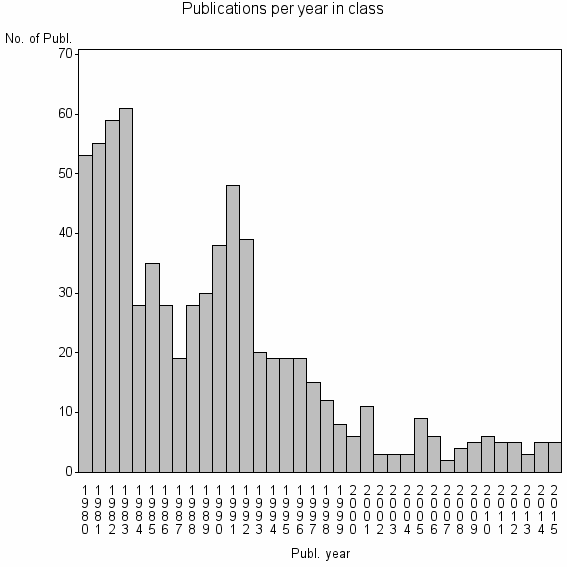
Class information for: |
Basic class information |
| ID | Publications | Average number of references |
Avg. shr. active ref. in WoS |
|---|---|---|---|
| 14278 | 714 | 42.8 | 48% |
Classes in level above (level 2) |
| ID, lev. above |
Publications | Label for level above |
|---|---|---|
| 501 | 14604 | RNA POLYMERASE//SITE SPECIFIC RECOMBINATION//H NS |
Terms with highest relevance score |
| Rank | Term | Type of term | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
|---|---|---|---|---|---|---|
| 1 | TERMINATION OF REPLICATION | Author keyword | 11 | 78% | 1% | 7 |
| 2 | REPLICATION TERMINATOR PROTEIN | Author keyword | 9 | 83% | 1% | 5 |
| 3 | FORK ARREST | Author keyword | 8 | 75% | 1% | 6 |
| 4 | TERMINATOR PROTEIN | Author keyword | 4 | 75% | 0% | 3 |
| 5 | DNA TERMINATOR | Author keyword | 3 | 100% | 0% | 3 |
| 6 | GLUCONATE METABOLISM | Author keyword | 2 | 50% | 0% | 3 |
| 7 | REPLICATION TERMINUS | Author keyword | 2 | 43% | 0% | 3 |
| 8 | TUS | Author keyword | 1 | 31% | 1% | 4 |
| 9 | CONTRAHELICASE ACTIVITY | Author keyword | 1 | 100% | 0% | 2 |
| 10 | FIT MUTATIONS | Author keyword | 1 | 100% | 0% | 2 |
Web of Science journal categories |
Author Key Words |
| Rank | Web of Science journal category | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
LCSH search | Wikipedia search |
|---|---|---|---|---|---|---|---|
| 1 | TERMINATION OF REPLICATION | 11 | 78% | 1% | 7 | Search TERMINATION+OF+REPLICATION | Search TERMINATION+OF+REPLICATION |
| 2 | REPLICATION TERMINATOR PROTEIN | 9 | 83% | 1% | 5 | Search REPLICATION+TERMINATOR+PROTEIN | Search REPLICATION+TERMINATOR+PROTEIN |
| 3 | FORK ARREST | 8 | 75% | 1% | 6 | Search FORK+ARREST | Search FORK+ARREST |
| 4 | TERMINATOR PROTEIN | 4 | 75% | 0% | 3 | Search TERMINATOR+PROTEIN | Search TERMINATOR+PROTEIN |
| 5 | DNA TERMINATOR | 3 | 100% | 0% | 3 | Search DNA+TERMINATOR | Search DNA+TERMINATOR |
| 6 | GLUCONATE METABOLISM | 2 | 50% | 0% | 3 | Search GLUCONATE+METABOLISM | Search GLUCONATE+METABOLISM |
| 7 | REPLICATION TERMINUS | 2 | 43% | 0% | 3 | Search REPLICATION+TERMINUS | Search REPLICATION+TERMINUS |
| 8 | TUS | 1 | 31% | 1% | 4 | Search TUS | Search TUS |
| 9 | CONTRAHELICASE ACTIVITY | 1 | 100% | 0% | 2 | Search CONTRAHELICASE+ACTIVITY | Search CONTRAHELICASE+ACTIVITY |
| 10 | FIT MUTATIONS | 1 | 100% | 0% | 2 | Search FIT+MUTATIONS | Search FIT+MUTATIONS |
Key Words Plus |
| Rank | Web of Science journal category | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
|---|---|---|---|---|---|
| 1 | FORK ARREST | 16 | 64% | 2% | 16 |
| 2 | ANTIGEN HELICASE | 15 | 82% | 1% | 9 |
| 3 | EXHIBIT POLAR INHIBITION | 13 | 71% | 1% | 10 |
| 4 | GLUCONOKINASE | 12 | 86% | 1% | 6 |
| 5 | POLAR CONTRAHELICASE | 12 | 86% | 1% | 6 |
