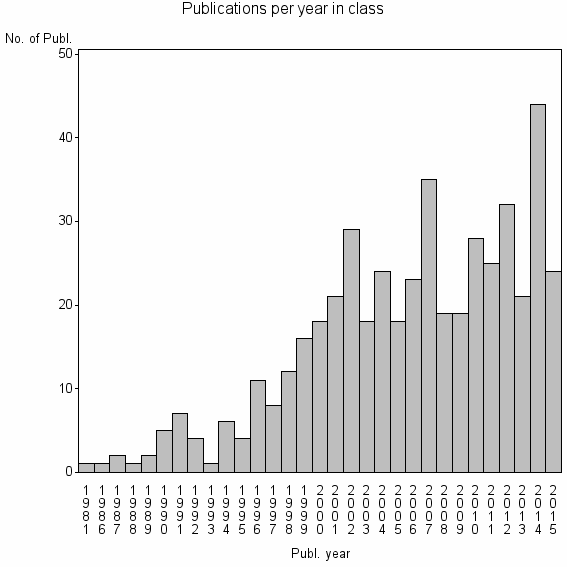
Class information for: |
Basic class information |
| ID | Publications | Average number of references |
Avg. shr. active ref. in WoS |
|---|---|---|---|
| 18592 | 479 | 41.3 | 87% |
Classes in level above (level 2) |
Terms with highest relevance score |
| Rank | Term | Type of term | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
|---|---|---|---|---|---|---|
| 1 | TRANS TRANSLATION | Author keyword | 161 | 91% | 14% | 68 |
| 2 | TMRNA | Author keyword | 155 | 82% | 19% | 89 |
| 3 | SMPB | Author keyword | 73 | 86% | 8% | 37 |
| 4 | 10SA RNA | Author keyword | 41 | 87% | 4% | 20 |
| 5 | PEPTIDYL TRNA HYDROLASE | Author keyword | 35 | 81% | 4% | 21 |
| 6 | SSRA | Author keyword | 18 | 65% | 4% | 17 |
| 7 | PELOTA | Author keyword | 11 | 78% | 1% | 7 |
| 8 | UP JE 2311 | Address | 7 | 67% | 1% | 6 |
| 9 | NONSTOP MRNA | Author keyword | 6 | 80% | 1% | 4 |
| 10 | UP JE2311 | Address | 6 | 80% | 1% | 4 |
Web of Science journal categories |
Author Key Words |
| Rank | Web of Science journal category | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
LCSH search | Wikipedia search |
|---|---|---|---|---|---|---|---|
| 1 | TRANS TRANSLATION | 161 | 91% | 14% | 68 | Search TRANS+TRANSLATION | Search TRANS+TRANSLATION |
| 2 | TMRNA | 155 | 82% | 19% | 89 | Search TMRNA | Search TMRNA |
| 3 | SMPB | 73 | 86% | 8% | 37 | Search SMPB | Search SMPB |
| 4 | 10SA RNA | 41 | 87% | 4% | 20 | Search 10SA+RNA | Search 10SA+RNA |
| 5 | PEPTIDYL TRNA HYDROLASE | 35 | 81% | 4% | 21 | Search PEPTIDYL+TRNA+HYDROLASE | Search PEPTIDYL+TRNA+HYDROLASE |
| 6 | SSRA | 18 | 65% | 4% | 17 | Search SSRA | Search SSRA |
| 7 | PELOTA | 11 | 78% | 1% | 7 | Search PELOTA | Search PELOTA |
| 8 | NONSTOP MRNA | 6 | 80% | 1% | 4 | Search NONSTOP+MRNA | Search NONSTOP+MRNA |
| 9 | ARFA | 6 | 71% | 1% | 5 | Search ARFA | Search ARFA |
| 10 | RIBOSOME RESCUE | 6 | 71% | 1% | 5 | Search RIBOSOME+RESCUE | Search RIBOSOME+RESCUE |
Key Words Plus |
| Rank | Web of Science journal category | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
|---|---|---|---|---|---|
| 1 | 10SA RNA | 180 | 79% | 24% | 117 |
| 2 | ESCHERICHIA COLI TMRNA | 158 | 98% | 9% | 42 |
| 3 | TRANSFER MESSENGER RNA | 146 | 86% | 15% | 74 |
| 4 | STALLED RIBOSOME | 112 | 97% | 7% | 32 |
| 5 | SMALL STABLE RNA | 98 | 84% | 11% | 53 |
| 6 | TRANS TRANSLATION | 92 | 71% | 16% | 75 |
| 7 | SMALL PROTEIN B | 79 | 91% | 7% | 32 |
| 8 | SMPB | 66 | 83% | 8% | 38 |
| 9 | RIBOSOME RESCUE | 63 | 71% | 11% | 51 |
| 10 | TMRNA | 52 | 64% | 11% | 51 |
