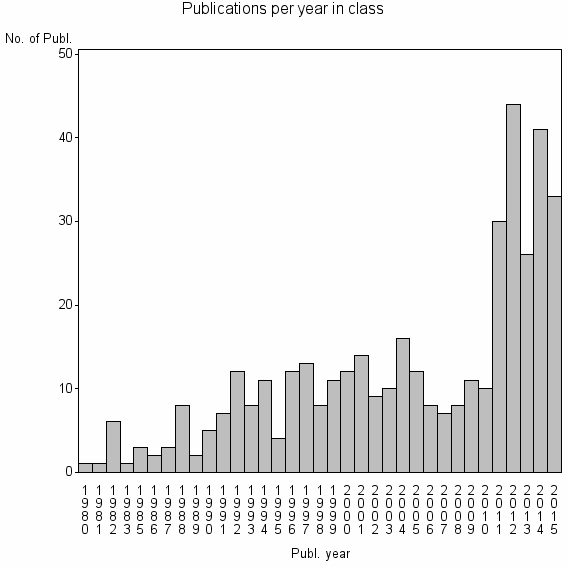
Class information for: |
Basic class information |
| ID | Publications | Average number of references |
Avg. shr. active ref. in WoS |
|---|---|---|---|
| 20181 | 409 | 38.3 | 75% |
Classes in level above (level 2) |
| ID, lev. above |
Publications | Label for level above |
|---|---|---|
| 946 | 10347 | LACCASE//MANGANESE PEROXIDASE//WHITE ROT FUNGI |
Terms with highest relevance score |
| Rank | Term | Type of term | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
|---|---|---|---|---|---|---|
| 1 | CELLOBIOSE DEHYDROGENASE | Author keyword | 137 | 73% | 26% | 105 |
| 2 | AA9 | Author keyword | 17 | 100% | 2% | 8 |
| 3 | GH61 | Author keyword | 15 | 68% | 3% | 13 |
| 4 | CELLOBIOSE OXIDASE | Author keyword | 13 | 80% | 2% | 8 |
| 5 | LYTIC POLYSACCHARIDE MONOOXYGENASE | Author keyword | 9 | 83% | 1% | 5 |
| 6 | CBM33 | Author keyword | 9 | 67% | 2% | 8 |
| 7 | LACTOBIONIC ACID | Author keyword | 7 | 29% | 5% | 20 |
| 8 | CELLOBIOSE QUINONE OXIDOREDUCTASE | Author keyword | 6 | 80% | 1% | 4 |
| 9 | LYTIC POLYSACCHARIDE MONOOXYGENASES | Author keyword | 6 | 71% | 1% | 5 |
| 10 | LYTIC POLYSACCHARIDE MONOOXYGENASE LPMO | Author keyword | 6 | 100% | 1% | 4 |
Web of Science journal categories |
Author Key Words |
| Rank | Web of Science journal category | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
LCSH search | Wikipedia search |
|---|---|---|---|---|---|---|---|
| 1 | CELLOBIOSE DEHYDROGENASE | 137 | 73% | 26% | 105 | Search CELLOBIOSE+DEHYDROGENASE | Search CELLOBIOSE+DEHYDROGENASE |
| 2 | AA9 | 17 | 100% | 2% | 8 | Search AA9 | Search AA9 |
| 3 | GH61 | 15 | 68% | 3% | 13 | Search GH61 | Search GH61 |
| 4 | CELLOBIOSE OXIDASE | 13 | 80% | 2% | 8 | Search CELLOBIOSE+OXIDASE | Search CELLOBIOSE+OXIDASE |
| 5 | LYTIC POLYSACCHARIDE MONOOXYGENASE | 9 | 83% | 1% | 5 | Search LYTIC+POLYSACCHARIDE+MONOOXYGENASE | Search LYTIC+POLYSACCHARIDE+MONOOXYGENASE |
| 6 | CBM33 | 9 | 67% | 2% | 8 | Search CBM33 | Search CBM33 |
| 7 | LACTOBIONIC ACID | 7 | 29% | 5% | 20 | Search LACTOBIONIC+ACID | Search LACTOBIONIC+ACID |
| 8 | CELLOBIOSE QUINONE OXIDOREDUCTASE | 6 | 80% | 1% | 4 | Search CELLOBIOSE+QUINONE+OXIDOREDUCTASE | Search CELLOBIOSE+QUINONE+OXIDOREDUCTASE |
| 9 | LYTIC POLYSACCHARIDE MONOOXYGENASES | 6 | 71% | 1% | 5 | Search LYTIC+POLYSACCHARIDE+MONOOXYGENASES | Search LYTIC+POLYSACCHARIDE+MONOOXYGENASES |
| 10 | LYTIC POLYSACCHARIDE MONOOXYGENASE LPMO | 6 | 100% | 1% | 4 | Search LYTIC+POLYSACCHARIDE+MONOOXYGENASE+LPMO | Search LYTIC+POLYSACCHARIDE+MONOOXYGENASE+LPMO |
Key Words Plus |
| Rank | Web of Science journal category | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
|---|---|---|---|---|---|
| 1 | SPOROTRICHUM PULVERULENTUM | 32 | 43% | 14% | 58 |
| 2 | FUNGUS SPOROTRICHUM PULVERULENTUM | 28 | 60% | 8% | 31 |
| 3 | CELLOBIOSE DEHYDROGENASE | 27 | 31% | 18% | 73 |
| 4 | GLYCOSIDE HYDROLASE FAMILY | 22 | 39% | 11% | 45 |
| 5 | HEMOFLAVOENZYME | 17 | 100% | 2% | 8 |
