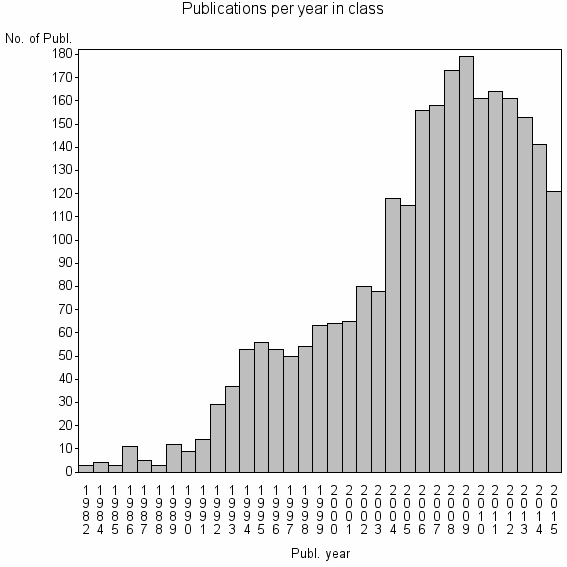
Class information for: |
Basic class information |
| ID | Publications | Average number of references |
Avg. shr. active ref. in WoS |
|---|---|---|---|
| 1481 | 2546 | 50.3 | 80% |
Classes in level above (level 2) |
| ID, lev. above |
Publications | Label for level above |
|---|---|---|
| 825 | 11237 | JOURNAL OF CHEMICAL INFORMATION AND MODELING//B IT//JOURNAL OF CHEMICAL INFORMATION AND COMPUTER SCIENCES |
Terms with highest relevance score |
| Rank | Term | Type of term | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
|---|---|---|---|---|---|---|
| 1 | JOURNAL OF CHEMICAL INFORMATION AND MODELING | Journal | 79 | 16% | 18% | 450 |
| 2 | VIRTUAL SCREENING | Author keyword | 58 | 17% | 12% | 308 |
| 3 | JOURNAL OF COMPUTER-AIDED MOLECULAR DESIGN | Journal | 53 | 17% | 11% | 281 |
| 4 | SCORING FUNCTION | Author keyword | 34 | 38% | 3% | 73 |
| 5 | PROTEIN LIGAND DOCKING | Author keyword | 20 | 44% | 1% | 35 |
| 6 | STRUCTURE BASED DRUG DESIGN | Author keyword | 16 | 16% | 4% | 97 |
| 7 | RECEPTOR FLEXIBILITY | Author keyword | 16 | 58% | 1% | 19 |
| 8 | FLEXIBLE DOCKING | Author keyword | 13 | 30% | 1% | 35 |
| 9 | SCORING FUNCTIONS | Author keyword | 12 | 34% | 1% | 30 |
| 10 | PROTEIN FLEXIBILITY | Author keyword | 11 | 20% | 2% | 48 |
Web of Science journal categories |
Author Key Words |
| Rank | Web of Science journal category | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
LCSH search | Wikipedia search |
|---|---|---|---|---|---|---|---|
| 1 | VIRTUAL SCREENING | 58 | 17% | 12% | 308 | Search VIRTUAL+SCREENING | Search VIRTUAL+SCREENING |
| 2 | SCORING FUNCTION | 34 | 38% | 3% | 73 | Search SCORING+FUNCTION | Search SCORING+FUNCTION |
| 3 | PROTEIN LIGAND DOCKING | 20 | 44% | 1% | 35 | Search PROTEIN+LIGAND+DOCKING | Search PROTEIN+LIGAND+DOCKING |
| 4 | STRUCTURE BASED DRUG DESIGN | 16 | 16% | 4% | 97 | Search STRUCTURE+BASED+DRUG+DESIGN | Search STRUCTURE+BASED+DRUG+DESIGN |
| 5 | RECEPTOR FLEXIBILITY | 16 | 58% | 1% | 19 | Search RECEPTOR+FLEXIBILITY | Search RECEPTOR+FLEXIBILITY |
| 6 | FLEXIBLE DOCKING | 13 | 30% | 1% | 35 | Search FLEXIBLE+DOCKING | Search FLEXIBLE+DOCKING |
| 7 | SCORING FUNCTIONS | 12 | 34% | 1% | 30 | Search SCORING+FUNCTIONS | Search SCORING+FUNCTIONS |
| 8 | PROTEIN FLEXIBILITY | 11 | 20% | 2% | 48 | Search PROTEIN+FLEXIBILITY | Search PROTEIN+FLEXIBILITY |
| 9 | COMPUTER AIDED DRUG DESIGN | 10 | 21% | 2% | 40 | Search COMPUTER+AIDED+DRUG+DESIGN | Search COMPUTER+AIDED+DRUG+DESIGN |
| 10 | BINDING AFFINITY PREDICTION | 9 | 55% | 0% | 12 | Search BINDING+AFFINITY+PREDICTION | Search BINDING+AFFINITY+PREDICTION |
Key Words Plus |
| Rank | Web of Science journal category | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
|---|---|---|---|---|---|
| 1 | FLEXIBLE DOCKING | 341 | 56% | 16% | 410 |
| 2 | MOLECULAR DOCKING | 192 | 34% | 18% | 469 |
| 3 | AUTOMATED DOCKING | 145 | 34% | 14% | 352 |
| 4 | SCORING FUNCTIONS | 119 | 43% | 8% | 213 |
| 5 | PROTEIN LIGAND DOCKING | 117 | 57% | 5% | 138 |
| 6 | SIDE CHAIN FLEXIBILITY | 97 | 66% | 3% | 89 |
| 7 | DIRECTED DRUG DESIGN | 88 | 89% | 2% | 40 |
