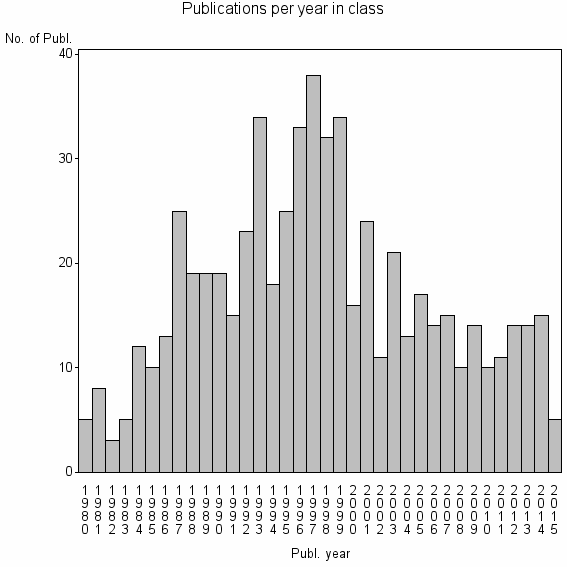
Class information for: |
Basic class information |
| ID | Publications | Average number of references |
Avg. shr. active ref. in WoS |
|---|---|---|---|
| 15952 | 614 | 37.7 | 77% |
Classes in level above (level 2) |
Terms with highest relevance score |
| Rank | Term | Type of term | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
|---|---|---|---|---|---|---|
| 1 | HRCA | Author keyword | 22 | 63% | 4% | 22 |
| 2 | RNA THERMOMETER | Author keyword | 18 | 83% | 2% | 10 |
| 3 | CIRCE | Author keyword | 15 | 62% | 3% | 16 |
| 4 | DNAK OPERON | Author keyword | 13 | 80% | 1% | 8 |
| 5 | SIGMA32 | Author keyword | 6 | 45% | 2% | 10 |
| 6 | BGAB | Author keyword | 4 | 75% | 0% | 3 |
| 7 | SIGMA 32 | Author keyword | 3 | 45% | 1% | 5 |
| 8 | RPOH | Author keyword | 3 | 32% | 1% | 7 |
| 9 | METHYLOVORUS SP STRAIN SS1 | Author keyword | 2 | 67% | 0% | 2 |
| 10 | RNA THERMOMETERS | Author keyword | 2 | 67% | 0% | 2 |
Web of Science journal categories |
Author Key Words |
| Rank | Web of Science journal category | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
LCSH search | Wikipedia search |
|---|---|---|---|---|---|---|---|
| 1 | HRCA | 22 | 63% | 4% | 22 | Search HRCA | Search HRCA |
| 2 | RNA THERMOMETER | 18 | 83% | 2% | 10 | Search RNA+THERMOMETER | Search RNA+THERMOMETER |
| 3 | CIRCE | 15 | 62% | 3% | 16 | Search CIRCE | Search CIRCE |
| 4 | DNAK OPERON | 13 | 80% | 1% | 8 | Search DNAK+OPERON | Search DNAK+OPERON |
| 5 | SIGMA32 | 6 | 45% | 2% | 10 | Search SIGMA32 | Search SIGMA32 |
| 6 | BGAB | 4 | 75% | 0% | 3 | Search BGAB | Search BGAB |
| 7 | SIGMA 32 | 3 | 45% | 1% | 5 | Search SIGMA+32 | Search SIGMA+32 |
| 8 | RPOH | 3 | 32% | 1% | 7 | Search RPOH | Search RPOH |
| 9 | METHYLOVORUS SP STRAIN SS1 | 2 | 67% | 0% | 2 | Search METHYLOVORUS+SP+STRAIN+SS1 | Search METHYLOVORUS+SP+STRAIN+SS1 |
| 10 | RNA THERMOMETERS | 2 | 67% | 0% | 2 | Search RNA+THERMOMETERS | Search RNA+THERMOMETERS |
Key Words Plus |
| Rank | Web of Science journal category | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
|---|---|---|---|---|---|
| 1 | GROESL OPERON | 31 | 57% | 6% | 36 |
| 2 | HTPR GENE | 23 | 79% | 2% | 15 |
| 3 | SIGMA 32 SYNTHESIS | 21 | 64% | 3% | 21 |
| 4 | DNAK OPERON | 17 | 43% | 5% | 31 |
| 5 | TRANSCRIPTION FACTOR SIGMA32 | 16 | 42% | 5% | 29 |
| 6 | THERMOSENSOR CONTROLS EXPRESSION | 15 | 53% | 3% | 20 |
| 7 | GROEL LIKE GENE | 15 | 88% | 1% | 7 |
| 8 | HRCA | 15 | 73% | 2% | 11 |
| 9 | FACTOR SIGMA 32 | 8 | 70% | 1% | 7 |
