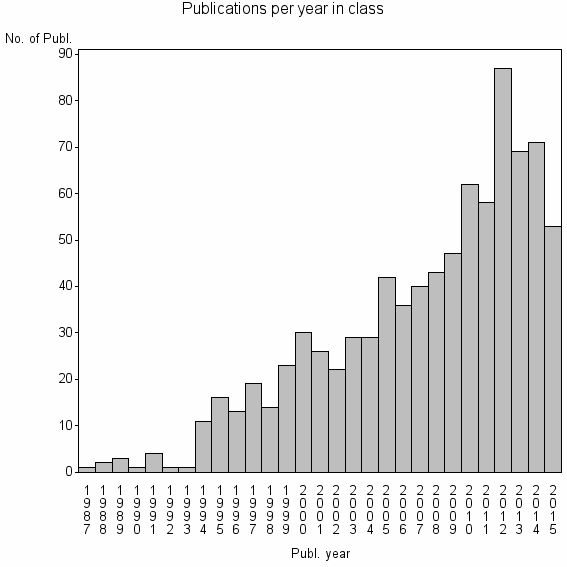
Class information for: |
Basic class information |
| ID | Publications | Average number of references |
Avg. shr. active ref. in WoS |
|---|---|---|---|
| 12283 | 853 | 48.4 | 92% |
Classes in level above (level 2) |
| ID, lev. above |
Publications | Label for level above |
|---|---|---|
| 1825 | 5671 | HUMAN ENDOGENOUS RETROVIRUS//ADAR//ENDOGENOUS RETROVIRUS |
Terms with highest relevance score |
| Rank | Term | Type of term | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
|---|---|---|---|---|---|---|
| 1 | ADAR | Author keyword | 127 | 78% | 10% | 83 |
| 2 | RNA EDITING | Author keyword | 66 | 25% | 27% | 234 |
| 3 | ADAR2 | Author keyword | 41 | 87% | 2% | 20 |
| 4 | ADAR1 | Author keyword | 26 | 55% | 4% | 33 |
| 5 | A TO I RNA EDITING | Author keyword | 21 | 73% | 2% | 16 |
| 6 | ADARS | Author keyword | 21 | 90% | 1% | 9 |
| 7 | ADENOSINE DEAMINATION | Author keyword | 19 | 76% | 2% | 13 |
| 8 | A TO I EDITING | Author keyword | 12 | 75% | 1% | 9 |
| 9 | SEROTONIN 2C RECEPTOR | Author keyword | 12 | 50% | 2% | 18 |
| 10 | INOSINE | Author keyword | 7 | 13% | 6% | 50 |
Web of Science journal categories |
Author Key Words |
| Rank | Web of Science journal category | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
LCSH search | Wikipedia search |
|---|---|---|---|---|---|---|---|
| 1 | ADAR | 127 | 78% | 10% | 83 | Search ADAR | Search ADAR |
| 2 | RNA EDITING | 66 | 25% | 27% | 234 | Search RNA+EDITING | Search RNA+EDITING |
| 3 | ADAR2 | 41 | 87% | 2% | 20 | Search ADAR2 | Search ADAR2 |
| 4 | ADAR1 | 26 | 55% | 4% | 33 | Search ADAR1 | Search ADAR1 |
| 5 | A TO I RNA EDITING | 21 | 73% | 2% | 16 | Search A+TO+I+RNA+EDITING | Search A+TO+I+RNA+EDITING |
| 6 | ADARS | 21 | 90% | 1% | 9 | Search ADARS | Search ADARS |
| 7 | ADENOSINE DEAMINATION | 19 | 76% | 2% | 13 | Search ADENOSINE+DEAMINATION | Search ADENOSINE+DEAMINATION |
| 8 | A TO I EDITING | 12 | 75% | 1% | 9 | Search A+TO+I+EDITING | Search A+TO+I+EDITING |
| 9 | SEROTONIN 2C RECEPTOR | 12 | 50% | 2% | 18 | Search SEROTONIN+2C+RECEPTOR | Search SEROTONIN+2C+RECEPTOR |
| 10 | INOSINE | 7 | 13% | 6% | 50 | Search INOSINE | Search INOSINE |
Key Words Plus |
| Rank | Web of Science journal category | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
|---|---|---|---|---|---|
| 1 | ADENOSINE DEAMINASES | 138 | 76% | 11% | 96 |
| 2 | GLUR B | 118 | 73% | 11% | 91 |
| 3 | HUMAN TRANSCRIPTOME | 90 | 44% | 18% | 155 |
| 4 | ENZYME ADAR2 | 77 | 96% | 3% | 24 |
| 5 | CANDIDATE ENZYME | 73 | 89% | 4% | 33 |
| 6 | ENZYME ADAR1 | 49 | 94% | 2% | 17 |
| 7 | SEROTONIN 2C RECEPTOR | 48 | 67% | 5% | 43 |