| 8 | INCREMENTAL CONSTRUCTION ALGORITHM | 86 | 55% | 4% | 107 |
| 9 | HIGH THROUGHPUT DOCKING | 86 | 59% | 4% | 97 |
| 10 | PROTEIN LIGAND INTERACTIONS | 73 | 37% | 6% | 160 |
Journals |
| Rank | Web of Science journal category | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
|---|---|---|---|---|---|
| 1 | JOURNAL OF CHEMICAL INFORMATION AND MODELING | 79 | 16% | 18% | 450 |
| 2 | JOURNAL OF COMPUTER-AIDED MOLECULAR DESIGN | 53 | 17% | 11% | 281 |
| 3 | JOURNAL OF CHEMINFORMATICS | 4 | 12% | 1% | 30 |
| 4 | PERSPECTIVES IN DRUG DISCOVERY AND DESIGN | 2 | 11% | 1% | 16 |
Reviews |
| Title | Publ. year | Cit. | Active references | % act. ref. to same field |
|---|---|---|---|---|
| Beware of docking! | 2015 | 3 | 163 | 50% |
| Can We Trust Docking Results? Evaluation of Seven Commonly Used Programs on PDBbind Database | 2011 | 83 | 51 | 80% |
| Structure-Based Virtual Screening for Drug Discovery: a Problem-Centric Review | 2012 | 65 | 92 | 84% |
| Latest developments in molecular docking: 2010-2011 in review | 2013 | 43 | 207 | 65% |
| Flexible ligand docking to multiple receptor conformations: a practical alternative | 2008 | 191 | 31 | 74% |
| Docking and scoring in virtual screening for drug discovery: Methods and applications | 2004 | 792 | 120 | 62% |
| Protein-ligand docking: Current status and future challenges | 2006 | 346 | 118 | 78% |
| Virtual screening strategies in drug discovery | 2007 | 134 | 37 | 84% |
| Computational Methods in Drug Discovery | 2014 | 29 | 473 | 21% |
| Challenges and advances in computational docking: 2009 in review | 2011 | 64 | 174 | 54% |
Address terms |
| Rank | Address term | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
|---|---|---|---|---|---|
| 1 | COMP AIDED MOL DESIGN GRP | 8 | 44% | 0.5% | 14 |
| 2 | GRP ORPAILLEUR | 4 | 75% | 0.1% | 3 |
| 3 | INHIBOX | 4 | 75% | 0.1% | 3 |
| 4 | PROT STRUCT ANAL DESIGN | 4 | 56% | 0.2% | 5 |
| 5 | PHARMACEUT INNOVAT VALUE CHAIN | 3 | 100% | 0.1% | 3 |
| 6 | S MERAM | 3 | 100% | 0.1% | 3 |
| 7 | NSF THEORET BIOL PHYS | 3 | 24% | 0.5% | 12 |
| 8 | ALGORITHMS SCI COMP | 3 | 23% | 0.4% | 11 |
| 9 | GPIN | 3 | 60% | 0.1% | 3 |
| 10 | GRP COMPUTAT MOL DESIGN | 3 | 60% | 0.1% | 3 |
Related classes at same level (level 1) |
| Rank | Relatedness score | Related classes |
|---|---|---|
| 1 | 0.0000241411 | B IT//LIFE SCI INFORMAT//LIMES PROGRAM UNIT CHEM BIOL MED CHEM |
| 2 | 0.0000128225 | ENVIRONM IND CHEM//ONE STEP PERTURBATION//SAMPL4 |
| 3 | 0.0000118288 | CAPRI//PROTEIN PROTEIN DOCKING//PROTEIN DOCKING |
| 4 | 0.0000112225 | COMPLEX SCI SOFTWARE//3D ZERNIKE DESCRIPTOR//PROTEIN SURFACE SHAPE |
| 5 | 0.0000111168 | NMR SCREENING//FRAGMENT BASED DRUG DISCOVERY//WATERLOGSY |
| 6 | 0.0000106680 | THREE DIMENSIONAL HOLOGRAPHIC VECTOR OF ATOMIC INTERACTION FIELD 3D HOVAIF//COMFA//3D QSAR |
| 7 | 0.0000092212 | POLYPHARMACOLOGY//NETWORK PHARMACOLOGY//DRUG REPOSITIONING |
| 8 | 0.0000085045 | JOURNAL OF CHEMINFORMATICS//INCHI//CHEMICAL MARKUP LANGUAGE |
| 9 | 0.0000079477 | NAKA WORKS//CYCLOALCANES//CONFORMATIONAL POPULATION |
| 10 | 0.0000078809 | PROTEIN HYDRATION//INTERNAL SOLVENT//SOLVENT MODELING |