| 6 | MYRIOCOCCUM THERMOPHILUM | 14 | 60% | 4% | 15 |
| 7 | WOOD DEGRADING ENZYME | 14 | 100% | 2% | 7 |
| 8 | LYTIC POLYSACCHARIDE MONOOXYGENASES | 9 | 83% | 1% | 5 |
| 9 | FLAVOCYTOCHROME | 9 | 29% | 6% | 26 |
| 10 | SCLEROTIUM ATHELIA ROLFSII | 8 | 70% | 2% | 7 |
Journals |
Reviews |
| Title | Publ. year | Cit. | Active references | % act. ref. to same field |
|---|---|---|---|---|
| A critical review of cellobiose dehydrogenases | 2000 | 203 | 85 | 76% |
| Cellobiose dehydrogenase - A flavocytochrome from wood-degrading, phytopathogenic and saprotropic fungi | 2006 | 96 | 126 | 68% |
| Novel enzymes for the degradation of cellulose | 2012 | 113 | 87 | 25% |
| Cellobiose dehydrogenase - an extracellular fungal flavocytochrome | 2001 | 59 | 54 | 81% |
| Cellulose Degradation by Polysaccharide Monooxygenases | 2015 | 1 | 92 | 47% |
| Cellobiose dehydrogenase modified electrodes: advances by materials science and biochemical engineering | 2013 | 12 | 144 | 45% |
| Direct electron transfer between ligninolytic redox enzymes and electrodes | 2004 | 67 | 152 | 29% |
| Recalcitrant polysaccharide degradation by novel oxidative biocatalysts | 2013 | 8 | 59 | 51% |
| New enzyme insights drive advances in commercial ethanol production | 2014 | 14 | 60 | 18% |
| Fungal polysaccharide monooxygenases: new players in the decomposition of cellulose | 2012 | 14 | 47 | 38% |
Address terms |
| Rank | Address term | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
|---|---|---|---|---|---|
| 1 | ANALYT CHEM BIOCHEM STRUCT BIOL | 4 | 33% | 2.4% | 10 |
| 2 | BCF UMR1163 | 2 | 44% | 1.0% | 4 |
| 3 | CELL MOL BIOL STRUCT BIOL | 2 | 67% | 0.5% | 2 |
| 4 | ESIL POLYTECH MARSEILLE | 2 | 67% | 0.5% | 2 |
| 5 | ANALYT CHEM BIOCHEM | 1 | 27% | 0.7% | 3 |
| 6 | GRP ENVIRONM ORGAN CHEM TECHNOL ENVOC | 1 | 25% | 0.7% | 3 |
| 7 | BIOTECHNOL CHAMPIGNONS FILAMENTEUX UMR1163 | 1 | 21% | 0.7% | 3 |
| 8 | AGR LIFE SIC | 1 | 50% | 0.2% | 1 |
| 9 | ARCHITECTURE FONCT MACROMOL BIOL UMR7257 | 1 | 50% | 0.2% | 1 |
| 10 | ECOL SUELOS HONGOS TROP | 1 | 50% | 0.2% | 1 |
Related classes at same level (level 1) |
| Rank | Relatedness score | Related classes |
|---|---|---|
| 1 | 0.0000196185 | BROWN ROT DECAY//BROWN ROT FUNGI//GLYOXYLATE OXIDASE |
| 2 | 0.0000189981 | 1 5 ANHYDRO D FRUCTOSE//1 5 ANHYDRO D GLUCITOL//PYRANOSE DEHYDROGENASE |
| 3 | 0.0000168683 | ENDOGLUCANASE//CELLOBIOHYDROLASE//CELLULASE |
| 4 | 0.0000151946 | BIOFUEL CELL//BIOFUEL CELLS//BILIRUBIN OXIDASE |
| 5 | 0.0000119384 | LACCASE//MANGANESE PEROXIDASE//LIGNIN PEROXIDASE |
| 6 | 0.0000095587 | XLNR//AMYR//TRICHODERMA REESEI |
| 7 | 0.0000082219 | LACTOSE DETERMINATION//OCTADECYLSILICA COLUMN//LACTOSE BIOSENSOR |
| 8 | 0.0000081315 | CYTOCHROME P450NOR//FUNGAL DENITRIFICATION//P450NOR |
| 9 | 0.0000079885 | FLAVOCYTOCHROME B2//TORULOPSIS GLABRATA//FRE 2930 |
| 10 | 0.0000063883 | GLUCURONOYL ESTERASE//4 O METHYL D GLUCURONIC ACID//HALOXYLON SALICORNICUM |