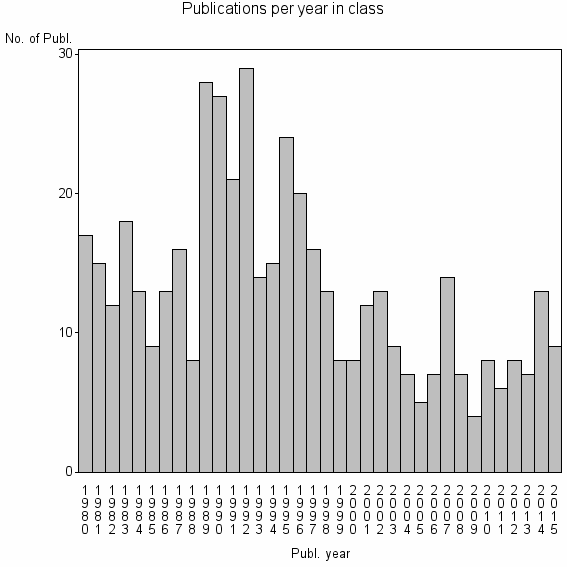
Class information for: |
Basic class information |
| ID | Publications | Average number of references |
Avg. shr. active ref. in WoS |
|---|---|---|---|
| 18702 | 473 | 37.1 | 59% |
Classes in level above (level 2) |
| ID, lev. above |
Publications | Label for level above |
|---|---|---|
| 501 | 14604 | RNA POLYMERASE//SITE SPECIFIC RECOMBINATION//H NS |
Terms with highest relevance score |
| Rank | Term | Type of term | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
|---|---|---|---|---|---|---|
| 1 | SKIRBALL PROGRAM MOL PATHOGENESIS | Address | 19 | 74% | 3% | 14 |
| 2 | BACTERIOPHAGE P2 | Author keyword | 12 | 86% | 1% | 6 |
| 3 | RETRON | Author keyword | 11 | 67% | 2% | 10 |
| 4 | MSDNA | Author keyword | 8 | 75% | 1% | 6 |
| 5 | PHAGE P4 | Author keyword | 3 | 100% | 1% | 3 |
| 6 | MOLECULAR PIRACY | Author keyword | 2 | 44% | 1% | 4 |
| 7 | SAPI | Author keyword | 2 | 44% | 1% | 4 |
| 8 | QUIM BIOQUIM BIOL MOL | Address | 2 | 18% | 2% | 10 |
| 9 | PROCAPSID | Author keyword | 2 | 25% | 1% | 6 |
| 10 | HELPER PHAGE | Author keyword | 2 | 43% | 1% | 3 |
Web of Science journal categories |
Author Key Words |
| Rank | Web of Science journal category | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
LCSH search | Wikipedia search |
|---|---|---|---|---|---|---|---|
| 1 | BACTERIOPHAGE P2 | 12 | 86% | 1% | 6 | Search BACTERIOPHAGE+P2 | Search BACTERIOPHAGE+P2 |
| 2 | RETRON | 11 | 67% | 2% | 10 | Search RETRON | Search RETRON |
| 3 | MSDNA | 8 | 75% | 1% | 6 | Search MSDNA | Search MSDNA |
| 4 | PHAGE P4 | 3 | 100% | 1% | 3 | Search PHAGE+P4 | Search PHAGE+P4 |
| 5 | MOLECULAR PIRACY | 2 | 44% | 1% | 4 | Search MOLECULAR+PIRACY | Search MOLECULAR+PIRACY |
| 6 | SAPI | 2 | 44% | 1% | 4 | Search SAPI | Search SAPI |
| 7 | PROCAPSID | 2 | 25% | 1% | 6 | Search PROCAPSID | Search PROCAPSID |
| 8 | HELPER PHAGE | 2 | 43% | 1% | 3 | Search HELPER+PHAGE | Search HELPER+PHAGE |
| 9 | BACTERIOPHAGE P2 P4 | 1 | 100% | 0% | 2 | Search BACTERIOPHAGE+P2+P4 | Search BACTERIOPHAGE+P2+P4 |
| 10 | MULTICOPY SINGLE STRANDED DNA | 1 | 100% | 0% | 2 | Search MULTICOPY+SINGLE+STRANDED+DNA | Search MULTICOPY+SINGLE+STRANDED+DNA |
Key Words Plus |
| Rank | Web of Science journal category | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
|---|---|---|---|---|---|
| 1 | SATELLITE BACTERIOPHAGE P4 | 46 | 75% | 7% | 33 |
| 2 | BRANCHED RNA | 37 | 73% | 6% | 29 |
| 3 | RNA LINKED MSDNA | 31 | 71% | 5% | 25 |
| 4 | MSDNA | 26 | 80% | 3% | 16 |
| 5 | PATHOGENICITY ISLAND SAPI1 | 24 | 91% | 2% | 10 |
| 6 | TEMPERATE COLIPHAGE 186 | 21 | 90% | 2% | 9 |
| 7 | SATELLITE PHAGE P4 | 20 | 100% | 2% | 9 |
