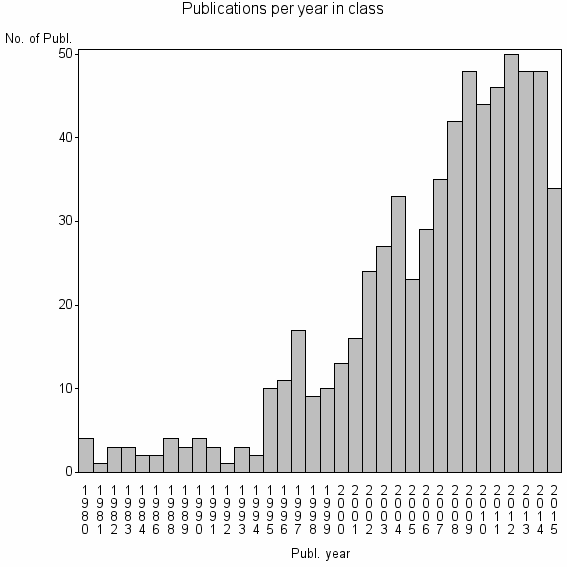
Class information for: |
Basic class information |
| ID | Publications | Average number of references |
Avg. shr. active ref. in WoS |
|---|---|---|---|
| 15271 | 652 | 43.3 | 90% |
Classes in level above (level 2) |
| ID, lev. above |
Publications | Label for level above |
|---|---|---|
| 1527 | 6895 | AUTOTRANSPORTER//PORIN//BACTERIAL CHEMOTAXIS |
Terms with highest relevance score |
| Rank | Term | Type of term | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
|---|---|---|---|---|---|---|
| 1 | HTRA1 | Author keyword | 28 | 48% | 7% | 43 |
| 2 | HTRA | Author keyword | 24 | 53% | 5% | 32 |
| 3 | HTRA2 | Author keyword | 21 | 53% | 4% | 28 |
| 4 | UCF 101 | Author keyword | 18 | 89% | 1% | 8 |
| 5 | OMI HTRA2 | Author keyword | 15 | 56% | 3% | 18 |
| 6 | HTRA3 | Author keyword | 15 | 73% | 2% | 11 |
| 7 | HTRA2 OMI | Author keyword | 11 | 56% | 2% | 14 |
| 8 | RSEB | Author keyword | 11 | 100% | 1% | 6 |
| 9 | EXTRACYTOPLASMIC STRESS | Author keyword | 11 | 78% | 1% | 7 |
| 10 | RSEA | Author keyword | 11 | 78% | 1% | 7 |
Web of Science journal categories |
Author Key Words |
| Rank | Web of Science journal category | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
LCSH search | Wikipedia search |
|---|---|---|---|---|---|---|---|
| 1 | HTRA1 | 28 | 48% | 7% | 43 | Search HTRA1 | Search HTRA1 |
| 2 | HTRA | 24 | 53% | 5% | 32 | Search HTRA | Search HTRA |
| 3 | HTRA2 | 21 | 53% | 4% | 28 | Search HTRA2 | Search HTRA2 |
| 4 | UCF 101 | 18 | 89% | 1% | 8 | Search UCF+101 | Search UCF+101 |
| 5 | OMI HTRA2 | 15 | 56% | 3% | 18 | Search OMI+HTRA2 | Search OMI+HTRA2 |
| 6 | HTRA3 | 15 | 73% | 2% | 11 | Search HTRA3 | Search HTRA3 |
| 7 | HTRA2 OMI | 11 | 56% | 2% | 14 | Search HTRA2+OMI | Search HTRA2+OMI |
| 8 | RSEB | 11 | 100% | 1% | 6 | Search RSEB | Search RSEB |
| 9 | EXTRACYTOPLASMIC STRESS | 11 | 78% | 1% | 7 | Search EXTRACYTOPLASMIC+STRESS | Search EXTRACYTOPLASMIC+STRESS |
| 10 | RSEA | 11 | 78% | 1% | 7 | Search RSEA | Search RSEA |
Key Words Plus |
| Rank | Web of Science journal category | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
|---|---|---|---|---|---|
| 1 | EXTRACYTOPLASMIC STRESS | 43 | 61% | 7% | 46 |
| 2 | SERINE PROTEASE HTRA1 | 37 | 75% | 4% | 27 |
| 3 | OMI HTRA2 | 28 | 57% | 5% | 34 |
| 4 | RPOE GENE | 27 | 76% | 3% | 19 |
| 5 | DEGP | 26 | 56% | 5% | 31 |
| 6 | ENVELOPE STRESS RESPONSE | 24 | 33% | 10% | 62 |
| 7 | RSEA | 24 | 59% | 4% | 27 |
| 8 | HIGH TEMPERATURE REQUIREMENT | 21 | 78% | 2% | 14 |
