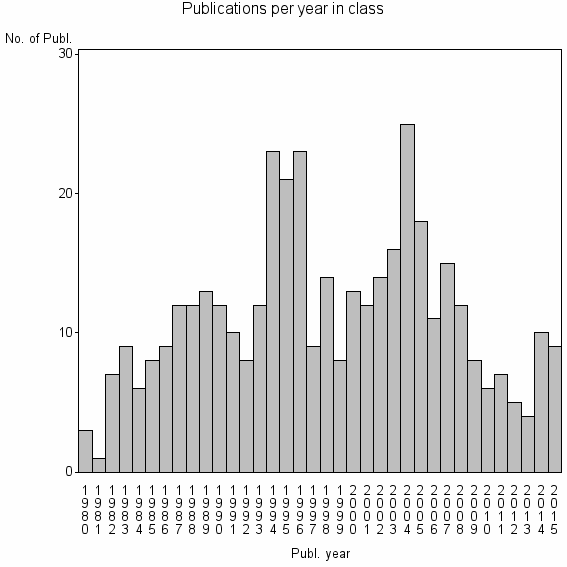
Class information for: |
Basic class information |
| ID | Publications | Average number of references |
Avg. shr. active ref. in WoS |
|---|---|---|---|
| 20276 | 405 | 32.6 | 68% |
Classes in level above (level 2) |
| ID, lev. above |
Publications | Label for level above |
|---|---|---|
| 1852 | 5569 | LESCH NYHAN SYNDROME//TIAZOFURIN//LESCH NYHAN DISEASE |
Terms with highest relevance score |
| Rank | Term | Type of term | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
|---|---|---|---|---|---|---|
| 1 | CARAB | Author keyword | 15 | 88% | 2% | 7 |
| 2 | PYRIMIDINE REGULATION | Author keyword | 6 | 100% | 1% | 4 |
| 3 | PYRR | Author keyword | 4 | 75% | 1% | 3 |
| 4 | TRANSCRIPTION ATTENUATION | Author keyword | 4 | 36% | 2% | 8 |
| 5 | TRYPTOPHAN BINDING | Author keyword | 3 | 57% | 1% | 4 |
| 6 | HUTP | Author keyword | 3 | 100% | 1% | 3 |
| 7 | TRP RNA BINDING ATTENUATION PROTEIN | Author keyword | 2 | 67% | 0% | 2 |
| 8 | HEDDLE INITIAT UNIT | Address | 2 | 33% | 1% | 4 |
| 9 | 2 OH | Author keyword | 1 | 50% | 0% | 2 |
| 10 | ARTIFICIAL CAPSID | Author keyword | 1 | 100% | 0% | 2 |
Web of Science journal categories |
Author Key Words |
| Rank | Web of Science journal category | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
LCSH search | Wikipedia search |
|---|---|---|---|---|---|---|---|
| 1 | CARAB | 15 | 88% | 2% | 7 | Search CARAB | Search CARAB |
| 2 | PYRIMIDINE REGULATION | 6 | 100% | 1% | 4 | Search PYRIMIDINE+REGULATION | Search PYRIMIDINE+REGULATION |
| 3 | PYRR | 4 | 75% | 1% | 3 | Search PYRR | Search PYRR |
| 4 | TRANSCRIPTION ATTENUATION | 4 | 36% | 2% | 8 | Search TRANSCRIPTION+ATTENUATION | Search TRANSCRIPTION+ATTENUATION |
| 5 | TRYPTOPHAN BINDING | 3 | 57% | 1% | 4 | Search TRYPTOPHAN+BINDING | Search TRYPTOPHAN+BINDING |
| 6 | HUTP | 3 | 100% | 1% | 3 | Search HUTP | Search HUTP |
| 7 | TRP RNA BINDING ATTENUATION PROTEIN | 2 | 67% | 0% | 2 | Search TRP+RNA+BINDING+ATTENUATION+PROTEIN | Search TRP+RNA+BINDING+ATTENUATION+PROTEIN |
| 8 | 2 OH | 1 | 50% | 0% | 2 | Search 2++OH | Search 2++OH |
| 9 | ARTIFICIAL CAPSID | 1 | 100% | 0% | 2 | Search ARTIFICIAL+CAPSID | Search ARTIFICIAL+CAPSID |
| 10 | HUTP PROTEIN | 1 | 100% | 0% | 2 | Search HUTP+PROTEIN | Search HUTP+PROTEIN |
Key Words Plus |
| Rank | Web of Science journal category | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
|---|---|---|---|---|---|
| 1 | MTRB | 31 | 92% | 3% | 12 |
| 2 | BINDING ATTENUATION PROTEIN | 29 | 51% | 10% | 41 |
| 3 | ANTI TRAP | 21 | 78% | 3% | 14 |
| 4 | ATTENUATION PROTEIN TRAP | 21 | 85% | 3% | 11 |
| 5 | TRANSCRIPTION ATTENUATION | 18 | 43% | 8% | 32 |
| 6 | PYRIMIDINE BIOSYNTHETIC OPERON | 17 | 79% | 3% | 11 |
| 7 | SUBTILIS TRP OPERON | 17 | 75% | 3% | 12 |
| 8 | MUTANT SUBUNITS | 15 | 88% | 2% | 7 |
| 9 | AUTOGENOUS TRANSCRIPTIONAL ATTENUATION | 14 | 100% | 2% | 7 |
| 10 | BACILLUS SUBTILIS BINDS | 11 | 100% | 1% | 6 |
Journals |
Reviews |
