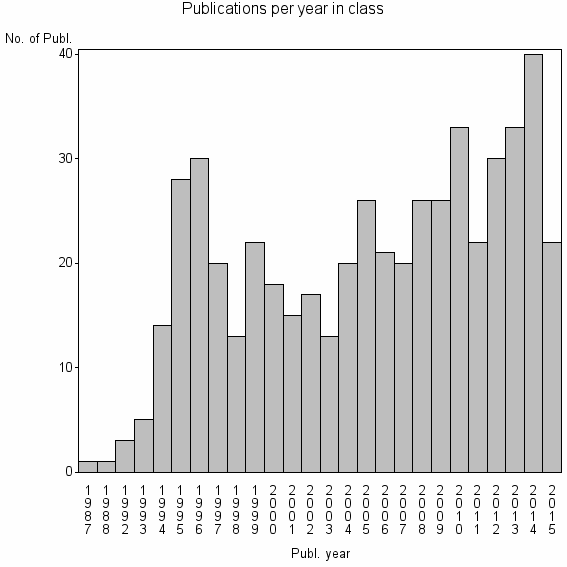
Class information for: |
Basic class information |
| ID | Publications | Average number of references |
Avg. shr. active ref. in WoS |
|---|---|---|---|
| 17791 | 519 | 46.6 | 94% |
Classes in level above (level 2) |
| ID, lev. above |
Publications | Label for level above |
|---|---|---|
| 313 | 17635 | X LINKED AGAMMAGLOBULINEMIA//BRUTONS TYROSINE KINASE//BTK |
Terms with highest relevance score |
| Rank | Term | Type of term | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
|---|---|---|---|---|---|---|
| 1 | P66SHC | Author keyword | 82 | 65% | 15% | 78 |
| 2 | SHC | Author keyword | 9 | 21% | 7% | 38 |
| 3 | SHC FAMILY | Author keyword | 6 | 80% | 1% | 4 |
| 4 | SHCA | Author keyword | 6 | 50% | 2% | 9 |
| 5 | SHC PROTEINS | Author keyword | 6 | 100% | 1% | 4 |
| 6 | SHC MOLECULES | Author keyword | 3 | 100% | 1% | 3 |
| 7 | MONTELUCE POLICLIN IST CLIN MED 1 | Address | 2 | 67% | 0% | 2 |
| 8 | N SHC | Author keyword | 2 | 67% | 0% | 2 |
| 9 | SHC3 | Author keyword | 2 | 67% | 0% | 2 |
| 10 | INTERDISCIPLINARY PROGRAM GENET GENOM | Address | 1 | 50% | 0% | 2 |
Web of Science journal categories |
Author Key Words |
| Rank | Web of Science journal category | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
LCSH search | Wikipedia search |
|---|---|---|---|---|---|---|---|
| 1 | P66SHC | 82 | 65% | 15% | 78 | Search P66SHC | Search P66SHC |
| 2 | SHC | 9 | 21% | 7% | 38 | Search SHC | Search SHC |
| 3 | SHC FAMILY | 6 | 80% | 1% | 4 | Search SHC+FAMILY | Search SHC+FAMILY |
| 4 | SHCA | 6 | 50% | 2% | 9 | Search SHCA | Search SHCA |
| 5 | SHC PROTEINS | 6 | 100% | 1% | 4 | Search SHC+PROTEINS | Search SHC+PROTEINS |
| 6 | SHC MOLECULES | 3 | 100% | 1% | 3 | Search SHC+MOLECULES | Search SHC+MOLECULES |
| 7 | N SHC | 2 | 67% | 0% | 2 | Search N+SHC | Search N+SHC |
| 8 | SHC3 | 2 | 67% | 0% | 2 | Search SHC3 | Search SHC3 |
| 9 | POLYOMA VIRUS MIDDLE T | 1 | 50% | 0% | 2 | Search POLYOMA+VIRUS+MIDDLE+T | Search POLYOMA+VIRUS+MIDDLE+T |
| 10 | SHC1 | 1 | 50% | 0% | 2 | Search SHC1 | Search SHC1 |
Key Words Plus |
| Rank | Web of Science journal category | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
|---|---|---|---|---|---|
| 1 | P66SHC | 36 | 58% | 8% | 42 |
| 2 | LONGEVITY GENE | 27 | 74% | 4% | 20 |
| 3 | 66 KDA SHC ISOFORM | 24 | 82% | 3% | 14 |
| 4 | N SHC | 24 | 82% | 3% | 14 |
| 5 | P66SHC PROTEIN | 15 | 82% | 2% | 9 |
| 6 | GENE PROTECTS | 15 | 88% | 1% | 7 |
| 7 | P66SHC GENE | 10 | 73% | 2% | 8 |
| 8 | FORKHEAD PROTEINS | 8 | 48% | 2% | 12 |
