Class information for: |
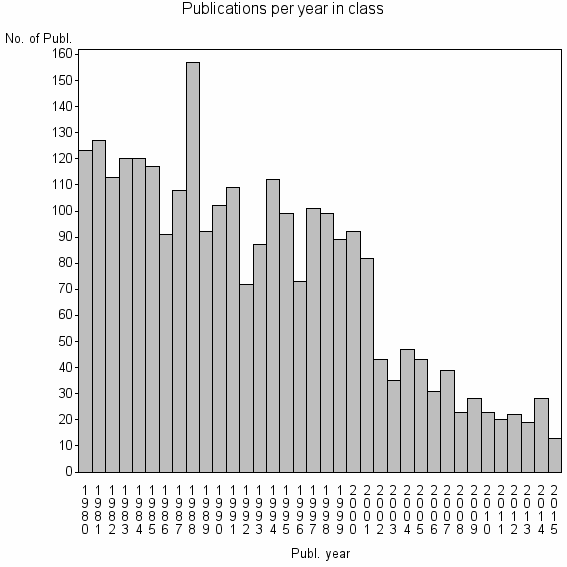
Basic class information |
| ID | Publications | Average number of references |
Avg. shr. active ref. in WoS |
|---|---|---|---|
| 1184 | 2699 | 38.2 | 58% |
Classes in level above (level 2) |
Terms with highest relevance score |
| Rank | Term | Type of term | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
|---|---|---|---|---|---|---|
| 1 | MRNA ANALOG | Author keyword | 53 | 95% | 1% | 18 |
| 2 | 80S RIBOSOME | Author keyword | 30 | 79% | 1% | 19 |
| 3 | HUMAN RIBOSOME | Author keyword | 19 | 76% | 0% | 13 |
| 4 | RIBOSOMAL PROTEIN S15 | Author keyword | 19 | 74% | 1% | 14 |
| 5 | RIBOSOME STRUCTURE | Author keyword | 17 | 50% | 1% | 25 |
| 6 | AG RIBOSOMEN | Address | 17 | 47% | 1% | 27 |
| 7 | RIBOSOMAL PROTEIN S7 | Author keyword | 17 | 75% | 0% | 12 |
| 8 | RIBOSOME | Author keyword | 16 | 10% | 6% | 149 |
| 9 | PEPTIDYL TRANSFERASE | Author keyword | 16 | 48% | 1% | 24 |
| 10 | 30 S SUBUNIT | Author keyword | 14 | 100% | 0% | 7 |
Web of Science journal categories |
Author Key Words |
| Rank | Web of Science journal category | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
LCSH search | Wikipedia search |
|---|---|---|---|---|---|---|---|
| 1 | MRNA ANALOG | 53 | 95% | 1% | 18 | Search MRNA+ANALOG | Search MRNA+ANALOG |
| 2 | 80S RIBOSOME | 30 | 79% | 1% | 19 | Search 80S+RIBOSOME | Search 80S+RIBOSOME |
| 3 | HUMAN RIBOSOME | 19 | 76% | 0% | 13 | Search HUMAN+RIBOSOME | Search HUMAN+RIBOSOME |
| 4 | RIBOSOMAL PROTEIN S15 | 19 | 74% | 1% | 14 | Search RIBOSOMAL+PROTEIN+S15 | Search RIBOSOMAL+PROTEIN+S15 |
| 5 | RIBOSOME STRUCTURE | 17 | 50% | 1% | 25 | Search RIBOSOME+STRUCTURE | Search RIBOSOME+STRUCTURE |
| 6 | RIBOSOMAL PROTEIN S7 | 17 | 75% | 0% | 12 | Search RIBOSOMAL+PROTEIN+S7 | Search RIBOSOMAL+PROTEIN+S7 |
| 7 | RIBOSOME | 16 | 10% | 6% | 149 | Search RIBOSOME | Search RIBOSOME |
| 8 | PEPTIDYL TRANSFERASE | 16 | 48% | 1% | 24 | Search PEPTIDYL+TRANSFERASE | Search PEPTIDYL+TRANSFERASE |
| 9 | 30 S SUBUNIT | 14 | 100% | 0% | 7 | Search 30+S+SUBUNIT | Search 30+S+SUBUNIT |
| 10 | INSTANT EVOLUTION | 14 | 100% | 0% | 7 | Search INSTANT+EVOLUTION | Search INSTANT+EVOLUTION |
Key Words Plus |
| Rank | Web of Science journal category | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
|---|---|---|---|---|---|
| 1 | ESCHERICHIA COLI RIBOSOMES | 86 | 35% | 7% | 197 |
| 2 | ESCHERICHIA COLI RIBOSOME | 80 | 39% | 6% | 163 |
| 3 | MESSENGER RNA ANALOGS | 80 | 80% | 2% | 49 |
| 4 | 16 S RNA | 65 | 93% | 1% | 25 |
| 5 | 16S RNA | 65 | 71% | 2% | 53 |
| 6 | P SITE | 52 | 43% | 3% | 92 |