| 9 | HTRA GENE | 20 | 59% | 3% | 22 |
| 10 | YAEL | 19 | 71% | 2% | 15 |
Journals |
Reviews |
| Title | Publ. year | Cit. | Active references |
% act. ref. to same field |
|---|---|---|---|---|
| HTRA proteases: regulated proteolysis in protein quality control | 2011 | 92 | 97 | 58% |
| Just scratching the surface: an expanding view of the Cpx envelope stress response | 2012 | 42 | 81 | 53% |
| Sensing external stress: watchdogs of the Escherichia coli cell envelope | 2005 | 178 | 43 | 86% |
| Architecture and regulation of HtrA-family proteins involved in protein quality control and stress response | 2013 | 15 | 86 | 66% |
| Regulation of the Escherichia coli sigma(E)-dependent envelope stress response | 2004 | 206 | 42 | 62% |
| Regulation by destruction: design of the sigma(E) envelope stress response | 2008 | 75 | 45 | 64% |
| The mitochondrial serine protease HtrA2/Omi: an overview | 2008 | 101 | 94 | 49% |
| The structural basis of mode of activation and functional diversity: A case study with HtrA family of serine proteases | 2011 | 21 | 85 | 71% |
| HtrA proteins as targets in therapy of cancer and other diseases | 2010 | 29 | 93 | 61% |
| Envelope stress responses and Gram-negative bacterial pathogenesis | 2005 | 159 | 87 | 52% |
Address terms |
| Rank | Address term | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
|---|---|---|---|---|---|
| 1 | FB BIOL GEOG | 9 | 64% | 1.4% | 9 |
| 2 | COMPARAT MEDMOL BACTERIOL GRP | 3 | 100% | 0.5% | 3 |
| 3 | CNRS URA1444 | 2 | 67% | 0.3% | 2 |
| 4 | MOL BACTERIOL GRP COMPARAT MED | 2 | 67% | 0.3% | 2 |
| 5 | MOL BACTERIOL GRP | 2 | 43% | 0.5% | 3 |
| 6 | CELL DEATH REGULAT | 2 | 20% | 1.1% | 7 |
| 7 | EXCELLENCE MOL MED | 2 | 20% | 1.1% | 7 |
| 8 | ABT PHYSIOL MIKROORGANISMEN | 1 | 100% | 0.3% | 2 |
| 9 | WUXI MATERN CHILDRENS HLTH HOSP | 1 | 100% | 0.3% | 2 |
| 10 | GENE FUNCT ANIM | 1 | 16% | 1.1% | 7 |
Related classes at same level (level 1) |
| Rank | Relatedness score | Related classes |
|---|---|---|
| 1 | 0.0000172209 | AUTOTRANSPORTER//AUTODISPLAY//BAM COMPLEX |
| 2 | 0.0000112860 | HRCA//RNA THERMOMETER//CIRCE |
| 3 | 0.0000111370 | ATP DEPENDENT PROTEASES//CLPB//FTSH |
| 4 | 0.0000087298 | ENVZ//OMPR//ENVZ OSMOSENSOR |
| 5 | 0.0000081916 | LIVIN//XIAP//SMAC MIMETICS |
| 6 | 0.0000067136 | TWIN ARGININE TRANSLOCATION//TWIN ARGININE SIGNAL PEPTIDE//TAT PATHWAY |
| 7 | 0.0000066423 | DSBA//DSBB//DSBD |
| 8 | 0.0000060759 | HFQ//S RNA//RYHB |
| 9 | 0.0000055250 | LPXC//LPXC INHIBITORS//LPXA |
| 10 | 0.0000050374 | MAGUK//PDZ DOMAIN//PSD 95 SAP90 |