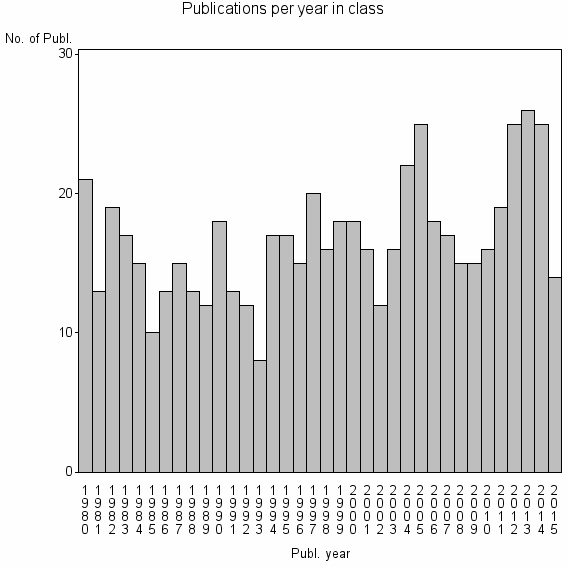
Class information for: |
Basic class information |
| ID | Publications | Average number of references |
Avg. shr. active ref. in WoS |
|---|---|---|---|
| 16195 | 601 | 38.9 | 61% |
Classes in level above (level 2) |
| ID, lev. above |
Publications | Label for level above |
|---|---|---|
| 300 | 17951 | BIODEGRADATION//REDUCTIVE DECHLORINATION//DEHALOCOCCOIDES |
Terms with highest relevance score |
| Rank | Term | Type of term | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
|---|---|---|---|---|---|---|
| 1 | CATECHOL DIOXYGENASE | Author keyword | 12 | 52% | 3% | 16 |
| 2 | GENTISATE 1 2 DIOXYGENASE | Author keyword | 11 | 78% | 1% | 7 |
| 3 | CATECHOL DIOXYGENASES | Author keyword | 9 | 67% | 1% | 8 |
| 4 | MONONUCLEAR IRONIII COMPLEXES | Author keyword | 8 | 100% | 1% | 5 |
| 5 | EXTRADIOL DIOXYGENASE | Author keyword | 6 | 37% | 2% | 13 |
| 6 | EXTRADIOL CLEAVAGE | Author keyword | 4 | 67% | 1% | 4 |
| 7 | INTRADIOL CLEAVAGE | Author keyword | 3 | 57% | 1% | 4 |
| 8 | KLEBSIELLA PNEUMONIAE M5A1 | Author keyword | 3 | 100% | 0% | 3 |
| 9 | EXTRADIOL | Author keyword | 3 | 45% | 1% | 5 |
| 10 | EXTRADIOL TYPE DIOXYGENASE | Author keyword | 3 | 60% | 0% | 3 |
Web of Science journal categories |
Author Key Words |
Key Words Plus |
| Rank | Web of Science journal category | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
|---|---|---|---|---|---|
| 1 | BREVIBACTERIUM FUSCUM | 86 | 96% | 4% | 26 |
| 2 | PROTOCATECHUATE 3 4 DIOXYGENASE | 82 | 51% | 19% | 116 |
| 3 | INTRADIOL | 61 | 86% | 5% | 31 |
| 4 | CATECHOLATOIRONIII COMPLEXES | 47 | 86% | 4% | 24 |
| 5 | EXTRADIOL OXYGENATIONS | 38 | 93% | 2% | 14 |
| 6 | FEII ACTIVE SITE | 35 | 89% | 3% | 16 |
| 7 | HOMOPROTOCATECHUATE 2 3 DIOXYGENASE | 34 | 79% | 4% | 22 |
| 8 | 3N LIGANDS | 27 | 92% | 2% | 11 |
| 9 | EXTRADIOL CLEAVAGE | 26 | 87% | 2% | 13 |
| 10 | 1 2 DIOXYGENASE | 19 | 33% | 8% | 47 |
Journals |
Reviews |
| Title | Publ. year | Cit. | Active references | % act. ref. to same field |
|---|---|---|---|---|
| lron(III) Complexes of Tripodal Monophenolate Ligands as Models for Non-Heme Catechol Dioxygenase Enzymes: Correlation of Dioxygenase Activity with Ligand Stereoelectronic Properties | 2009 | 23 | 89 | 87% |
| Ring-Cleaving Dioxygenases with a Cupin Fold | 2012 | 24 | 120 | 32% |
| The ins and outs of ring-cleaving dioxygenases | 2006 | 148 | 161 | 32% |
