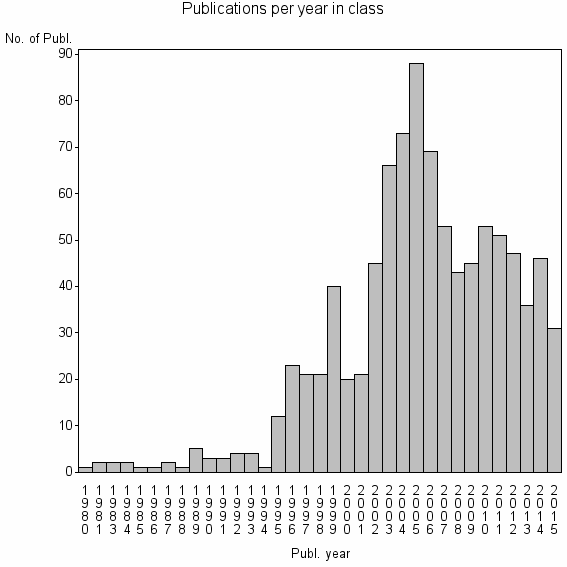
Class information for: |
Basic class information |
| ID | Publications | Average number of references |
Avg. shr. active ref. in WoS |
|---|---|---|---|
| 11177 | 936 | 20.6 | 46% |
Classes in level above (level 2) |
| ID, lev. above |
Publications | Label for level above |
|---|---|---|
| 1160 | 9033 | MEMBRANE COMPUTING//GRP NAT COMP//THEORETICAL COMPUTER SCIENCE |
Terms with highest relevance score |
| Rank | Term | Type of term | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
|---|---|---|---|---|---|---|
| 1 | DNA COMPUTING | Author keyword | 120 | 45% | 21% | 201 |
| 2 | SPLICING SYSTEMS | Author keyword | 44 | 85% | 2% | 23 |
| 3 | ADLEMAN LIPTON MODEL | Author keyword | 42 | 94% | 2% | 15 |
| 4 | DNA COMPUTATION | Author keyword | 33 | 56% | 4% | 40 |
| 5 | PL DNA COMP | Address | 24 | 91% | 1% | 10 |
| 6 | DNA BASED COMPUTING | Author keyword | 21 | 75% | 2% | 15 |
| 7 | HAIRPIN COMPLETION | Author keyword | 18 | 83% | 1% | 10 |
| 8 | STICKER MODEL | Author keyword | 15 | 68% | 1% | 13 |
| 9 | EVOLUTIONARY PROCESSOR | Author keyword | 15 | 88% | 1% | 7 |
| 10 | SPLICING OPERATION | Author keyword | 15 | 88% | 1% | 7 |
Web of Science journal categories |
Author Key Words |
| Rank | Web of Science journal category | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
LCSH search | Wikipedia search |
|---|---|---|---|---|---|---|---|
| 1 | DNA COMPUTING | 120 | 45% | 21% | 201 | Search DNA+COMPUTING | Search DNA+COMPUTING |
| 2 | SPLICING SYSTEMS | 44 | 85% | 2% | 23 | Search SPLICING+SYSTEMS | Search SPLICING+SYSTEMS |
| 3 | ADLEMAN LIPTON MODEL | 42 | 94% | 2% | 15 | Search ADLEMAN+LIPTON+MODEL | Search ADLEMAN+LIPTON+MODEL |
| 4 | DNA COMPUTATION | 33 | 56% | 4% | 40 | Search DNA+COMPUTATION | Search DNA+COMPUTATION |
| 5 | DNA BASED COMPUTING | 21 | 75% | 2% | 15 | Search DNA+BASED+COMPUTING | Search DNA+BASED+COMPUTING |
| 6 | HAIRPIN COMPLETION | 18 | 83% | 1% | 10 | Search HAIRPIN+COMPLETION | Search HAIRPIN+COMPLETION |
| 7 | STICKER MODEL | 15 | 68% | 1% | 13 | Search STICKER+MODEL | Search STICKER+MODEL |
| 8 | EVOLUTIONARY PROCESSOR | 15 | 88% | 1% | 7 | Search EVOLUTIONARY+PROCESSOR | Search EVOLUTIONARY+PROCESSOR |
| 9 | SPLICING OPERATION | 15 | 88% | 1% | 7 | Search SPLICING+OPERATION | Search SPLICING+OPERATION |
| 10 | MOLECULAR COMPUTING | 14 | 29% | 4% | 40 | Search MOLECULAR+COMPUTING | Search MOLECULAR+COMPUTING |
Key Words Plus |
| Rank | Web of Science journal category | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
|---|---|---|---|---|---|
| 1 | MOLECULAR COMPUTATION | 87 | 51% | 13% | 123 |
| 2 | DNA SOLUTION | 28 | 64% | 3% | 28 |
| 3 | ONE SIDED CONTEXTS | 11 | 100% | 1% | 6 |
| 4 | WORD DESIGN STRATEGY | 9 | 83% | 1% | 5 |
| 5 | STOCHASTIC LOCAL SEARCH | 7 | 57% | 1% | 8 |
