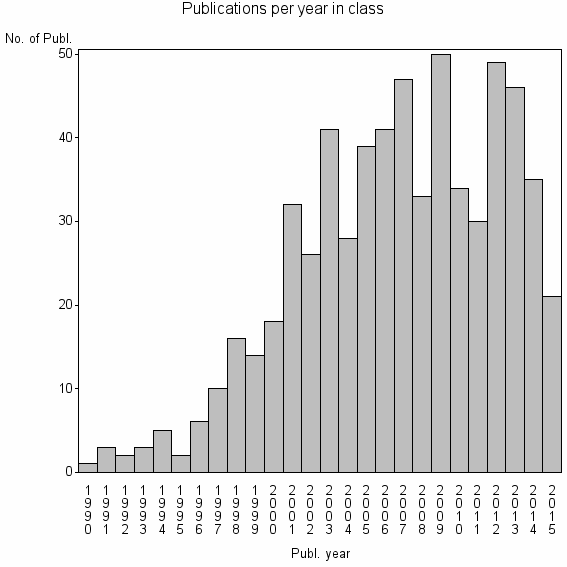
Class information for: |
Basic class information |
| ID | Publications | Average number of references |
Avg. shr. active ref. in WoS |
|---|---|---|---|
| 15617 | 632 | 46.8 | 90% |
Classes in level above (level 2) |
| ID, lev. above |
Publications | Label for level above |
|---|---|---|
| 1315 | 8032 | PROTEIN TRANSLOCATION//PROTEIN IMPORT//SECA |
Terms with highest relevance score |
| Rank | Term | Type of term | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
|---|---|---|---|---|---|---|
| 1 | TWIN ARGININE TRANSLOCATION | Author keyword | 105 | 89% | 8% | 48 |
| 2 | TWIN ARGININE SIGNAL PEPTIDE | Author keyword | 62 | 92% | 4% | 24 |
| 3 | TAT PATHWAY | Author keyword | 47 | 79% | 5% | 30 |
| 4 | TWIN ARGININE | Author keyword | 40 | 76% | 4% | 28 |
| 5 | DMSD | Author keyword | 24 | 91% | 2% | 10 |
| 6 | TWIN ARGININE TRANSLOCASE | Author keyword | 17 | 75% | 2% | 12 |
| 7 | TAT SYSTEM | Author keyword | 13 | 61% | 2% | 14 |
| 8 | SEC INDEPENDENT | Author keyword | 12 | 86% | 1% | 6 |
| 9 | TAT PROTEIN TRANSPORT | Author keyword | 11 | 100% | 1% | 6 |
| 10 | TORD | Author keyword | 11 | 100% | 1% | 6 |
Web of Science journal categories |
Author Key Words |
| Rank | Web of Science journal category | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
LCSH search | Wikipedia search |
|---|---|---|---|---|---|---|---|
| 1 | TWIN ARGININE TRANSLOCATION | 105 | 89% | 8% | 48 | Search TWIN+ARGININE+TRANSLOCATION | Search TWIN+ARGININE+TRANSLOCATION |
| 2 | TWIN ARGININE SIGNAL PEPTIDE | 62 | 92% | 4% | 24 | Search TWIN+ARGININE+SIGNAL+PEPTIDE | Search TWIN+ARGININE+SIGNAL+PEPTIDE |
| 3 | TAT PATHWAY | 47 | 79% | 5% | 30 | Search TAT+PATHWAY | Search TAT+PATHWAY |
| 4 | TWIN ARGININE | 40 | 76% | 4% | 28 | Search TWIN+ARGININE | Search TWIN+ARGININE |
| 5 | DMSD | 24 | 91% | 2% | 10 | Search DMSD | Search DMSD |
| 6 | TWIN ARGININE TRANSLOCASE | 17 | 75% | 2% | 12 | Search TWIN+ARGININE+TRANSLOCASE | Search TWIN+ARGININE+TRANSLOCASE |
| 7 | TAT SYSTEM | 13 | 61% | 2% | 14 | Search TAT+SYSTEM | Search TAT+SYSTEM |
| 8 | SEC INDEPENDENT | 12 | 86% | 1% | 6 | Search SEC+INDEPENDENT | Search SEC+INDEPENDENT |
| 9 | TAT PROTEIN TRANSPORT | 11 | 100% | 1% | 6 | Search TAT+PROTEIN+TRANSPORT | Search TAT+PROTEIN+TRANSPORT |
| 10 | TORD | 11 | 100% | 1% | 6 | Search TORD | Search TORD |
Key Words Plus |
| Rank | Web of Science journal category | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
|---|---|---|---|---|---|
| 1 | SEC INDEPENDENT PROTEIN | 190 | 86% | 15% | 97 |
| 2 | EXPORT PATHWAY | 169 | 65% | 26% | 162 |
| 3 | TWIN ARGININE TRANSLOCASE | 94 | 76% | 10% | 65 |
| 4 | LEADER BINDING PROTEIN | 81 | 96% | 4% | 25 |