| Non-heme iron-dependent dioxygenases: unravelling catalytic mechanisms for complex enzymatic oxidations | 2008 | 88 | 33 | 52% |
| (Catecholato)iron(III) complexes: structural and functional models for the catechol-bound iron(III) form of catechol dioxygenases | 2002 | 55 | 35 | 97% |
| MECHANISTIC ASPECTS OF DIHYDROXYBENZOATE DIOXYGENASES | 1992 | 140 | 59 | 75% |
| Dioxygenase enzymes: catalytic mechanisms and chemical models | 2003 | 156 | 134 | 30% |
| Manganese(II)-dependent extradiol-cleaving catechol dioxygenases | 2000 | 39 | 42 | 86% |
| Enzymatic cleavage of aromatic rings: mechanistic aspects of the catechol dioxygenases and later enzymes of bacterial oxidative cleavage pathways | 1998 | 87 | 120 | 49% |
| Dioxygen activation by enzymes with mononuclear non-heme iron active sites | 1996 | 568 | 145 | 25% |
Address terms |
| Rank | Address term | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
|---|---|---|---|---|---|
| 1 | CAS AGR ENVIRONM MICROBIOL | 1 | 50% | 0.2% | 1 |
| 2 | CRYSTAL STRUCT CHEM GRP | 1 | 50% | 0.2% | 1 |
| 3 | FUGAL PATHOGENESIS | 1 | 50% | 0.2% | 1 |
| 4 | PHARMAZEUT CHEM RUMENTELLE ANALYT CHEM | 1 | 50% | 0.2% | 1 |
| 5 | PROGRAMME ENVIRONM MICROBIOL | 0 | 20% | 0.3% | 2 |
| 6 | CLEANING PROD | 0 | 33% | 0.2% | 1 |
| 7 | HGEBEIT WASSERREINHALTUNG | 0 | 33% | 0.2% | 1 |
| 8 | OORGANISK KEMI | 0 | 33% | 0.2% | 1 |
| 9 | CHEM ANAL TRAINING | 0 | 15% | 0.3% | 2 |
| 10 | DUT KTH | 0 | 25% | 0.2% | 1 |
Related classes at same level (level 1) |
| Rank | Relatedness score | Related classes |
|---|---|---|
| 1 | 0.0000229316 | MET BIOCATALYSIS//BIOMIMET SYST//NONHEME IRON ENZYMES |
| 2 | 0.0000184795 | BIPHENYL DIOXYGENASE//BPH GENES//BIPHENYL DEGRADATION |
| 3 | 0.0000150516 | QUERCETINASE//BAI LIGANDS//QUERCETIN 2 3 DIOXYGENASE |
| 4 | 0.0000147441 | MUCONATE CYCLOISOMERASE//MODIFIED ORTHO CLEAVAGE PATHWAY//CATECHOL 1 2 DIOXYGENASE |
| 5 | 0.0000145702 | XYLS//TOL PLASMID//XYLR |
| 6 | 0.0000135902 | VALENCE TAUTOMERISM//O IMINOBENZOSEMIQUINONE//GA RAZUVAEV ORGANOMETALL CHEM |
| 7 | 0.0000133482 | P HYDROXYBENZOATE HYDROXYLASE//3 HYDROXYBENZOATE 6 HYDROXYLASE//4 AMINOBENZOATE HYDROXYLASE |
| 8 | 0.0000132956 | MODIFIED META CLEAVAGE PATHWAY//2 AMINO 5 CARBOXYMUCONIC 6 SEMIALDEHYDE DEAMINASE//3 NITROBENZOATE |
| 9 | 0.0000114556 | PURPLE ACID PHOSPHATASES//METHANE MONOOXYGENASE//PURPLE ACID PHOSPHATASE |
| 10 | 0.0000078196 | CATECHOLASE ACTIVITY//CATECHOL OXIDASE//MET BIOCATALYSIS |