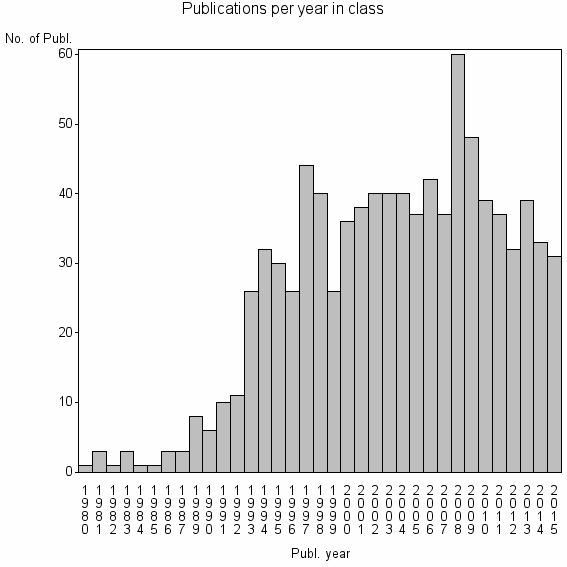
Class information for: |
Basic class information |
| ID | Publications | Average number of references |
Avg. shr. active ref. in WoS |
|---|---|---|---|
| 11591 | 904 | 47.2 | 86% |
Classes in level above (level 2) |
Terms with highest relevance score |
| Rank | Term | Type of term | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
|---|---|---|---|---|---|---|
| 1 | DSBA | Author keyword | 62 | 68% | 6% | 54 |
| 2 | DSBB | Author keyword | 38 | 89% | 2% | 17 |
| 3 | DSBD | Author keyword | 35 | 86% | 2% | 18 |
| 4 | CYTOCHROME C MATURATION | Author keyword | 31 | 76% | 2% | 22 |
| 5 | HEME LYASE | Author keyword | 30 | 100% | 1% | 12 |
| 6 | HEME ATTACHMENT | Author keyword | 27 | 92% | 1% | 11 |
| 7 | CYTOCHROME C BIOGENESIS | Author keyword | 27 | 74% | 2% | 20 |
| 8 | DSBC | Author keyword | 20 | 67% | 2% | 18 |
| 9 | PROTEIN DISULFIDE OXIDOREDUCTASE | Author keyword | 12 | 75% | 1% | 9 |
| 10 | DSB PROTEINS | Author keyword | 12 | 86% | 1% | 6 |
Web of Science journal categories |
Author Key Words |
| Rank | Web of Science journal category | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
LCSH search | Wikipedia search |
|---|---|---|---|---|---|---|---|
| 1 | DSBA | 62 | 68% | 6% | 54 | Search DSBA | Search DSBA |
| 2 | DSBB | 38 | 89% | 2% | 17 | Search DSBB | Search DSBB |
| 3 | DSBD | 35 | 86% | 2% | 18 | Search DSBD | Search DSBD |
| 4 | CYTOCHROME C MATURATION | 31 | 76% | 2% | 22 | Search CYTOCHROME+C+MATURATION | Search CYTOCHROME+C+MATURATION |
| 5 | HEME LYASE | 30 | 100% | 1% | 12 | Search HEME+LYASE | Search HEME+LYASE |
| 6 | HEME ATTACHMENT | 27 | 92% | 1% | 11 | Search HEME+ATTACHMENT | Search HEME+ATTACHMENT |
| 7 | CYTOCHROME C BIOGENESIS | 27 | 74% | 2% | 20 | Search CYTOCHROME+C+BIOGENESIS | Search CYTOCHROME+C+BIOGENESIS |
| 8 | DSBC | 20 | 67% | 2% | 18 | Search DSBC | Search DSBC |
| 9 | PROTEIN DISULFIDE OXIDOREDUCTASE | 12 | 75% | 1% | 9 | Search PROTEIN+DISULFIDE+OXIDOREDUCTASE | Search PROTEIN+DISULFIDE+OXIDOREDUCTASE |
| 10 | DSB PROTEINS | 12 | 86% | 1% | 6 | Search DSB+PROTEINS | Search DSB+PROTEINS |
Key Words Plus |
| Rank | Web of Science journal category | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
|---|---|---|---|---|---|
| 1 | FORMATION INVIVO | 111 | 64% | 12% | 110 |
| 2 | HEME CHAPERONE CCME | 73 | 88% | 4% | 35 |
| 3 | BOND FORMATION INVIVO | 69 | 72% | 6% | 54 |
| 4 | MEMBRANE PROTEIN DSBD | 58 | 92% | 3% | 23 |
| 5 | ISOMERASE LIKE PROTEIN | 57 | 95% | 2% | 19 |
| 6 | CYTOCHROME C MATURATION | 56 | 72% | 5% | 44 |
| 7 | CHAPERONE CCME | 49 | 94% | 2% | 17 |
| 8 | ISOMERASE DSBC | 49 | 94% | 2% | 17 |