| 7 | P SITES | 50 | 58% | 2% | 58 |
| 8 | DIRECTED CROSS LINKING | 45 | 45% | 3% | 75 |
| 9 | A SITES | 44 | 61% | 2% | 47 |
| 10 | 3 MAJOR DOMAIN | 41 | 90% | 1% | 18 |
Journals |
Reviews |
| Title | Publ. year | Cit. | Active references | % act. ref. to same field |
|---|---|---|---|---|
| Crystal structure of the 30 S ribosomal subunit from Thermus thermophilus: Structure of the proteins and their interactions with 16 S RNA | 2002 | 241 | 120 | 77% |
| LESSONS FROM AN EVOLVING RIBOSOMAL-RNA - 16S AND 23S RIBOSOMAL-RNA STRUCTURES FROM A COMPARATIVE PERSPECTIVE | 1994 | 549 | 54 | 59% |
| Assembly of Bacterial Ribosomes | 2011 | 112 | 117 | 45% |
| Ribosomes and translation | 1997 | 305 | 172 | 76% |
| X-ray crystal structures of 70S ribosome functional complexes | 1999 | 444 | 91 | 62% |
| RIBOSOMAL-RNA AND TRANSLATION | 1991 | 394 | 91 | 79% |
| RNA folding and ribosome assembly | 2008 | 44 | 47 | 66% |
| Molecular movement inside the translational engine | 1998 | 143 | 64 | 67% |
| Ribosome biogenesis and the translation process in Eschetichia coli | 2007 | 149 | 170 | 25% |
| The Ribosome Emerges from a Black Box | 2014 | 7 | 54 | 50% |
Address terms |
| Rank | Address term | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
|---|---|---|---|---|---|
| 1 | AG RIBOSOMEN | 17 | 47% | 1.0% | 27 |
| 2 | ARRHENIUS S E5 | 9 | 64% | 0.3% | 9 |
| 3 | PROGRAM CELL MOL DEV BIOL BIOPHYS | 8 | 70% | 0.3% | 7 |
| 4 | MOL BIOL RNA | 5 | 16% | 1.1% | 31 |
| 5 | FAK MED MIKROBIOL | 4 | 44% | 0.3% | 7 |
| 6 | RNA REGULAT | 4 | 41% | 0.3% | 7 |
| 7 | JW WILSON | 4 | 33% | 0.3% | 9 |
| 8 | RIBOSOMAL STRUCT FUNCT | 3 | 100% | 0.1% | 3 |
| 9 | WITTMANN ABT | 3 | 60% | 0.1% | 3 |
| 10 | NAT SCI SECT | 2 | 27% | 0.3% | 7 |
Related classes at same level (level 1) |
| Rank | Relatedness score | Related classes |
|---|---|---|
| 1 | 0.0000220232 | ELONGATION FACTOR TU//EF G//ELONGATION FACTOR G |
| 2 | 0.0000190334 | RIBOSOMAL STALK//ACIDIC RIBOSOMAL PROTEIN//ACIDIC RIBOSOMAL P PROTEINS |
| 3 | 0.0000144297 | 5S RRNA//5S RIBOSOMAL RNA//ARBEITSGRP MAKROMOL STRUKTURANAL |
| 4 | 0.0000135870 | PLASMID PKK232 8//RIBOSOME RECONSTITUTION//16S 23S INTERGENIC SPACER |
| 5 | 0.0000121819 | SD SEQUENCE//RIBOSOMAL PROTEIN S1//SHINE DALGARNO SEQUENCE |
| 6 | 0.0000110496 | RIBOSOMAL PROTEIN//RIBOSOMAL PROTEIN MRNA//SURFEIT LOCUS |
| 7 | 0.0000104224 | POLARIZED TARGET//NA58//LITHIUM DEUTERIDE |
| 8 | 0.0000087571 | OBG//DRG2//BACTERIAL GTPASE |
| 9 | 0.0000066652 | FRAMESHIFTING//RIBOSOMAL FRAMESHIFTING//RNA PSEUDOKNOT |
| 10 | 0.0000051542 | RNA SECONDARY STRUCTURE//RNA SECONDARY STRUCTURE PREDICTION//RNA STRUCTURE PREDICTION |