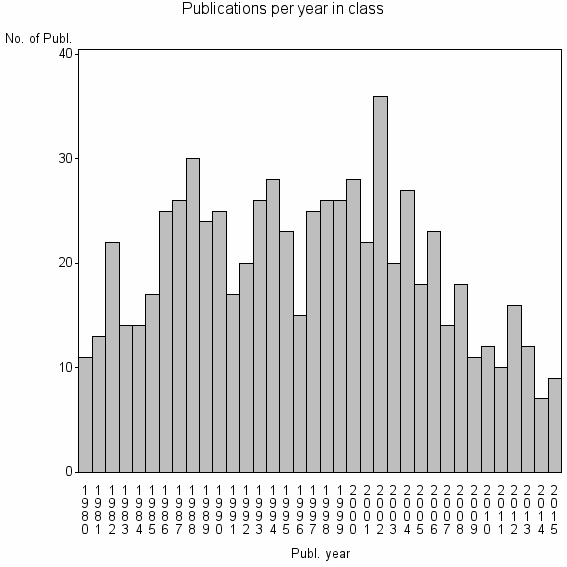
Class information for: |
Basic class information |
| ID | Publications | Average number of references |
Avg. shr. active ref. in WoS |
|---|---|---|---|
| 14329 | 710 | 38.0 | 65% |
Classes in level above (level 2) |
| ID, lev. above |
Publications | Label for level above |
|---|---|---|
| 1002 | 9892 | PHOSPHOLIPASE D//DIACYLGLYCEROL KINASE//PROTEIN KINASE C |
Terms with highest relevance score |
| Rank | Term | Type of term | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
|---|---|---|---|---|---|---|
| 1 | DNA BOUND LIPIDS | Author keyword | 9 | 83% | 1% | 5 |
| 2 | NUCLEAR PHOSPHOLIPIDS | Author keyword | 6 | 80% | 1% | 4 |
| 3 | PIKE A | Author keyword | 6 | 100% | 1% | 4 |
| 4 | NUCL LIPID BIOPATHOL | Address | 5 | 55% | 1% | 6 |
| 5 | CELLULAR SIGNALLING | Address | 4 | 14% | 4% | 30 |
| 6 | BIOMED SCI DIBINEM | Address | 4 | 75% | 0% | 3 |
| 7 | SIGNAL TRANSDUCT UNIT CELL BIOL | Address | 4 | 75% | 0% | 3 |
| 8 | PHYSIOPATHOL SECT | Address | 4 | 56% | 1% | 5 |
| 9 | INOSITOL LIPIDS | Author keyword | 4 | 36% | 1% | 9 |
| 10 | FRIEND CELLS | Author keyword | 4 | 46% | 1% | 6 |
Web of Science journal categories |
Author Key Words |
| Rank | Web of Science journal category | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
LCSH search | Wikipedia search |
|---|---|---|---|---|---|---|---|
| 1 | DNA BOUND LIPIDS | 9 | 83% | 1% | 5 | Search DNA+BOUND+LIPIDS | Search DNA+BOUND+LIPIDS |
| 2 | NUCLEAR PHOSPHOLIPIDS | 6 | 80% | 1% | 4 | Search NUCLEAR+PHOSPHOLIPIDS | Search NUCLEAR+PHOSPHOLIPIDS |
| 3 | PIKE A | 6 | 100% | 1% | 4 | Search PIKE+A | Search PIKE+A |
| 4 | INOSITOL LIPIDS | 4 | 36% | 1% | 9 | Search INOSITOL+LIPIDS | Search INOSITOL+LIPIDS |
| 5 | FRIEND CELLS | 4 | 46% | 1% | 6 | Search FRIEND+CELLS | Search FRIEND+CELLS |
| 6 | ENDONUCLEAR LIPIDS | 3 | 100% | 0% | 3 | Search ENDONUCLEAR+LIPIDS | Search ENDONUCLEAR+LIPIDS |
| 7 | NUCLEAR SIGNAL TRANSDUCTION | 3 | 60% | 0% | 3 | Search NUCLEAR+SIGNAL+TRANSDUCTION | Search NUCLEAR+SIGNAL+TRANSDUCTION |
| 8 | PHOSPHOLIPASE C BETA 1 | 3 | 32% | 1% | 7 | Search PHOSPHOLIPASE+C+BETA+1 | Search PHOSPHOLIPASE+C+BETA+1 |
| 9 | NUCLEAR PI 3 KINASE | 2 | 67% | 0% | 2 | Search NUCLEAR+PI+3+KINASE | Search NUCLEAR+PI+3+KINASE |
| 10 | HEPATOCYTE NUCLEI | 2 | 50% | 0% | 3 | Search HEPATOCYTE+NUCLEI | Search HEPATOCYTE+NUCLEI |
Key Words Plus |
| Rank | Web of Science journal category | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
|---|---|---|---|---|---|
| 1 | FRIEND CELLS | 54 | 71% | 6% | 44 |
| 2 | PI PLC BETA 1 | 27 | 92% | 2% | 11 |
| 3 | RAT LIVER NUCLEI | 15 | 19% | 10% | 72 |
| 4 | OSTEOSARCOMA SAOS 2 CELLS | 15 | 71% | 2% | 12 |
| 5 | G2 M PHASE TRANSITION | 14 | 64% | 2% | 14 |
| 6 | CHROMATIN PHOSPHOLIPIDS | 11 | 78% | 1% | 7 |
| 7 | PHOSPHOINOSITIDASE C BETA | 10 | 73% | 1% | 8 |
