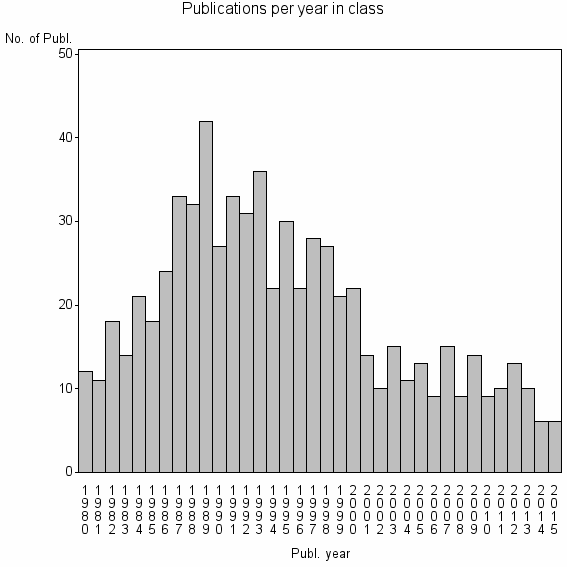
Class information for: |
Basic class information |
| ID | Publications | Average number of references |
Avg. shr. active ref. in WoS |
|---|---|---|---|
| 14676 | 688 | 31.1 | 69% |
Classes in level above (level 2) |
Terms with highest relevance score |
| Rank | Term | Type of term | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
|---|---|---|---|---|---|---|
| 1 | ROLLING CIRCLE REPLICATION | Author keyword | 11 | 29% | 5% | 32 |
| 2 | PLASMID REPLICATION CONTROL | Author keyword | 6 | 71% | 1% | 5 |
| 3 | CRYPTIC PLASMID | Author keyword | 6 | 27% | 3% | 19 |
| 4 | RCR PLASMID | Author keyword | 6 | 100% | 1% | 4 |
| 5 | SINGLE STRAND ORIGIN | Author keyword | 6 | 100% | 1% | 4 |
| 6 | COPG | Author keyword | 4 | 67% | 1% | 4 |
| 7 | ROLLING CIRCLE REPLICATION PLASMID | Author keyword | 4 | 75% | 0% | 3 |
| 8 | MOBILIZATION PROTEIN | Author keyword | 3 | 100% | 0% | 3 |
| 9 | PJB01 | Author keyword | 3 | 100% | 0% | 3 |
| 10 | PT181 | Author keyword | 3 | 100% | 0% | 3 |
Web of Science journal categories |
Author Key Words |
| Rank | Web of Science journal category | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
LCSH search | Wikipedia search |
|---|---|---|---|---|---|---|---|
| 1 | ROLLING CIRCLE REPLICATION | 11 | 29% | 5% | 32 | Search ROLLING+CIRCLE+REPLICATION | Search ROLLING+CIRCLE+REPLICATION |
| 2 | PLASMID REPLICATION CONTROL | 6 | 71% | 1% | 5 | Search PLASMID+REPLICATION+CONTROL | Search PLASMID+REPLICATION+CONTROL |
| 3 | CRYPTIC PLASMID | 6 | 27% | 3% | 19 | Search CRYPTIC+PLASMID | Search CRYPTIC+PLASMID |
| 4 | RCR PLASMID | 6 | 100% | 1% | 4 | Search RCR+PLASMID | Search RCR+PLASMID |
| 5 | SINGLE STRAND ORIGIN | 6 | 100% | 1% | 4 | Search SINGLE+STRAND+ORIGIN | Search SINGLE+STRAND+ORIGIN |
| 6 | COPG | 4 | 67% | 1% | 4 | Search COPG | Search COPG |
| 7 | ROLLING CIRCLE REPLICATION PLASMID | 4 | 75% | 0% | 3 | Search ROLLING+CIRCLE+REPLICATION+PLASMID | Search ROLLING+CIRCLE+REPLICATION+PLASMID |
| 8 | MOBILIZATION PROTEIN | 3 | 100% | 0% | 3 | Search MOBILIZATION+PROTEIN | Search MOBILIZATION+PROTEIN |
| 9 | PJB01 | 3 | 100% | 0% | 3 | Search PJB01 | Search PJB01 |
| 10 | PT181 | 3 | 100% | 0% | 3 | Search PT181 | Search PT181 |
Key Words Plus |
| Rank | Web of Science journal category | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
|---|---|---|---|---|---|
| 1 | PLS1 | 86 | 90% | 5% | 37 |
| 2 | STAPHYLOCOCCAL PLASMIDS | 54 | 80% | 5% | 33 |
| 3 | PUB110 | 30 | 68% | 4% | 27 |
| 4 | PT181 ENCODED REPC PROTEIN | 23 | 100% | 1% | 10 |
| 5 | PC194 | 22 | 68% | 3% | 19 |
| 6 | PT181 REPLICATION | 21 | 90% | 1% | 9 |
| 7 | PT181 | 17 | 62% | 3% | 18 |
| 8 | PT181 FAMILY | 12 | 86% | 1% | 6 |
| 9 | NUMBER CONTROL | 11 | 65% | 2% | 11 |