| 8 | C1 REPRESSOR | 15 | 82% | 2% | 9 |
| 9 | CDNA PRIMING REACTION | 14 | 100% | 1% | 7 |
| 10 | GENETIC ELEMENT P4 | 14 | 100% | 1% | 7 |
Journals |
Reviews |
| Title | Publ. year | Cit. | Active references | % act. ref. to same field |
|---|---|---|---|---|
| The phage-related chromosomal islands of Gram-positive bacteria | 2010 | 87 | 39 | 38% |
| Pirates of the Caudovirales | 2012 | 12 | 100 | 60% |
| Evolution of P2-like phages and their impact on bacterial evolution | 2007 | 15 | 31 | 74% |
| MECHANISMS OF GENOME PROPAGATION AND HELPER EXPLOITATION BY SATELLITE PHAGE-P4 | 1993 | 81 | 132 | 71% |
| Bacteriophage-mediated spread of bacterial virulence genes | 2015 | 2 | 46 | 37% |
| The plasmid status of satellite bacteriophage P4 | 2001 | 31 | 80 | 64% |
| The msDNAs of bacteria | 2001 | 12 | 61 | 84% |
| Genome of bacteriophage P1 | 2004 | 71 | 280 | 20% |
| BACTERIOPHAGE-MEDIATED GENERALIZED TRANSDUCTION IN ESCHERICHIA-COLI AND SALMONELLA-TYPHIMURIUM | 1991 | 145 | 21 | 33% |
| REVERSE TRANSCRIPTION IN BACTERIA | 1989 | 36 | 10 | 80% |
Address terms |
| Rank | Address term | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
|---|---|---|---|---|---|
| 1 | SKIRBALL PROGRAM MOL PATHOGENESIS | 19 | 74% | 3.0% | 14 |
| 2 | QUIM BIOQUIM BIOL MOL | 2 | 18% | 2.1% | 10 |
| 3 | AGROBIOTECNOL RECURSOS NAT | 1 | 15% | 1.3% | 6 |
| 4 | EDIFICIO SEMINARIO | 1 | 50% | 0.2% | 1 |
| 5 | JET PROP 125 224 | 1 | 50% | 0.2% | 1 |
| 6 | DIPARTIMENTO GENET BIOL MICROORGANISMI | 0 | 11% | 0.8% | 4 |
| 7 | INVEST TECNOL ANIM | 0 | 11% | 0.8% | 4 |
| 8 | EXPT MARINE BIOL OCEANOL | 0 | 33% | 0.2% | 1 |
| 9 | INFECT DIS GENOM | 0 | 33% | 0.2% | 1 |
| 10 | MOL BIOSCI BIOCHEM | 0 | 17% | 0.4% | 2 |
Related classes at same level (level 1) |
| Rank | Relatedness score | Related classes |
|---|---|---|
| 1 | 0.0000130312 | SITE SPECIFIC RECOMBINATION//SERINE RECOMBINASE//PHI C31 INTEGRASE |
| 2 | 0.0000128342 | DNA PACKAGING//BACTERIOPHAGE P22//PORTAL PROTEIN |
| 3 | 0.0000110375 | REX EXCLUSION//BACTERIOPHAGE LAMBDA LAMBDA//CI REXA REXB OPERON |
| 4 | 0.0000083711 | REPLICATION CONTROL//PLASMID R6K//PLASMID RK2 |
| 5 | 0.0000072954 | TRP REPRESSOR//LAC REPRESSOR//LACTOSE REPRESSOR |
| 6 | 0.0000069258 | EPIDEMIOLOGICAL SIGNIFICANCE//SIMILARITY IDENTIFICATION//AIRBORNE ESCHERICHIA COLI |
| 7 | 0.0000067277 | POLYLINKER//STREPTOMYCIN SENSITIVITY//INSERTIONAL INACTIVATION |
| 8 | 0.0000062372 | TERMINATION OF REPLICATION//REPLICATION TERMINATOR PROTEIN//FORK ARREST |
| 9 | 0.0000058881 | BACTERIOPHAGE ALPHA 3//MICROVIRUS//BACTERIOPHAGE PHI X174 |
| 10 | 0.0000056019 | PHAGE THERAPY//BACTERIOPHAGE//BACTERIOPHAGE THERAPY |