| 6 | DELETION SYSTEMS | 6 | 80% | 0% | 4 |
| 7 | LEFT INSTRUCTIONS | 6 | 80% | 0% | 4 |
| 8 | WORD DESIGN | 6 | 80% | 0% | 4 |
| 9 | HYBRID NETWORKS | 6 | 41% | 1% | 12 |
| 10 | ADLEMAN LIPTON MODEL | 5 | 63% | 1% | 5 |
Journals |
Reviews |
| Title | Publ. year | Cit. | Active references |
% act. ref. to same field |
|---|---|---|---|---|
| A novel text and image encryption method based on chaos theory and DNA computing | 2013 | 6 | 7 | 86% |
| New field of cryptography: DNA cryptography | 2006 | 28 | 17 | 53% |
| A review on DNA computing models | 2007 | 14 | 35 | 89% |
| A critical review on waste paper sorting techniques | 2014 | 3 | 14 | 29% |
| Sticker DNA computer model - Part I: Theory | 2004 | 12 | 14 | 100% |
| Molecular computation and evolutionary wetware: A cutting-edge technology for artificial life and nanobiotechnologies | 2007 | 5 | 38 | 42% |
| Biologically Relevant Molecular Finite Automata | 2011 | 1 | 29 | 48% |
| Sticker DNA computer model - Part II: Application | 2004 | 1 | 12 | 75% |
| From molecular biology to nanotechnology and nanomedicine | 2002 | 76 | 80 | 14% |
| The emerging discipline of biomolecular computation in the US | 2002 | 7 | 30 | 50% |
Address terms |
| Rank | Address term | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
|---|---|---|---|---|---|
| 1 | PL DNA COMP | 24 | 91% | 1.1% | 10 |
| 2 | GRP MATH LINGUIST | 11 | 25% | 4.2% | 39 |
| 3 | GRP INFORMAT FONDAMENTALE METZ | 4 | 75% | 0.3% | 3 |
| 4 | BIOMOL COMP | 4 | 56% | 0.5% | 5 |
| 5 | BIO X DNA COMP CONSORTIUM | 3 | 57% | 0.4% | 4 |
| 6 | AFFAIR OFF | 3 | 50% | 0.4% | 4 |
| 7 | COMPUTAT BIOMODELLING | 3 | 60% | 0.3% | 3 |
| 8 | FORMAL METHODS COMP SCI FMI | 3 | 60% | 0.3% | 3 |
| 9 | ADV DESIGN INTELLIGENT COMP | 2 | 16% | 1.5% | 14 |
| 10 | LANGUAGES PROJECTS COMP INFORMAT SYST | 2 | 67% | 0.2% | 2 |
Related classes at same level (level 1) |
| Rank | Relatedness score | Related classes |
|---|---|---|
| 1 | 0.0000152985 | MEMBRANE COMPUTING//GRP NAT COMP//P SYSTEMS |
| 2 | 0.0000131679 | PRIMITIVE WORDS//UNAVOIDABLE SETS//SHUFFLE ON TRAJECTORIES |
| 3 | 0.0000130774 | DNA ORIGAMI//DNA NANOTECHNOLOGY//DNA NANOSTRUCTURES |
| 4 | 0.0000078711 | ENZYME LOGIC//MOLECULAR LOGIC//LOGIC GATES |
| 5 | 0.0000057876 | TECHNOL MANAGEMENT POLICY PROGRAMME//TECHM P//PERVAS ICT |
| 6 | 0.0000054072 | GENERATIVE POWER//PARALLEL COMMUNICATING GRAMMAR SYSTEMS//GRAMMAR SYSTEMS |
| 7 | 0.0000052273 | NONSPECIFIC HYBRIDIZATIONS//NEAREST NEIGHBOR PROPERTIES//NUCLEIC ACID OLIGOMERS |
| 8 | 0.0000049541 | POST CORRESPONDENCE PROBLEM//WATSON CRICK COMPLEMENTARITY//D0L SYSTEM |
| 9 | 0.0000048301 | PICTURE LANGUAGES//RANDOM CONTEXT GRAMMARS//TWO DIMENSIONAL LANGUAGES |
| 10 | 0.0000041728 | SYNCHRONIZING AUTOMATA//SYNCHRONIZING WORD//CERNY CONJECTURE |