| 5 | TAT PATHWAY | 58 | 68% | 8% | 52 |
| 6 | PROTEIN TRANSPORT SYSTEM | 53 | 89% | 4% | 24 |
| 7 | DELTA PH PATHWAY | 50 | 88% | 4% | 23 |
| 8 | SHEWANELLA MASSILIA | 48 | 100% | 3% | 17 |
| 9 | FOLDING QUALITY CONTROL | 39 | 80% | 4% | 24 |
| 10 | SIGNAL PEPTIDE | 38 | 21% | 26% | 165 |
Journals |
Reviews |
| Title | Publ. year | Cit. | Active references | % act. ref. to same field |
|---|---|---|---|---|
| The twin-arginine translocation (Tat) protein export pathway | 2012 | 114 | 140 | 79% |
| Transport and proofreading of proteins by the twin-arginine translocation (Tat) system in bacteria | 2011 | 42 | 78 | 94% |
| The Tat system of Gram-positive bacteria | 2014 | 9 | 149 | 78% |
| The Twin-Arginine Protein Franslocation Pathway | 2015 | 1 | 102 | 88% |
| Protein transport by the bacterial Tat pathway | 2014 | 6 | 97 | 94% |
| The bacterial twin-arginine translocation pathway | 2006 | 156 | 132 | 78% |
| Twin-arginine-dependent translocation of folded proteins | 2012 | 33 | 156 | 97% |
| Possible function of VIPP1 in maintaining chloroplast membranes | 2015 | 1 | 53 | 45% |
| Export of complex cofactor-containing proteins by the bacterial Tat pathway | 2005 | 131 | 47 | 94% |
| The importance of the twin-arginine translocation pathway for bacterial virulence | 2008 | 55 | 48 | 65% |
Address terms |
| Rank | Address term | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
|---|---|---|---|---|---|
| 1 | METALLOPROT SPECT BIOL | 4 | 19% | 3.3% | 21 |
| 2 | BIO GEOWISSEN 1 | 4 | 75% | 0.5% | 3 |
| 3 | HORT SCI PLANT MOL CELLULAR BIOL PROGRAM | 3 | 60% | 0.5% | 3 |
| 4 | UPR9043 | 2 | 16% | 2.2% | 14 |
| 5 | UPR9043 CNRS | 1 | 50% | 0.3% | 2 |
| 6 | BPSI | 1 | 33% | 0.3% | 2 |
| 7 | MET PROT SPECT BIOL | 1 | 14% | 0.8% | 5 |
| 8 | AG SYNTHET MICROBIOL | 1 | 50% | 0.2% | 1 |
| 9 | ARBEITSGRP MOL ZELLBIOL | 1 | 50% | 0.2% | 1 |
| 10 | AVRON EVEN ARI MINERVA PHOTOSYNTH | 1 | 50% | 0.2% | 1 |
Related classes at same level (level 1) |
| Rank | Relatedness score | Related classes |
|---|---|---|
| 1 | 0.0000241965 | CHLOROPLAST PROTEIN IMPORT//TOC75//PROTEIN IMPORT |
| 2 | 0.0000153525 | SECA//SECB//SECY |
| 3 | 0.0000107379 | STREPTOMYCES SUBTILISIN INHIBITOR//DNASE INHIBITOR//M ECO RI |
| 4 | 0.0000102140 | YIDC//OXA1//COX17 |
| 5 | 0.0000085613 | MOLYBDOPTERIN//DIMETHYL SULFOXIDE REDUCTASE//MOLYBDENUM COFACTOR |
| 6 | 0.0000067136 | HTRA1//HTRA//HTRA2 |
| 7 | 0.0000061619 | TYPE II SECRETION//TYPE IV PILI//GENERAL SECRETORY PATHWAY |
| 8 | 0.0000044622 | NIFE HYDROGENASE//HYDROGENASE//HYDROGENASES |
| 9 | 0.0000042776 | ABRB//SPO0A//TRANSITION STATE REGULATOR |
| 10 | 0.0000040009 | SIGNAL RECOGNITION PARTICLE//SRP RECEPTOR//FTSY |