| 9 | PERIPLASMIC THIOREDOXIN | 48 | 100% | 2% | 17 |
| 10 | FORMATION PATHWAY | 45 | 94% | 2% | 16 |
Journals |
Reviews |
| Title | Publ. year | Cit. | Active references | % act. ref. to same field |
|---|---|---|---|---|
| Protein disulfide bond formation in prokaryotes | 2003 | 344 | 100 | 78% |
| Cytochrome c biogenesis: the Ccm system | 2010 | 60 | 60 | 93% |
| Disulfide bond formation in prokaryotes: History, diversity and design | 2014 | 11 | 141 | 56% |
| Mechanisms of Oxidative Protein Folding in the Bacterial Cell Envelope | 2010 | 55 | 111 | 85% |
| Disulfide Bond Formation in the Bacterial Periplasm: Major Achievements and Challenges Ahead | 2013 | 21 | 48 | 67% |
| The disulfide bond formation (Dsb) system | 2008 | 78 | 62 | 79% |
| Composition and function of cytochrome c biogenesis System II | 2011 | 35 | 60 | 88% |
| DSB proteins and bacterial pathogenicity | 2009 | 90 | 98 | 66% |
| Cytochrome c Biogenesis: Mechanisms for Covalent Modifications and Trafficking of Heme and for Heme-Iron Redox Control | 2009 | 82 | 156 | 69% |
| Production of disulfide-bonded proteins in Escherichia coli | 2012 | 27 | 138 | 55% |
Address terms |
| Rank | Address term | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
|---|---|---|---|---|---|
| 1 | PROT CHEMHIGASHI KU | 2 | 67% | 0.2% | 2 |
| 2 | POST GENOME SCI | 2 | 43% | 0.3% | 3 |
| 3 | BCHM GRM | 1 | 100% | 0.2% | 2 |
| 4 | CRISTALLOGENESE CRISTALLISAT PROT | 1 | 100% | 0.2% | 2 |
| 5 | DIP BIOL STRUTTURALE FUNZ | 1 | 40% | 0.2% | 2 |
| 6 | ABORATING REFERENCE CAMPYLOBACTER | 1 | 50% | 0.1% | 1 |
| 7 | ANIM DIS HUMAN HLTH SICHUAN | 1 | 50% | 0.1% | 1 |
| 8 | AUSTRALAIN COUNCIL SPECIAL FUNCT | 1 | 50% | 0.1% | 1 |
| 9 | BEADLE N118 | 1 | 50% | 0.1% | 1 |
| 10 | BEADLE N200 | 1 | 50% | 0.1% | 1 |
Related classes at same level (level 1) |
| Rank | Relatedness score | Related classes |
|---|---|---|
| 1 | 0.0000216524 | PROTEIN DISULFIDE ISOMERASE//ERP29//PROTEIN DISULPHIDE ISOMERASE |
| 2 | 0.0000113651 | THIOREDOXIN H//PHOSPHORIBULOKINASE//CP12 |
| 3 | 0.0000102171 | GLUTAREDOXIN//THIOLTRANSFERASE//GLUTATHIONYLATION |
| 4 | 0.0000096898 | ALKALINE TRANSITION//ALKALINE ISOMERIZATION//CYTOCHROME C2 |
| 5 | 0.0000090165 | CYTOCHROME P460//N N BIS CARBOXYMETHYL N N DINITROSO 1 4 PHENYLENEDIAMINE//NITROSYLHEME |
| 6 | 0.0000082103 | CYTOCHROME C3//MULTIHEME CYTOCHROME//MULTIHEME CYTOCHROMES |
| 7 | 0.0000070955 | PPSR//CRTJ//FNRL |
| 8 | 0.0000066423 | HTRA1//HTRA//HTRA2 |
| 9 | 0.0000064034 | BPTI//BAKER CHEM CHEM BIOL//SCRAMBLED PROTEINS |
| 10 | 0.0000061006 | AUGMENTER OF LIVER REGENERATION//HEPATIC STIMULATOR SUBSTANCE//SULFHYDRYL OXIDASE |