| 6 | CONTRAHELICASE | 11 | 100% | 1% | 6 |
| 7 | TUS TER COMPLEX | 8 | 75% | 1% | 6 |
| 8 | TERMINUS REGION | 7 | 25% | 4% | 26 |
| 9 | TUS PROTEIN | 6 | 80% | 1% | 4 |
| 10 | TERMINATOR PROTEIN | 6 | 41% | 2% | 11 |
Journals |
Reviews |
| Title | Publ. year | Cit. | Active references | % act. ref. to same field |
|---|---|---|---|---|
| Termination of DNA replication of bacterial and plasmid chromosomes | 1999 | 53 | 32 | 88% |
| Replication termination in Escherichia coli: Structure and antihelicase activity of the Tus-Ter complex | 2005 | 53 | 159 | 42% |
| Mechanisms of polar arrest of a replication fork | 2009 | 11 | 33 | 64% |
| ARREST OF BACTERIAL-DNA REPLICATION | 1992 | 66 | 72 | 71% |
| REPLICATION ARREST | 1995 | 28 | 19 | 84% |
| A COLLECTION OF STRAINS CONTAINING GENETICALLY LINKED ALTERNATING ANTIBIOTIC-RESISTANCE ELEMENTS FOR GENETIC-MAPPING OF ESCHERICHIA-COLI | 1989 | 649 | 39 | 21% |
| COLIBRI - A FUNCTIONAL DATA-BASE FOR THE ESCHERICHIA-COLI GENOME | 1993 | 58 | 23 | 57% |
| Mechanism and physiological significance of programmed replication termination | 2014 | 2 | 86 | 44% |
| TUS AND THE TERMINATORS - THE ARREST OF REPLICATION IN PROKARYOTES | 1989 | 51 | 18 | 100% |
| What's for dinner? Entner-Doudoroff metabolism in Escherichia coli | 1998 | 107 | 45 | 27% |
Address terms |
| Rank | Address term | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
|---|---|---|---|---|---|
| 1 | CUD | 1 | 50% | 0.1% | 1 |
| 2 | JOSEPHINE BAY PAUL COMP MOL BIOL EVOLUT | 1 | 50% | 0.1% | 1 |
| 3 | BIO BRAIN | 0 | 25% | 0.1% | 1 |
| 4 | CARDIFF PURE PL BIOL | 0 | 25% | 0.1% | 1 |
| 5 | GENE TECHNOL SECTOR | 0 | 25% | 0.1% | 1 |
| 6 | SCI INFORMAT ANAL MANAGEMENT | 0 | 25% | 0.1% | 1 |
| 7 | MOL MICROBIAL PATHOGENESIS UNIT | 0 | 20% | 0.1% | 1 |
| 8 | NORDEA HEALTHY AGING | 0 | 20% | 0.1% | 1 |
| 9 | CHEM BIOMOL ENGN BIOCENTURY | 0 | 14% | 0.1% | 1 |
| 10 | ENGN BIOCHEM ENGN | 0 | 11% | 0.1% | 1 |
Related classes at same level (level 1) |
| Rank | Relatedness score | Related classes |
|---|---|---|
| 1 | 0.0000118788 | HISTIDINE BIOSYNTHESIS//IMIDAZOLEGLYCEROL PHOSPHATE DEHYDRATASE//HISTIDINOL DEHYDROGENASE |
| 2 | 0.0000105795 | CYCLIC AMP RECEPTOR PROTEIN//CATABOLITE ACTIVATOR PROTEIN CAP//CAMP RECEPTOR PROTEIN |
| 3 | 0.0000103716 | TYRR//SHIKIMIC ACID PRODUCTION//CIS CIS MUCONIC ACID |
| 4 | 0.0000101984 | DNAA PROTEIN//DNAA//ORIC |
| 5 | 0.0000097980 | FNR//NAR PROMOTER//SPIROSIN |
| 6 | 0.0000095867 | KAUFFMANN WHITE SCHEME//I CEUI//HMU UCFM INFECT GENOM |
| 7 | 0.0000093751 | DNA TRANSPOSITION//PHAGE MU//TN10 |
| 8 | 0.0000089073 | HOLLIDAY JUNCTION//RECBCD ENZYME//RECBCD |
| 9 | 0.0000088796 | THREONINE DEHYDROGENASE//SERINE DEHYDRATASE//L THREONINE DEHYDROGENASE |
| 10 | 0.0000086956 | CARAB//PYRIMIDINE REGULATION//PYRR |