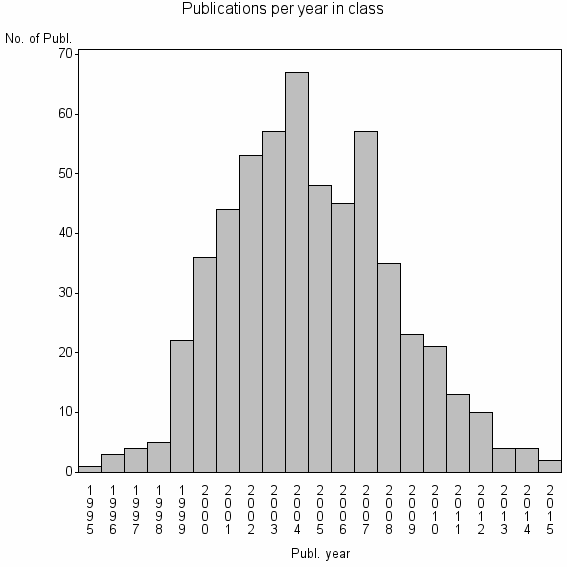
Class information for: |
Basic class information |
| ID | Publications | Average number of references |
Avg. shr. active ref. in WoS |
|---|---|---|---|
| 17090 | 554 | 35.5 | 88% |
Classes in level above (level 2) |
| ID, lev. above |
Publications | Label for level above |
|---|---|---|
| 1339 | 7890 | BIOTECHNIQUES//PCR-METHODS AND APPLICATIONS//DIFFERENTIAL DISPLAY |
Terms with highest relevance score |
| Rank | Term | Type of term | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
|---|---|---|---|---|---|---|
| 1 | SERIAL ANALYSIS OF GENE EXPRESSION | Author keyword | 12 | 28% | 7% | 38 |
| 2 | SAGE | Author keyword | 10 | 13% | 13% | 74 |
| 3 | GLGI | Author keyword | 6 | 100% | 1% | 4 |
| 4 | SERIAL ANALYSIS OF GENE EXPRESSION SAGE | Author keyword | 5 | 32% | 3% | 14 |
| 5 | SAGE TECHNIQUE | Author keyword | 3 | 100% | 1% | 3 |
| 6 | ENH | Address | 2 | 21% | 2% | 10 |
| 7 | FLUID MICROARRAY | Author keyword | 2 | 67% | 0% | 2 |
| 8 | MICROSAGE | Author keyword | 2 | 67% | 0% | 2 |
| 9 | MOL ENDOCRINOL ONCOL ANAT PHYSIO | Address | 2 | 67% | 0% | 2 |
| 10 | PATTERN DISCOVERY AND VISUALIZATION | Author keyword | 2 | 67% | 0% | 2 |
Web of Science journal categories |
Author Key Words |
| Rank | Web of Science journal category | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
LCSH search | Wikipedia search |
|---|---|---|---|---|---|---|---|
| 1 | SERIAL ANALYSIS OF GENE EXPRESSION | 12 | 28% | 7% | 38 | Search SERIAL+ANALYSIS+OF+GENE+EXPRESSION | Search SERIAL+ANALYSIS+OF+GENE+EXPRESSION |
| 2 | SAGE | 10 | 13% | 13% | 74 | Search SAGE | Search SAGE |
| 3 | GLGI | 6 | 100% | 1% | 4 | Search GLGI | Search GLGI |
| 4 | SERIAL ANALYSIS OF GENE EXPRESSION SAGE | 5 | 32% | 3% | 14 | Search SERIAL+ANALYSIS+OF+GENE+EXPRESSION+SAGE | Search SERIAL+ANALYSIS+OF+GENE+EXPRESSION+SAGE |
| 5 | SAGE TECHNIQUE | 3 | 100% | 1% | 3 | Search SAGE+TECHNIQUE | Search SAGE+TECHNIQUE |
| 6 | FLUID MICROARRAY | 2 | 67% | 0% | 2 | Search FLUID+MICROARRAY | Search FLUID+MICROARRAY |
| 7 | MICROSAGE | 2 | 67% | 0% | 2 | Search MICROSAGE | Search MICROSAGE |
| 8 | PATTERN DISCOVERY AND VISUALIZATION | 2 | 67% | 0% | 2 | Search PATTERN+DISCOVERY+AND+VISUALIZATION | Search PATTERN+DISCOVERY+AND+VISUALIZATION |
| 9 | DIGITAL GENE EXPRESSION DISPLAYER | 1 | 100% | 0% | 2 | Search DIGITAL+GENE+EXPRESSION+DISPLAYER | Search DIGITAL+GENE+EXPRESSION+DISPLAYER |
| 10 | HBIOT2 | 1 | 100% | 0% | 2 | Search HBIOT2 | Search HBIOT2 |
Key Words Plus |
| Rank | Web of Science journal category | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
|---|---|---|---|---|---|
| 1 | SERIAL ANALYSIS | 66 | 25% | 42% | 230 |
| 2 | TRANSCRIPT PROFILES | 10 | 57% | 2% | 12 |
| 3 | SAGE | 8 | 13% | 11% | 60 |
| 4 | GENE EXPRESSION SAGE | 8 | 56% | 2% | 10 |
| 5 | REVEAL CHANGES | 7 | 56% | 2% | 9 |