| 10 | ROLLING CIRCLE REPLICATION | 10 | 20% | 6% | 44 |
Journals |
Reviews |
| Title | Publ. year | Cit. | Active references | % act. ref. to same field |
|---|---|---|---|---|
| Plasmid rolling-circle replication: highlights of two decades of research | 2005 | 105 | 73 | 84% |
| Plasmid rolling-circle replication: recent developments | 2000 | 67 | 29 | 86% |
| Rolling-circle replication of bacterial plasmids | 1997 | 167 | 149 | 71% |
| Replication and control of circular bacterial plasmids | 1998 | 467 | 298 | 29% |
| THE FAMILY OF HIGHLY INTERRELATED SINGLE-STRANDED DEOXYRIBONUCLEIC-ACID PLASMIDS | 1989 | 401 | 72 | 68% |
| ROLLING CIRCLE-REPLICATING PLASMIDS FROM GRAM-POSITIVE AND GRAM-NEGATIVE BACTERIA - A WALL FALLS | 1993 | 119 | 31 | 81% |
| PLASMID ROLLING CIRCLE REPLICATION AND ITS CONTROL | 1995 | 51 | 24 | 92% |
| Small rolling circle plasmids in Bacillus subtilis and related species: Organization, distribution, and their possible role in host physiology | 2007 | 9 | 109 | 69% |
| STAPHYLOCOCCAL PLASMIDS AND THEIR REPLICATION | 1989 | 278 | 76 | 55% |
| The patchwork nature of rolling-circle plasmids: comparison of six plasmids from two distinct Bacillus thuringiensis serotypes | 2003 | 36 | 104 | 45% |
Address terms |
| Rank | Address term | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
|---|---|---|---|---|---|
| 1 | 131 TAKARA MACHI | 2 | 67% | 0.3% | 2 |
| 2 | IND MICROBIOL SECT | 1 | 16% | 0.9% | 6 |
| 3 | PHARMACEUT SCI MICROBIOL | 1 | 16% | 0.7% | 5 |
| 4 | QUIM ANALIT NUTR BROMATOL TECNOL ALIMENTOS | 1 | 50% | 0.1% | 1 |
| 5 | VIROL IMMUNOL CELLULAIRE | 1 | 50% | 0.1% | 1 |
| 6 | MOL MICROBIOL INFECT BIOL | 0 | 14% | 0.4% | 3 |
| 7 | DEF DEV CANADA SUFFIELD | 0 | 33% | 0.1% | 1 |
| 8 | CHRISTIAN DOPPLER GENETICALLY ENGN LACT ACID | 0 | 25% | 0.1% | 1 |
| 9 | INVEST BIOMED CANARIAS | 0 | 25% | 0.1% | 1 |
| 10 | LEK PHARMACEUT | 0 | 25% | 0.1% | 1 |
Related classes at same level (level 1) |
| Rank | Relatedness score | Related classes |
|---|---|---|
| 1 | 0.0000252537 | GENOME VECTOR//NIPPONKO KU//LONG ACCURATE PCR |
| 2 | 0.0000229964 | REPLICATION CONTROL//PLASMID R6K//PLASMID RK2 |
| 3 | 0.0000201115 | FELIX HERELLE REFERENCE BACTERIAL VIRUSES//PHAGE RESISTANCE//LACTOCOCCUS LACTIS |
| 4 | 0.0000198526 | BACTERIOPHAGE ALPHA 3//MICROVIRUS//BACTERIOPHAGE PHI X174 |
| 5 | 0.0000177076 | PHAGE TYPING BACTERIAL GENET//VTB ENZYMTECHNOL//DEGSU |
| 6 | 0.0000146228 | ERM METHYLTRANSFERASE//CMLA//MLS ANTIBIOTIC RESISTANCE |
| 7 | 0.0000129727 | TN916//CONJUGATIVE TRANSPOSON//CONJUGATIVE TRANSPOSONS |
| 8 | 0.0000126733 | THERMUS//THERMACEAE//MEIOTHERMUS |
| 9 | 0.0000111403 | EMANUEL SYNDROME//PARTIAL TRISOMY 11Q//DUPLICATION 11Q |
| 10 | 0.0000110257 | BACTERIAL CONJUGATION//COUPLING PROTEIN//R391 |