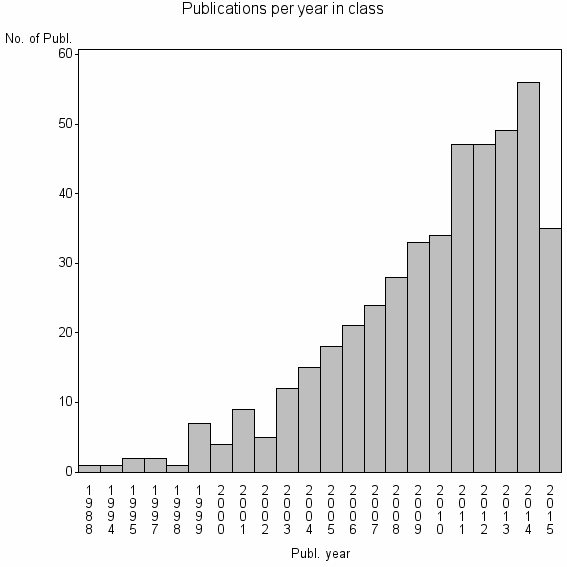
Class information for: |
Basic class information |
| ID | Publications | Average number of references |
Avg. shr. active ref. in WoS |
|---|---|---|---|
| 19200 | 451 | 44.7 | 88% |
Classes in level above (level 2) |
| ID, lev. above |
Publications | Label for level above |
|---|---|---|
| 42 | 31225 | PROTEINS-STRUCTURE FUNCTION AND BIOINFORMATICS//PROTEIN STRUCTURE PREDICTION//PROTEIN FOLDING |
Terms with highest relevance score |
| Rank | Term | Type of term | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
|---|---|---|---|---|---|---|
| 1 | INTERACTION PROPENSITY | Author keyword | 8 | 100% | 1% | 5 |
| 2 | RNA BINDING RESIDUES | Author keyword | 4 | 75% | 1% | 3 |
| 3 | COMPUTAT INTELLIGENCE LEARNING DISCOVERY | Address | 3 | 50% | 1% | 4 |
| 4 | DNA BINDING RESIDUES | Author keyword | 2 | 67% | 0% | 2 |
| 5 | INVARIANT CONTACTS | Author keyword | 2 | 67% | 0% | 2 |
| 6 | VARIABLE CONTACTS | Author keyword | 2 | 67% | 0% | 2 |
| 7 | GOLDEN AUDIT | Address | 2 | 43% | 1% | 3 |
| 8 | CROSS LINKING AND IMMUNOPRECIPITATION | Author keyword | 1 | 100% | 0% | 2 |
| 9 | DNA PROTEIN RECOGNITION MODULE | Author keyword | 1 | 100% | 0% | 2 |
| 10 | PROTEIN RNA DOCKING | Author keyword | 1 | 100% | 0% | 2 |
Web of Science journal categories |
Author Key Words |
Key Words Plus |
| Rank | Web of Science journal category | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
|---|---|---|---|---|---|
| 1 | ALTERNATING CHARGE CLUSTERS | 14 | 100% | 2% | 7 |
| 2 | EFFICIENT PREDICTION | 11 | 69% | 2% | 9 |
| 3 | PAR CLIP | 10 | 35% | 5% | 24 |
| 4 | NUCLEIC ACID RECOGNITION | 5 | 21% | 5% | 21 |
| 5 | STRUCTURAL INFORMATION | 3 | 10% | 6% | 26 |
| 6 | RIBOSOMAL RNA BINDING | 3 | 38% | 1% | 6 |
| 7 | NUCLEOTIDE RESOLUTION | 3 | 10% | 5% | 24 |
| 8 | STRUCTURE BASED PREDICTION | 2 | 18% | 2% | 11 |
| 9 | HITS CLIP | 2 | 36% | 1% | 5 |
| 10 | ALPHA SHAPE | 2 | 67% | 0% | 2 |
Journals |
Reviews |
| Title | Publ. year | Cit. | Active references | % act. ref. to same field |
|---|---|---|---|---|
| The "Observer Effect" in Genome-wide Surveys of Protein-RNA Interactions | 2013 | 14 | 25 | 32% |
| Prediction of RNA binding proteins comes of age from low resolution to high resolution | 2013 | 5 | 63 | 52% |