Journals |
Reviews |
| Title | Publ. year | Cit. | Active references | % act. ref. to same field |
|---|---|---|---|---|
| The tmRNA system for translational surveillance and ribosome rescue | 2007 | 151 | 131 | 70% |
| Biology of trans-Translation | 2008 | 89 | 108 | 73% |
| Ribosome rescue systems in bacteria | 2015 | 2 | 144 | 74% |
| Trans-translation: The tmRNA-mediated surveillance mechanism for ribosome rescue, directed protein degradation, and nonstop mRNA decay | 2007 | 44 | 96 | 74% |
| The task force that rescues stalled ribosomes in bacteria | 2013 | 12 | 77 | 75% |
| Protecting the proteome: Eukaryotic cotranslational quality control pathways | 2014 | 15 | 98 | 32% |
| A salvage pathway for protein synthesis: tmRNA and trans-translation | 2003 | 103 | 100 | 69% |
| The SsrA-SmpB system for protein tagging, directed degradation and ribosome rescue | 2000 | 240 | 62 | 48% |
| Quality control systems for aberrant mRNAs induced by aberrant translation elongation and termination | 2013 | 17 | 91 | 55% |
| Beyond ribosome rescue: tmRNA and co-translational processes | 2010 | 34 | 63 | 67% |
Address terms |
| Rank | Address term | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
|---|---|---|---|---|---|
| 1 | UP JE 2311 | 7 | 67% | 1.3% | 6 |
| 2 | UP JE2311 | 6 | 80% | 0.8% | 4 |
| 3 | UP JEUNE EQUIPE 2311 | 4 | 75% | 0.6% | 3 |
| 4 | BIOCHIM PHARMACEUT | 4 | 38% | 1.7% | 8 |
| 5 | UP JEUNE EQUIPE 2311IFR 97 | 3 | 100% | 0.6% | 3 |
| 6 | ALTHOUSE 401 | 2 | 67% | 0.4% | 2 |
| 7 | ANTENNE CHIM SUBST NAT | 2 | 67% | 0.4% | 2 |
| 8 | EQUIPE STRUCT DYNAM MACROMOL | 1 | 100% | 0.4% | 2 |
| 9 | TRANSLAT FOLDING TEAM | 1 | 50% | 0.4% | 2 |
| 10 | UMR 7654 CNRS | 1 | 50% | 0.4% | 2 |
Related classes at same level (level 1) |
| Rank | Relatedness score | Related classes |
|---|---|---|
| 1 | 0.0000142365 | TRIGGER FACTOR//NASCENT CHAIN//COTRANSLATIONAL FOLDING |
| 2 | 0.0000128356 | FRAMESHIFTING//RIBOSOMAL FRAMESHIFTING//RNA PSEUDOKNOT |
| 3 | 0.0000126966 | ELONGATION FACTOR TU//EF G//ELONGATION FACTOR G |
| 4 | 0.0000116780 | RNASE E//RNASE G//POLYNUCLEOTIDE PHOSPHORYLASE |
| 5 | 0.0000112554 | ATP DEPENDENT PROTEASES//CLPB//FTSH |
| 6 | 0.0000108336 | SD SEQUENCE//RIBOSOMAL PROTEIN S1//SHINE DALGARNO SEQUENCE |
| 7 | 0.0000089871 | SUP35//TRANSLATION TERMINATION//URE3 |
| 8 | 0.0000074809 | UPF1//NMD//NONSENSE MEDIATED MRNA DECAY |
| 9 | 0.0000071568 | DAP3//ICT1//MITOCHONDRIAL RIBOSOMAL PROTEINS |
| 10 | 0.0000069279 | TOXIN ANTITOXIN//TOXIN ANTITOXIN SYSTEM//TOXIN ANTITOXIN SYSTEMS |