| 6 | LONGSAGE | 6 | 58% | 1% | 7 |
| 7 | SAGE LIBRARIES | 6 | 58% | 1% | 7 |
| 8 | TRANSCRIPT PROFILING ANALYSIS | 5 | 55% | 1% | 6 |
| 9 | TRANSCRIPTOME CHARACTERIZATION | 4 | 50% | 1% | 6 |
| 10 | MPSS | 3 | 35% | 1% | 8 |
Journals |
Reviews |
| Title | Publ. year | Cit. | Active references | % act. ref. to same field |
|---|---|---|---|---|
| Understanding SAGE data | 2007 | 47 | 47 | 72% |
| Serial Analysis of Gene Expression (SAGE): 13 Years of Application in Research | 2008 | 29 | 128 | 66% |
| Statistical evaluation of SAGE libraries: consequences for experimental design | 2002 | 71 | 16 | 81% |
| SuperSAGE: A Modern Platform for Genome-Wide Quantitative Transcript Profiling | 2008 | 22 | 20 | 45% |
| In silico gene expression analysis - an overview | 2007 | 23 | 34 | 38% |
| Tag-based approaches for transcriptome research and genome annotation | 2005 | 89 | 51 | 29% |
| Clustering-based approaches to SAGE data mining | 2008 | 4 | 41 | 73% |
| PET-Tool: a software suite for comprehensive processing and managing of Paired-End diTag (PET) sequence data | 2006 | 8 | 15 | 67% |
| Global Expression Profiling in Epileptogenesis: Does It Add to the Confusion? | 2010 | 15 | 94 | 20% |
| Serial analysis of gene expression - Technical considerations and applications to cardiovascular biology | 2002 | 30 | 39 | 67% |
Address terms |
| Rank | Address term | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
|---|---|---|---|---|---|
| 1 | ENH | 2 | 21% | 1.8% | 10 |
| 2 | MOL ENDOCRINOL ONCOL ANAT PHYSIO | 2 | 67% | 0.4% | 2 |
| 3 | EQUIPE SIGNALISAT IDENT CELLULAI | 1 | 100% | 0.4% | 2 |
| 4 | FUNCT GENOM ANAT PHYSIOL | 1 | 100% | 0.4% | 2 |
| 5 | NEUROZINTUIGEN | 1 | 20% | 0.9% | 5 |
| 6 | BIOINFO USP | 1 | 40% | 0.4% | 2 |
| 7 | NETWORK SERV SOLUT BUSINESS GRP | 1 | 40% | 0.4% | 2 |
| 8 | GRAIN FOODS CRC | 1 | 19% | 0.7% | 4 |
| 9 | IMAGE CONSORTIUM | 1 | 33% | 0.4% | 2 |
| 10 | MICROBIOL MOL VIROL | 1 | 17% | 0.7% | 4 |
Related classes at same level (level 1) |
| Rank | Relatedness score | Related classes |
|---|---|---|
| 1 | 0.0000225587 | MKIAA//SMART CDNA LIBRARY//CDNA SEQUENCING |
| 2 | 0.0000116042 | DIFFERENTIAL DISPLAY//MRNA DIFFERENTIAL DISPLAY//ANIM NUTR FEED YUNNAN PROV |
| 3 | 0.0000112077 | TAISHO FUNCT GENOM//MULTIPHOTON DETECTION//ADAPTER TAGGED COMPETITIVE PCR |
| 4 | 0.0000111719 | WRAP53//OVERLAPPING GENES//CUSTOM BIOTECHNOL SERV GRP |
| 5 | 0.0000107270 | RNA SEQ//DE NOVO ASSEMBLY//TRANSCRIPTOME ASSEMBLY |
| 6 | 0.0000090921 | MICROARRAY IMAGE PROCESSING//GENE FILTERING//MAQC |
| 7 | 0.0000070857 | BIDIRECTIONAL PROMOTER//NEIGHBORING GENES//GENOME NETWORK PROJECT CORE GRPGENOM SCI |
| 8 | 0.0000068284 | RNA AMPLIFICATION//LASER CAPTURE MICRODISSECTION//LASER MICRODISSECTION |
| 9 | 0.0000068113 | VOXELATION//GPR88//GENE EXPRESSION TOMOGRAPHY |
| 10 | 0.0000067615 | PLANT MICROBIAL GENOM//LRT1//NONADDITIVE PROTEIN |