| 8 | ADAR DEAMINASES | 47 | 84% | 3% | 26 |
| 9 | DEAMINASES | 42 | 74% | 4% | 31 |
| 10 | ADAR2 | 41 | 81% | 3% | 25 |
Journals |
Reviews |
| Title | Publ. year | Cit. | Active references | % act. ref. to same field |
|---|---|---|---|---|
| Functions and Regulation of RNA Editing by ADAR Deaminases | 2010 | 251 | 149 | 81% |
| RNA editing by adenosine deaminases that act on RNA | 2002 | 557 | 114 | 65% |
| Molecular diversity through RNA editing: a balancing act | 2010 | 65 | 90 | 79% |
| A-to-I RNA Editing: Effects on Proteins Key to Neural Excitability | 2012 | 34 | 54 | 76% |
| Deciphering the functions and regulation of brain-enriched A-to-I RNA editing | 2013 | 17 | 42 | 83% |
| Elucidating the inosinome: global approaches to adenosine-to-inosine RNA editing | 2011 | 35 | 37 | 81% |
| ADAR editing in double-stranded UTRs and other noncoding RNA sequences | 2010 | 48 | 68 | 72% |
| Adenosine deaminases acting on RNA (ADARs) are both antiviral and proviral | 2011 | 51 | 182 | 59% |
| Adenosine-to-inosine RNA editing and human disease | 2013 | 16 | 77 | 70% |
| RNA Editing by Mammalian ADARs | 2011 | 36 | 119 | 70% |
Address terms |
| Rank | Address term | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
|---|---|---|---|---|---|
| 1 | RNA EDITING | 30 | 100% | 1.4% | 12 |
| 2 | INTER PROGRAM BIOCHEM MOL BIOL | 2 | 29% | 0.6% | 5 |
| 3 | COMP SCI COMMUN CSC | 1 | 50% | 0.2% | 2 |
| 4 | OHIO STATE RNA BIOL | 1 | 40% | 0.2% | 2 |
| 5 | PROGRAM HUMAN MED GENET | 1 | 40% | 0.2% | 2 |
| 6 | ONCOHAEMATOL | 1 | 12% | 0.9% | 8 |
| 7 | INTERDEPARTMENTAL PROGRAM BIOCHEM MOL BIOL | 1 | 25% | 0.4% | 3 |
| 8 | PEDIAT ONCOHAEMATOL | 1 | 33% | 0.2% | 2 |
| 9 | BIOL INTRACELLULAR BACTERIA UNIT | 1 | 50% | 0.1% | 1 |
| 10 | BIOMED UNIT ASSOCIATED | 1 | 50% | 0.1% | 1 |
Related classes at same level (level 1) |
| Rank | Relatedness score | Related classes |
|---|---|---|
| 1 | 0.0000185291 | DYSCHROMATOSIS SYMMETRICA HEREDITARIA//DOWLING DEGOS DISEASE//DSRAD GENE |
| 2 | 0.0000145552 | P54NRB//POLYPYRIMIDINE TRACT BINDING PROTEIN ASSOCIATED SPLICING FACTOR//MATRIN 3 |
| 3 | 0.0000125696 | PKR//VA RNA//NF90 |
| 4 | 0.0000109179 | WRAP53//OVERLAPPING GENES//CUSTOM BIOTECHNOL SERV GRP |
| 5 | 0.0000080376 | CYCLOTHIAZIDE//KAINATE RECEPTOR//KAINATE RECEPTORS |
| 6 | 0.0000071684 | 5 HT2C RECEPTOR//5 HT2C RECEPTORS//5 HT2A RECEPTOR |
| 7 | 0.0000062431 | CYTIDINE DEAMINASE//ANTIBIOTIC FUNGICIDE//BLASTICIDIN S DEAMINASE |
| 8 | 0.0000053770 | HEPATITIS DELTA VIRUS//HEPATITIS D VIRUS//HDV RNA |
| 9 | 0.0000052262 | TRNA NUCLEOTIDYLTRANSFERASE//CCA ADDING ENZYME//TRNAHIS GUANYLYLTRANSFERASE |
| 10 | 0.0000052140 | ENDONUCLEASE V//S CAROLINA EXPT STN//OXANINE |