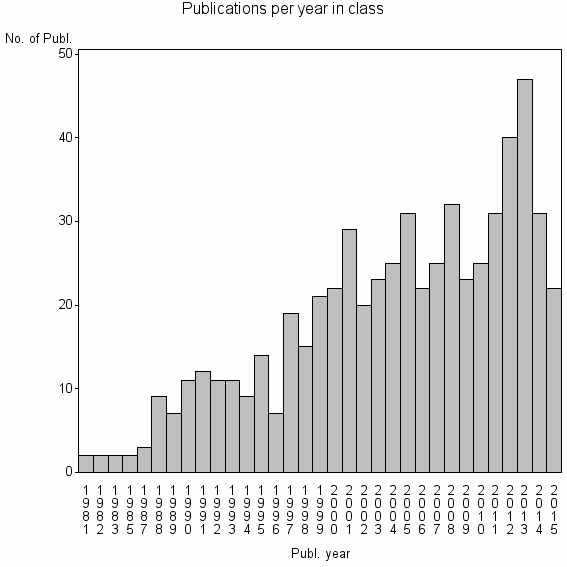
Class information for: |
Basic class information |
| ID | Publications | Average number of references |
Avg. shr. active ref. in WoS |
|---|---|---|---|
| 16120 | 605 | 44.0 | 86% |
Classes in level above (level 2) |
Terms with highest relevance score |
| Rank | Term | Type of term | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
|---|---|---|---|---|---|---|
| 1 | MITHRAMYCIN | Author keyword | 22 | 46% | 6% | 35 |
| 2 | AUREOLIC ACID | Author keyword | 10 | 73% | 1% | 8 |
| 3 | TOLFENAMIC ACID | Author keyword | 9 | 32% | 4% | 23 |
| 4 | SPECIFICITY PROTEIN 1 | Author keyword | 8 | 35% | 3% | 18 |
| 5 | MITHRAMYCIN A | Author keyword | 5 | 42% | 2% | 10 |
| 6 | SP3 | Author keyword | 5 | 16% | 5% | 29 |
| 7 | SP TRANSCRIPTION FACTORS | Author keyword | 4 | 39% | 1% | 9 |
| 8 | LCRG1 | Author keyword | 3 | 100% | 0% | 3 |
| 9 | CHROMOMYCIN A3 | Author keyword | 3 | 20% | 2% | 13 |
| 10 | SP PROTEINS | Author keyword | 3 | 50% | 1% | 4 |
Web of Science journal categories |
Author Key Words |
| Rank | Web of Science journal category | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
LCSH search | Wikipedia search |
|---|---|---|---|---|---|---|---|
| 1 | MITHRAMYCIN | 22 | 46% | 6% | 35 | Search MITHRAMYCIN | Search MITHRAMYCIN |
| 2 | AUREOLIC ACID | 10 | 73% | 1% | 8 | Search AUREOLIC+ACID | Search AUREOLIC+ACID |
| 3 | TOLFENAMIC ACID | 9 | 32% | 4% | 23 | Search TOLFENAMIC+ACID | Search TOLFENAMIC+ACID |
| 4 | SPECIFICITY PROTEIN 1 | 8 | 35% | 3% | 18 | Search SPECIFICITY+PROTEIN+1 | Search SPECIFICITY+PROTEIN+1 |
| 5 | MITHRAMYCIN A | 5 | 42% | 2% | 10 | Search MITHRAMYCIN+A | Search MITHRAMYCIN+A |
| 6 | SP3 | 5 | 16% | 5% | 29 | Search SP3 | Search SP3 |
| 7 | SP TRANSCRIPTION FACTORS | 4 | 39% | 1% | 9 | Search SP+TRANSCRIPTION+FACTORS | Search SP+TRANSCRIPTION+FACTORS |
| 8 | LCRG1 | 3 | 100% | 0% | 3 | Search LCRG1 | Search LCRG1 |
| 9 | CHROMOMYCIN A3 | 3 | 20% | 2% | 13 | Search CHROMOMYCIN+A3 | Search CHROMOMYCIN+A3 |
| 10 | SP PROTEINS | 3 | 50% | 1% | 4 | Search SP+PROTEINS | Search SP+PROTEINS |
Key Words Plus |
| Rank | Web of Science journal category | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
|---|---|---|---|---|---|
| 1 | MG2 COMPLEXES | 53 | 95% | 3% | 18 |
| 2 | PRODUCER STREPTOMYCES ARGILLACEUS | 30 | 79% | 3% | 19 |
| 3 | TRANSCRIPTION FACTOR SP1 | 17 | 19% | 13% | 78 |
| 4 | MITHRAMYCIN | 11 | 29% | 6% | 34 |
| 5 | SP FAMILY | 11 | 26% | 6% | 36 |
| 6 | BOX BINDING PROTEINS | 10 | 18% | 8% | 49 |
| 7 | CHROMOMYCIN A3 | 8 | 28% | 4% | 23 |