| 9 | SRC HOMOLOGY | 7 | 67% | 1% | 6 |
| 10 | P66 SHC | 6 | 80% | 1% | 4 |
Journals |
Reviews |
| Title | Publ. year | Cit. | Active references | % act. ref. to same field |
|---|---|---|---|---|
| Teaching an old dogma new tricks: twenty years of Shc adaptor signalling | 2012 | 15 | 150 | 70% |
| Apoptosis and aging: Role of p66(Shc) Redox protein | 2006 | 56 | 47 | 64% |
| Apoptosis and Oxidative Stress-Related Diseases: The p66Shc Connection | 2009 | 21 | 48 | 75% |
| The Lifespan-Regulator p66Shc in Mitochondria: Redox Enzyme or Redox Sensor? | 2010 | 29 | 78 | 41% |
| P66Shc signals to age | 2009 | 33 | 47 | 47% |
| The Role of p66shc in Oxidative Stress and Apoptosis | 2010 | 7 | 52 | 79% |
| p66Shc, Mitochondria, and the Generation of Reactive Oxygen Species | 2013 | 4 | 30 | 73% |
| Molecular pathways of arterial aging | 2015 | 2 | 100 | 20% |
| Nitric Oxide, Oxidative Stress, and p66(Shc) Interplay in Diabetic Endothelial Dysfunction | 2014 | 6 | 148 | 25% |
| Final common molecular pathways of aging and cardiovascular disease - Role of the p66(Shc) protein | 2008 | 52 | 52 | 29% |
Address terms |
| Rank | Address term | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
|---|---|---|---|---|---|
| 1 | MONTELUCE POLICLIN IST CLIN MED 1 | 2 | 67% | 0.4% | 2 |
| 2 | INTERDISCIPLINARY PROGRAM GENET GENOM | 1 | 50% | 0.4% | 2 |
| 3 | PROGRAM PROTECTING BRAIN | 1 | 33% | 0.6% | 3 |
| 4 | SAMUEL LUNENFELD MOLEC DEV BIOL | 1 | 19% | 1.0% | 5 |
| 5 | SECT DEGENERAT INFLAMMATORY NEUROL DIS | 1 | 21% | 0.6% | 3 |
| 6 | BIOCHIM BIOFIS CHIM MACROMOL | 1 | 50% | 0.2% | 1 |
| 7 | EGA WOMEN HLTH | 1 | 50% | 0.2% | 1 |
| 8 | FDN POLICLIN S MATTEO IRCCS | 1 | 50% | 0.2% | 1 |
| 9 | MED SURGTRANSPLANT | 1 | 50% | 0.2% | 1 |
| 10 | MOL GENET AGING INTERVENT | 1 | 50% | 0.2% | 1 |
Related classes at same level (level 1) |
| Rank | Relatedness score | Related classes |
|---|---|---|
| 1 | 0.0000205727 | GRB2//SH2 DOMAIN//SH3 DOMAIN |
| 2 | 0.0000129207 | FE65//ALCADEIN//X11 ALPHA |
| 3 | 0.0000106857 | INSULIN RECEPTOR SUBSTRATE//INSULIN RECEPTOR SUBSTRATE 1//IRS 1 |
| 4 | 0.0000099345 | SHIP2//INOSITOL 5 PHOSPHATASE//SHIP1 |
| 5 | 0.0000079086 | METABOLIC MEMORY//HYPERGLYCEMIC MEMORY//EPIGEN PROFILING IL |
| 6 | 0.0000067564 | SLP 76//LINKER FOR ACTIVATION OF T CELLS//SECT LYMPHOCYTE SIGNALING |
| 7 | 0.0000066378 | VASCULAR AGING//REYNOLDS OKLAHOMA AGING//ENDOTHELIUM DEPENDENT DILATION |
| 8 | 0.0000063934 | ROMO1//ANTICANC MOL ONCOL//NOX 1 |
| 9 | 0.0000061734 | EPS8//MTSS1//I BAR |
| 10 | 0.0000050088 | GRB7//GRB14//GRB10 |