| Title | Publ. year | Cit. | Active references | % act. ref. to same field |
|---|---|---|---|---|
| Complexity in regulation of tryptophan biosynthesis in Bacillus subtilis | 2005 | 86 | 60 | 85% |
| Regulation of pyrimidine biosynthetic gene expression in bacteria: Repression without repressors | 2008 | 50 | 150 | 61% |
| Regulation of transcription attenuation and translation initiation by allosteric control of an RNA-binding protein: the Bacillus subtilis TRAP protein | 2004 | 38 | 40 | 98% |
| The different roles of tryptophan transfer RNA in regulating trp operon expression in E-coli versus B-subtilis | 2004 | 36 | 28 | 64% |
| Regulation of tryptophan biosynthesis: Trp-ing the TRAP or how Bacillus subtilis reinvented the wheel | 1997 | 55 | 25 | 92% |
| Posttranscription initiation control of tryptophan metabolism in Bacillus subtilis by the trp RNA-binding attenuation protein (TRAP), anti-TRAP, and RNA structure | 2001 | 41 | 39 | 87% |
| RNA-based regulation of genes of tryptophan synthesis and degradation, in bacteria | 2007 | 35 | 43 | 35% |
| Regulation of gene expression by reiterative transcription | 2011 | 10 | 34 | 38% |
| Structural basis of HutP-mediated transcription anti-termination | 2006 | 4 | 19 | 68% |
| Nucleotide metabolism and its control in lactic acid bacteria | 2005 | 52 | 138 | 22% |
Address terms |
| Rank | Address term | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
|---|---|---|---|---|---|
| 1 | HEDDLE INITIAT UNIT | 2 | 33% | 1.0% | 4 |
| 2 | DYNAM EXP S EVOLUT GENOMES MICROORGANISME | 1 | 100% | 0.5% | 2 |
| 3 | LIGHT IND TECHNOL ENGN | 1 | 100% | 0.5% | 2 |
| 4 | 4 RUE BLAISE PASCAL | 1 | 50% | 0.2% | 1 |
| 5 | BIOL OUCES FUNCT | 1 | 50% | 0.2% | 1 |
| 6 | CNRS GENET MOL GENOM MICROBIOL | 1 | 50% | 0.2% | 1 |
| 7 | DYNAM EVOLUT EXP S GENOME MICROORG | 1 | 50% | 0.2% | 1 |
| 8 | MOL MED PHARMACOLMOL BIOL | 1 | 50% | 0.2% | 1 |
| 9 | UMR792 INGN SYST BIOL PROCEDESUMR5504 | 1 | 50% | 0.2% | 1 |
| 10 | METAB SIGNALING REGULAT GRP | 1 | 25% | 0.5% | 2 |
Related classes at same level (level 1) |
| Rank | Relatedness score | Related classes |
|---|---|---|
| 1 | 0.0000160070 | RIBOSWITCH//RIBOSWITCHES//STREPTOMYCES DAVAWENSIS |
| 2 | 0.0000158141 | DIHYDROOROTASE//CARBAMOYL PHOSPHATE SYNTHETASE//ASPARTATE CARBAMOYLTRANSFERASE |
| 3 | 0.0000150210 | ARGININE REPRESSOR//ARGININE REPRESSOR PROTEIN//ARGR |
| 4 | 0.0000143747 | PHOSPHORIBOSYLPYROPHOSPHATE SYNTHETASE//PHOSPHORIBOSYL PYROPHOSPHATE SYNTHETASE//ARTHROBACTER A302 |
| 5 | 0.0000131987 | PHOSPHORIBOSYLTRANSFERASE//MOL PARASITOL DRUG DESIGN//HGXPRT |
| 6 | 0.0000127904 | PERCHLORIC ACID SOLUBLE PROTEIN//G SISINI//UK101 |
| 7 | 0.0000110577 | CYCLOPENTENYL CYTOSINE//CTP SYNTHASE//CTP SYNTHETASE |
| 8 | 0.0000109205 | NUSG//TRANSCRIPTION TERMINATION//RHO FACTOR |
| 9 | 0.0000096155 | ADENYLOSUCCINATE LYASE DEFICIENCY//ADENYLOSUCCINATE LYASE//PURINE BIOSYNTHESIS |
| 10 | 0.0000086956 | TERMINATION OF REPLICATION//REPLICATION TERMINATOR PROTEIN//FORK ARREST |