| 8 | TRANSCRIPTION FACTOR SP3 | 7 | 41% | 2% | 13 |
| 9 | SMALL DNA DUPLEXES | 6 | 80% | 1% | 4 |
| 10 | SPECIFICITY PROTEINS | 6 | 80% | 1% | 4 |
Journals |
Reviews |
| Title | Publ. year | Cit. | Active references | % act. ref. to same field |
|---|---|---|---|---|
| Sp1 and the 'hallmarks of cancer' | 2015 | 3 | 369 | 19% |
| The Sp-family of transcription factors | 1999 | 785 | 78 | 37% |
| Sp1 Phosphorylation and Its Regulation of Gene Transcription | 2009 | 126 | 73 | 30% |
| Transcription factor Sp1, also known as specificity protein 1 as a therapeutic target | 2014 | 4 | 97 | 59% |
| Sp transcription factor family and its role in cancer | 2005 | 270 | 108 | 25% |
| Regulation of the activity of Sp1-related transcription factors | 2002 | 314 | 123 | 37% |
| Sp1: Emerging roles - Beyond constitutive activation of TATA-less housekeeping genes | 2008 | 186 | 267 | 19% |
| Transcriptional regulation by post-transcriptional modification-Role of phosphorylation in Sp1 transcriptional activity | 2012 | 20 | 100 | 35% |
| Spl: Regulation of gene expression by phosphorylation | 2005 | 177 | 84 | 23% |
| Anti-tumor activity of non-steroidal anti-inflammatory drugs: Cyclooxygenase-independent targets | 2014 | 10 | 96 | 17% |
Address terms |
| Rank | Address term | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
|---|---|---|---|---|---|
| 1 | TRANSLAT ANIM | 2 | 67% | 0.3% | 2 |
| 2 | BIOL FUNDAMENTAL CIENCIES SALUT | 2 | 50% | 0.5% | 3 |
| 3 | VET PHYSIOL TOXICOL | 1 | 100% | 0.3% | 2 |
| 4 | AFFILIATED E HOSP | 1 | 33% | 0.5% | 3 |
| 5 | INFECT DIS SIGNAL TRANSDUCT | 1 | 22% | 0.7% | 4 |
| 6 | ORAL DIS RELATED CPDS | 1 | 33% | 0.3% | 2 |
| 7 | ORAL PHARMACOL | 1 | 13% | 0.8% | 5 |
| 8 | BIO OURCE BIOL SYSTGENE ENGN | 1 | 50% | 0.2% | 1 |
| 9 | CHEM TRANSFORMAT ANTIBIOT | 1 | 50% | 0.2% | 1 |
| 10 | CNRSUMR 49 | 1 | 50% | 0.2% | 1 |
Related classes at same level (level 1) |
| Rank | Relatedness score | Related classes |
|---|---|---|
| 1 | 0.0000104937 | KLF6//KLF4//KRUPPEL LIKE FACTOR 4 |
| 2 | 0.0000082240 | ZBP 89//ZNF148//APOPTOSIS EFFECTOR GENES |
| 3 | 0.0000077719 | ZNF703//C8ORF4//8P12 |
| 4 | 0.0000068889 | MGDC//VERRUCARIN A//RADIAT TUMOR PHYSIOL |
| 5 | 0.0000066351 | VANCOSAMINE//ACOSAMINE//AMICETOSE |
| 6 | 0.0000064884 | NF Y//NF YB//CCAAT BOX |
| 7 | 0.0000063940 | YY1//YIN YANG 1//GON4L |
| 8 | 0.0000056495 | REVERSINE//MITOTIC CATASTROPHE//WP 631 |
| 9 | 0.0000054255 | ARTIFICIAL TRANSCRIPTION FACTOR//ARTIFICIAL TRANSCRIPTION FACTORS//RECOGNITION CODE |
| 10 | 0.0000048950 | AN 9//SABARUBICIN//AN 7 |