| 10 | GROEL LIKE GENES | 8 | 70% | 1% | 7 |
Journals |
Reviews |
| Title | Publ. year | Cit. | Active references | % act. ref. to same field |
|---|---|---|---|---|
| Bacterial RNA thermometers: molecular zippers and switches | 2012 | 58 | 83 | 36% |
| Negative regulation of bacterial heat shock genes | 1999 | 153 | 43 | 70% |
| Regulation of the heat-shock response | 1999 | 141 | 61 | 57% |
| Convergence of molecular, modeling, and systems approaches for an understanding of the Escherichia coli heat shock response | 2008 | 96 | 68 | 32% |
| REGULATION OF THE HEAT-SHOCK RESPONSE IN BACTERIA | 1993 | 343 | 106 | 51% |
| Heat-shock and general stress response in Bacillus subtilis | 1996 | 370 | 44 | 25% |
| Cyanobacterial heat-shock response: role and regulation of molecular chaperones | 2014 | 4 | 99 | 28% |
| Translational control of bacterial heat shock and virulence genes by temperature-sensing mRNAs | 2010 | 17 | 44 | 41% |
| Negative regulation of the heat shock response in Streptomyces | 2001 | 44 | 38 | 55% |
| REGULATION OF THE ESCHERICHIA-COLI HEAT-SHOCK RESPONSE | 1993 | 242 | 63 | 46% |
Address terms |
| Rank | Address term | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
|---|---|---|---|---|---|
| 1 | LEHRSTUHL BIOL MIKROORGANISMEN | 2 | 13% | 2.0% | 12 |
| 2 | AGR MICROBIAL TECHNOL MICROBIAL SCI | 1 | 50% | 0.3% | 2 |
| 3 | NYSGXRC | 1 | 50% | 0.2% | 1 |
| 4 | PILOT PLANT NAT PROD | 1 | 50% | 0.2% | 1 |
| 5 | SCI EXPT CLIN VET MED | 1 | 50% | 0.2% | 1 |
| 6 | TARGET BIOTECHNOL | 1 | 50% | 0.2% | 1 |
| 7 | PHARMACEUT SCI MICROBIOL HIGASHI KU | 0 | 33% | 0.2% | 1 |
| 8 | PROGRAMA GENOM EVOLUTIVA | 0 | 33% | 0.2% | 1 |
| 9 | PROMUBIECONICET | 0 | 33% | 0.2% | 1 |
| 10 | MECHNIKOV MICROBIOL IMMUNOL | 0 | 14% | 0.3% | 2 |
Related classes at same level (level 1) |
| Rank | Relatedness score | Related classes |
|---|---|---|
| 1 | 0.0000254474 | DNAJ//GRPE//DNAK |
| 2 | 0.0000182068 | ATP DEPENDENT PROTEASES//CLPB//FTSH |
| 3 | 0.0000140140 | CHAPERONIN//GROEL//GROES |
| 4 | 0.0000116369 | COLD SHOCK PROTEIN//GLYCINE RICH RNA BINDING PROTEIN//CIRP |
| 5 | 0.0000115412 | PHARM YAMASHIRO CHO//TRYPSIN LIKE PROTEINASE//INTRACELLULAR SERINE PROTEASE |
| 6 | 0.0000112860 | HTRA1//HTRA//HTRA2 |
| 7 | 0.0000096899 | RNA POLYMERASE//ESCHERICHIA COLI RNA POLYMERASE//SIGMA70 |
| 8 | 0.0000085887 | CAULOBACTER//CTRA//CAULOBACTER CRESCENTUS |
| 9 | 0.0000076808 | TSAF8//ATP AGAROSE//HSP28 |
| 10 | 0.0000075216 | ABRB//SPO0A//TRANSITION STATE REGULATOR |