| 8 | LIPID PHOSPHORYLATION | 7 | 64% | 1% | 7 |
| 9 | PHOSPHOLIPASE C BETA1 | 7 | 50% | 1% | 10 |
| 10 | PLC BETA1 | 6 | 80% | 1% | 4 |
Journals |
Reviews |
| Title | Publ. year | Cit. | Active references | % act. ref. to same field |
|---|---|---|---|---|
| Nuclear lipid signalling | 2003 | 239 | 127 | 44% |
| INOSITIDES AND THE NUCLEUS AND INOSITIDES IN THE NUCLEUS | 1993 | 196 | 16 | 56% |
| Nuclear sphingolipids: metabolism and signaling | 2008 | 44 | 85 | 31% |
| The role of intranuclear lipids | 2004 | 73 | 75 | 67% |
| The nuclear phosphoinositide 3-kinase/AKT pathway: a new second messenger system | 2002 | 121 | 70 | 37% |
| Nuclear phospholipid signaling: phosphatidylinositol-specific phospholipase C and phosphoinositide 3-kinase | 2007 | 19 | 83 | 66% |
| Nuclear phospholipase C: Involvement in signal transduction | 2005 | 42 | 112 | 57% |
| Nuclear Sphingolipid Metabolism | 2012 | 15 | 152 | 26% |
| Lipid Mediators in the Neural Cell Nucleus: Their Metabolism, Signaling, and Association with Neurological Disorders | 2009 | 24 | 76 | 39% |
| Nuclear phospholipase C signaling through type 1 IGF receptor and its involvement in cell growth and differentiation | 2005 | 15 | 20 | 85% |
Address terms |
| Rank | Address term | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
|---|---|---|---|---|---|
| 1 | NUCL LIPID BIOPATHOL | 5 | 55% | 0.8% | 6 |
| 2 | CELLULAR SIGNALLING | 4 | 14% | 4.2% | 30 |
| 3 | BIOMED SCI DIBINEM | 4 | 75% | 0.4% | 3 |
| 4 | SIGNAL TRANSDUCT UNIT CELL BIOL | 4 | 75% | 0.4% | 3 |
| 5 | PHYSIOPATHOL SECT | 4 | 56% | 0.7% | 5 |
| 6 | CLIN EXPT MED PHYSIOPATHOL | 3 | 100% | 0.4% | 3 |
| 7 | IST CITOMORFOL NORMALE PATOL | 3 | 19% | 2.3% | 16 |
| 8 | HUMAN ANAT SCI | 3 | 21% | 1.8% | 13 |
| 9 | DIPARTIMENTO MORFOL EMBRIOL | 3 | 19% | 2.0% | 14 |
| 10 | DIPARTIMENTO MORFOL UMANA NORMALE | 3 | 22% | 1.5% | 11 |
Related classes at same level (level 1) |
| Rank | Relatedness score | Related classes |
|---|---|---|
| 1 | 0.0000183306 | PHOSPHOLIPASE C DELTA 1//PHOSPHOLIPASE C GAMMA 1//GENOME BIOSIGNAL |
| 2 | 0.0000141942 | DIACYLGLYCEROL KINASE//DIACYLGLYCEROL KINASE DGK//DGK ZETA |
| 3 | 0.0000140711 | NUCLEAR ION CHANNELS//NUCLEOPLASMIC RETICULUM//NUCLEAR CA2 |
| 4 | 0.0000089739 | PIKFYVE//PTDINS5P//GOLPH3 |
| 5 | 0.0000064844 | INTACT NUCLEI//TIP FAK ARASTIRMA HASTANESI NOROSIRURJI KLIN//EXPT CLIN PHYSIOPATHOL GRP BIO296 |
| 6 | 0.0000062149 | PROTEIN KINASE C ISOFORMS//PROTEIN KINASE C ISOZYMES//PROTEIN KINASE C SUBSPECIES |
| 7 | 0.0000061886 | DCP LA//ALPHA 7 ACH RECEPTOR//ALPHA 7 ACETYLCHOLINE RECEPTOR |
| 8 | 0.0000060223 | FATTY ACID REMODELING//TRANSACYLASE//COA INDEPENDENT TRANSACYLASE |
| 9 | 0.0000056887 | NUCLEAR MATRIX//MATRIX ATTACHMENT REGION//NUCLEAR SCAFFOLD |
| 10 | 0.0000056511 | HEPATIC NUCLEAR BINDING//P65 GENE//ANTI P65 ANTIBODY |