| Methods for comprehensive experimental identification of RNA-protein interactions | 2014 | 9 | 102 | 18% |
| Computational Approaches for Predicting the Binding Sites and Understanding the Recognition Mechanism of Protein-DNA Complexes | 2013 | 1 | 100 | 56% |
| Understanding the Recognition Mechanism of Protein-RNA Complexes Using Energy Based Approach | 2010 | 3 | 34 | 53% |
| An Overview of the Prediction of Protein DNA-Binding Sites | 2015 | 0 | 94 | 50% |
| RNA-binding proteins in Mendelian disease | 2013 | 27 | 74 | 9% |
| UV Cross-Linking of Interacting RNA and Protein in Cultured Cells | 2014 | 0 | 6 | 50% |
| Protein-RNA interactions: new genomic technologies and perspectives | 2012 | 42 | 44 | 16% |
| Computational tools for protein-DNA interactions | 2012 | 1 | 87 | 46% |
Address terms |
| Rank | Address term | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
|---|---|---|---|---|---|
| 1 | COMPUTAT INTELLIGENCE LEARNING DISCOVERY | 3 | 50% | 0.9% | 4 |
| 2 | GOLDEN AUDIT | 2 | 43% | 0.7% | 3 |
| 3 | CEPBA IBM | 1 | 50% | 0.2% | 1 |
| 4 | GENET NETWORKS PROGRAM | 1 | 50% | 0.2% | 1 |
| 5 | GENOME OURCE ANAL UNIT GRAS | 1 | 50% | 0.2% | 1 |
| 6 | HORMONES DEV ABORAT INNOVAT TIA | 1 | 50% | 0.2% | 1 |
| 7 | ICREA PROF | 1 | 50% | 0.2% | 1 |
| 8 | INTEGRAT TRANSLAT GEN | 1 | 50% | 0.2% | 1 |
| 9 | OKINAWA SCI | 1 | 50% | 0.2% | 1 |
| 10 | QUIM BIOL FISIOL | 1 | 50% | 0.2% | 1 |
Related classes at same level (level 1) |
| Rank | Relatedness score | Related classes |
|---|---|---|
| 1 | 0.0000183342 | CAPRI//PROTEIN PROTEIN DOCKING//PROTEIN DOCKING |
| 2 | 0.0000137003 | COMPLEX SCI SOFTWARE//3D ZERNIKE DESCRIPTOR//PROTEIN SURFACE SHAPE |
| 3 | 0.0000120154 | PSEUDO AMINO ACID COMPOSITION//COMPUTAT MUTAT PROJECT//JACKKNIFE TEST |
| 4 | 0.0000118204 | ARTIFICIAL TRANSCRIPTION FACTOR//ARTIFICIAL TRANSCRIPTION FACTORS//RECOGNITION CODE |
| 5 | 0.0000117816 | MOTIF DISCOVERY//CLOSEST STRING PROBLEM//MOTIF FINDING |
| 6 | 0.0000085497 | CYTOPLASMIC POLYADENYLATION//PUMILIO//CPEB |
| 7 | 0.0000084562 | HUR//TRISTETRAPROLIN//AU RICH ELEMENT |
| 8 | 0.0000078209 | DNA CURVATURE//A TRACT//DNA FLEXIBILITY |
| 9 | 0.0000076521 | HNRNP B1//HETEROGENEOUS NUCLEAR RIBONUCLEOPROTEINS HNRNPS//RNP CONCENSUS RNA BINDING DOMAIN |
| 10 | 0.0000075307 | LIN28//LIN28B//LIN28A |